Abstract
There is evidence that progesterone plays a role in the aetiology of invasive epithelial ovarian cancer. Therefore, genes involved in pathways that regulate progesterone may be candidates for susceptibility to this disease. Previous studies have suggested that genetic variants in the progesterone receptor gene (PGR) may be associated with ovarian cancer risk, although results have been inconsistent. We have established an international consortium to pool resources and data from many ovarian cancer case–control studies in an effort to identify variants that influence risk. In this study, three PGR single nucleotide polymorphisms (SNPs), for which previous data have suggested they affect ovarian cancer risk, were examined. These were +331 C/T (rs10895068), PROGINS (rs1042838), and a 3′ variant (rs608995). A total of 4788 ovarian cancer cases and 7614 controls from 12 case–control studies were included in this analysis. Unconditional logistic regression was used to model the association between each SNP and ovarian cancer risk and two-sided P-values are reported. Overall, risk of ovarian cancer was not associated with any of the three variants studied. However, in histopathological subtype analyses, we found a statistically significant association between risk of endometrioid ovarian cancer and the PROGINS allele (n=651, OR=1.17, 95% CI=1.01–1.36, P=0.036). We also observed borderline evidence of an association between risk of endometrioid ovarian cancer and the +331C/T variant (n=725 cases; OR=0.80, 95% CI 0.62–1.04, P=0.100). These data suggest that while these three variants in the PGR are not associated with ovarian cancer overall, the PROGINS variant may play a modest role in risk of endometrioid ovarian cancer.
Similar content being viewed by others
Main
Several lines of evidence support a role for progesterone in the aetiology of ovarian cancer (Risch, 1998). Epidemiological studies have consistently shown a significant protective effect of parity. The protective effect increases steadily with each birth and pregnancy is associated with high progesterone levels (Hartge et al, 1989; Cooper et al, 1999; Titus-Ernstoff et al, 2001; Whiteman et al, 2003; Pike et al, 2004). In the third trimester, progesterone levels are some 10–15 times higher than in the luteal phase of the normal menstrual cycle. Oral contraceptives are also protective against ovarian cancer (Study, 1987; Rosenblatt et al, 1992, 1994; Ness et al, 2000; Royar et al, 2001; Schildkraut et al, 2002) and use of progestin-containing oral contraceptives increases average circulating progesterone levels to 9.2 ngm−1 compared to ∼3.5 ngm−1 during the normal menstrual cycle (Norman and Litwack, 1997). The protective effect of oral contraceptives per month of use is less than the protection from births, in line with the concentrations of progesterone. There is also some evidence that oral contraceptives with higher progestin content afford more protection against ovarian cancer (Schildkraut et al, 2002).
Animal models and in vitro data also suggest that progesterone has a significant influence on the ovary and on ovarian cancer. Studies in macaques suggest an apoptotic effect of progestins on the surface of the ovary (Rodriguez et al, 1998). In vitro treatment of both benign and malignant ovarian tumour cells with progestins results in an antiproliferative response (Zhou et al, 2002).
Progesterone binds to the progesterone receptor (PR) to initiate signalling. Two progesterone receptor isoforms (PR-A, PR-B) are encoded by a single gene (PGR). Except for a 164 amino-acid sequence at the N-terminal end of PR-B that is absent from PR-A, the PR isoforms are identical but their actions are divergent (Kastner et al, 1990). PR-B acts as a transcription activator whereas PR-A inhibits PR-B (and other members of the nuclear receptor superfamily) (Vegeto et al, 1993).
The PGR has long been hypothesised as a candidate gene for ovarian cancer susceptibility and its variation has been widely studied. Originally, an ALU in intron 7 named PROGINS was identified and found to be associated with increased risk of ovarian cancer (McKenna et al, 1995; Rowe et al, 1995). Subsequent characterisation of the coding region of the gene identified a non-synonymous single nucleotide polymorphism (SNP) in exon 4 and a synonymous SNP in exon 5 that were in perfect linkage disequilibrium with the PROGINS (De Vivo et al, 2002). The PROGINS (or variants in which it is in perfect linkage disequilibrium) has been studied by many groups in relation to ovarian cancer risk. The results are, however, equivocal (McKenna et al, 1995; Manolitsas et al, 1997; Lancaster et al, 1998, 2003; Spurdle et al, 2001; Tong et al, 2001; Agoulnik et al, 2004; Pearce et al, 2005; Terry et al, 2005; Romano et al, 2006). Pearce et al (2005), suggested that a variant 3′ of the PGR (rs608995), in partial linkage disequilibrium with the PROGINS, might be a better marker of ovarian cancer risk, but this has not been confirmed by other investigators.
In addition, a putative functional SNP, +331C/T (sometimes denoted as +331G/A), in the promoter region of the PGR that may affect the relative transcription of the PR-A and PR-B isoforms has been found to be associated with a reduced risk of ovarian cancer in studies from North Carolina and Australia (Berchuck et al, 2004). This association was particularly strong among clear cell/endometrioid subtypes. However, Risch et al (2006) observed an increased risk of ovarian cancer associated with this SNP.
The inconsistent results with the PROGINS and the +331C/T SNP are not surprising. Genetic association studies are plagued by conflicting results that can be explained by heterogeneity across study populations as well as false-positive and -negative results. Lohmueller et al (2003) demonstrated that approximately two-thirds of genetic associations do not hold up on meta-analysis. Large sample sizes and pooling of data are therefore critical to evaluate the association between a phenotype and genetic variation with confidence.
To clarify the association between variation at the PGR locus and ovarian cancer risk, including histological subtype associations, 12 groups from the Ovarian Cancer Association Consortium (OCAC) have pooled their data to examine the +331C/T variant (rs10895068), the PROGINS allele (measured by the exon 4 non-synonymous SNP; rs1042838) and a variant 3′ of the PGR (rs608995) in relation to ovarian cancer risk. The results are reported here.
Materials and methods
Approval and consent
All study participants provided written informed consent prior to the collection of biological samples or interview/clinical data. Each group involved in the OCAC has Institutional Review Board/ethics approval for this analysis and the University of Southern California and Duke University have Institutional Review Board approval to serve as data coordinating centres for the OCAC.
Study populations
The OCAC comprises investigators who collaborate on promising genetic associations by combining data from their individual ovarian cancer case–control studies. The participating groups for this PGR study are the Australian Cancer Study, (Merritt et al, 2008) the Australian Ovarian Cancer Study (Merritt et al, 2008), the Connecticut Ovary Study (CONN) (Risch et al, 2006), the Family Registry for Ovarian Cancer Study (Auranen et al, 2005; Song et al, 2006), the Hormones and Ovarian Cancer Prediction Study, the Danish Malignant Ovarian Cancer Study (MALOVA) (Auranen et al, 2005; Song et al, 2006), the Mayo Clinic Ovarian Cancer Case–Control Study (Sellers et al, 2005), the North Carolina Ovarian Cancer Study (Berchuck et al, 2004), the New England-based Case–Control Study (NECC) (Terry et al, 2005), the Polish Ovarian Cancer Study (POCS) (García-Closas et al, 2007), the UK SEARCH Ovarian Cancer Study (SEARCH) (Auranen et al, 2005; Song et al, 2006) and the USC/Los Angeles County Case–Control Studies of Ovarian Cancer (USC) (Pearce et al, 2005). Details of these studies have been published previously (Gayther et al, 2007); Table 1 shows the basic information for each study. The cases analysed here are restricted to women diagnosed with invasive epithelial ovarian cancer.
Genotyping and quality control
The three SNPs genotyped in this study were the +331C/T (rs10895068), PROGINS (measured by the exon 4 SNP rs1042838) and rs608995 (a variant 3′ of the PGR). The allele designations are based on the forward strand as given in the University of California at Santa Cruz genome browser.
All groups used the 5′ nuclease Taqman allelic discrimination assay (Taqman; Applied Biosystems, Foster City, CA, USA) to genotype samples with the exception of the Australian Cancer Study and Australian Ovarian Cancer Study, which used the iPlex Sequenom MassArray system (Sequenom Inc., San Diego, CA, USA), CONN that used dot blotting (Risch et al, 2006), and Mayo Clinic Ovarian Cancer Case–Control Study that used Pyrosequencing for PROGINS and rs608995.
To confirm that laboratory to laboratory quality control was adequate, five SNPs were genotyped in the HAPMAPPT01 panel of CEPH-Utah trios-standard plate provided by Coriell (http://locus.umdny.edu/nigms/nigms_cgi/panel.cgi?id=2&query=HAPMAP01). This 96-well plate contains 90 different DNA samples, five duplicate samples, and a negative template control. Genotyping call rates and concordance between studies were compared. Call rates for these five SNPs ranged from 96 to 99% and the concordance of results across the laboratories was >99%.
Hardy–Weinberg Equilibrium (HWE) was checked among controls by the racial/ethnic group. Data from one study (CONN) for two of the SNPs (PROGINS and rs608995) were excluded for gross deviations (P<10−4) from HWE. The genotyping calls for studies with minor deviations from HWE (0.01<P<0.05) were examined to monitor the quality of the genotyping. There were no obvious reasons for deviation from HWE (e.g., genotyping irregularities), and therefore the minor deviations were assumed most likely due to chance. In addition, results were unchanged when excluding those studies with HWE P-values between 0.01 and 0.05 (data not shown). Concordance between duplicate samples was 100% across all studies for the three variants for all data included in these analyses.
Results were available for 12 groups for the +331C/T variant. The NECC study did not genotype rs608995. Results for the PROGINS and rs608995 were excluded for CONN due to significant deviations from HWE. Therefore, results were available for 11 studies for the PROGINS and 10 studies for the rs608995 3′ variant.
Statistical analysis
The variables available for this analysis were study, race/ethnicity (White, Latina, African-American), age, stage of disease (FIGO), histology (serous, mucinous, clear cell, and endometrioid), and time from diagnosis to blood collection (cases only).
Unconditional logistic regression was used to model the association between each SNP and risk of ovarian cancer stratified on study, age, and race/ethnicity. All single SNP models were log additive. Goodness of fit P-values were calculated to evaluate heterogeneity across the study populations. Statistical analyses were carried out using both SAS (Version 9, Cary, NC, USA) and STATA (Version 9, StataCorp, College Station TX, USA). All statistical significance levels (P-values) quoted are two-sided. All odds ratios are expressed per copy of the minor allele.
Results
A total of 4788 invasive epithelial ovarian cancer cases and 7614 controls were available for the current analysis (Table 1). Overall, 92.0% of cases and 93.0% of controls were White and the mean ages were 56.7 and 54.3 years respectively. Information on stage at diagnosis was available on 73.7% of cases, the majority of which were FIGO stage III/IV (63.0%) and 55.5% had a serous histology (Supplementary Table 1).
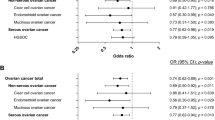
Across the studies, the minor allele frequencies in White controls ranged from 4.6 to 7.3% for +331C/T (rs10895068), 9.2 to 19.0% for the PROGINS (rs1042838) and 20.0 to 26.6% for the 3′ variant (rs608995; Supplementary Table 2). The study-specific and summary effect estimates are shown in Figure 1 for all cases and endometrioid subtype associations.
There was no association with the +331C/T (rs10895068) variant among all cases (per allele OR=1.00; 95% CI 0.89–1.13; P=1.0; Table 2a). In cell type-specific subgroup analyses, a suggestive association was observed with carrying a T allele and risk of endometrioid invasive ovarian cancers (per allele OR=0.80; 95% CI 0.62–1.04; P=0.100; Figure 1). Risk of clear cell ovarian cancer with this variant was reduced to a similar degree (OR=0.83; Table 2b). No associations were observed between serous or mucinous subtypes and this allele (Table 2b).
No overall association was observed with risk of ovarian cancer and the PROGINS allele (rs1042838; OR=1.04; 95% CI 0.96–1.12; P=0.38; Table 2a). However, risk was statistically significantly elevated among endometrioid ovarian cancer cases (OR=1.17, 95% CI 1.01–1.36, P=0.036; Table 2b).
In a joint effects analysis, risk of endometrioid ovarian cancer associated with the PROGINS was observed only among non-carriers of the +331 minor allele (OR=1.22, 95% CI 1.01–1.46, P=0.037). Although not statistically significant, the protective effect of the +331 minor allele persisted among non-carriers of the PROGINS (OR=0.76, 95% CI 0.55–1.06, P=0.11) and carriers of the PROGINS (OR=0.79, 95% CI 0.40–1.57, P=0.50).
No statistically significant association was observed between the 3′ variant (rs608995) and risk of ovarian cancer when all cases were considered (Table 2a). In subtype analysis, a borderline statistically significant association was observed between endometrioid cases and the rs608995 variant (OR=1.14, 95% CI 0.99–1.31, P=0.076), however, this effect was limited to individuals also carrying at least one copy of the PROGINS (data not shown).
Discussion
Since the publication of the first paper examining the relationship between the PROGINS and ovarian cancer risk more than 10 years ago, there has been substantial interest in the role of the PGR in risk of this disease. We have evaluated three SNPs, +331C/T (rs10895068), PROGINS (rs1042838), and a 3′ variant (rs608995), in the PGR in a pooled ovarian cancer dataset from 12 groups around the world and have found no overall role for this gene in disease risk.
The pooled analysis does provide statistically significant evidence of an association between the PROGINS and risk of invasive endometrioid ovarian cancer. The restriction of an association to this subtype only provides an explanation for the equivocal nature of the published results on the PROGINS and ovarian cancer risk, given that the proportion of endometrioid ovarian cancer cases likely varied by published study and typically accounts for no more than 15–20% of cases.
We also found suggestive evidence of an association between endometrioid ovarian cancer and the +331C/T variant (OR=0.80, 95% CI 0.62–1.04, P=0.100). As suggested by Berchuck et al (2004), combining endometrioid and clear cell histologies in which the effect is similar, resulted in a borderline statistically significant association (n=1088 cases, OR=0.81, 95% CI 0.65–1.01, P=0.058).
Pearce et al (2005) had previously suggested that rs608995 may explain the PROGINS-ovarian cancer association, however, in this larger dataset in which the effect was restricted to endometrioid cases, this was not supported. When examining the joint effects of the PROGINS and rs608995, the OR for endometrioid ovarian cancer associated with the rs608995 minor allele was 0.79 (95% CI 0.59–1.07, P=0.12) in the absence of the PROGINS allele. This suggests that the PROGINS allele or a marker in linkage disequilibrium with the PROGINS is responsible for the association and not the rs608995 variant.
Both the +331 variant and the PROGINS have been studied with regard to their functional effect. The T allele of the +331 favours an increase in the transcription of PR-B relative to PR-A (De Vivo et al, 2002); PR-B acts as a classic steroid receptor whereas PR-A acts as a repressor of both PR-B and other steroid receptors. PR-A therefore may lessen overall progesterone responsiveness through its repressive effect. Any variation which increases PR-B relative to PR-A may reduce risk of ovarian cancer by increasing exposure to the beneficial effects of progesterone. In a small study of 107 ovarian cancer cases, decreased risk of death was observed among cases positive for PRB (labelling index>10) relative to cases negative for PRB (P=0.037). However, this finding was amongst all cell types (Akahira et al, 2000). There is also a suggestion that the PROGINS allele as defined by the V660L exon five variant (as examined in the present study) decreases overall response to progesterone which would be consistent with an increased risk of disease associated with this variant (Romano et al, 2007).
In this collaborative effort, there were 4788 ovarian cancer cases, of which 766 (16.0%) were endometrioid tumours. With the samples sizes available in this current OCAC study, we had 80% power to detect odds ratios of 0.83 and 1.12 for the +331, and PROGINS variants, respectively for all cases using a log additive genetic model and a two-sided α of 0.05. Among endometrioid subtypes, we had 80% power to detect odds ratios of 0.67 and 1.25 for the +331 and PROGINS variants, respectively using a log additive genetic model and a two-sided α of 0.05. Although the power in the current OCAC study is still quite limited, it underscores the importance of collaborative efforts, as the largest individual OCAC study had only 124 endometrioid ovarian cancers. Thus the power of subgroups analyses is clear and will be enhanced in the future with continued patient accrual to existing studies and additional investigators contributing to OCAC studies.
Alternatively, the findings of an association with the +331C/T and PROGINS, variants with the endometrioid histology may simply be due to chance. By assigning priors of 0.05, 0.10, and 0.15, the resulting false positive report probabilities (Wacholder et al, 2004) are approximately 0.78, 0.62, and 0.52 for the +331C/T variant and 0.61, 0.42, and 0.32 for the PROGINS, respectively. Thus they may represent false positive findings.
Our analysis is the largest report describing the association between ovarian cancer risk and variants in the PGR. However, there remain several limitations to the study. For example, it is possible that environmental modifiers, such as oral contraceptive use, may be important in refining the PGR ovarian cancer risk associations and such analyses are planned in the future. There are also weaknesses of this study. Firstly, there are variable participation rates for cases between studies (Table 1). If any or all of the variants analysed is related to survival, then the low participation rates among cases might be expected to influence the results. Efforts to evaluate this include the analyses of data stratified by FIGO stage and time from diagnosis to blood collection. None of the results differed significantly when conducting these analyses. Second, as is the nature of collaborative projects, each study had a different level of pathology review and random misclassification cannot be ruled out, which would bias results towards the null in histologic-specific analyses suggesting that our results may be attenuated. Lastly, while we evaluated the best PGR candidate variants suggested by the literature, it remains possible that other, as yet unidentified variants at the locus, influence ovarian cancer risk.
Also, we observed significant heterogeneity of effect for the PROGINS allele and risk of ovarian cancer overall. Evaluation of the heterogeneity by removing one study at a time revealed that the NECC study population had a significantly different odds ratio (OR=0.75, heterogeneity P=0.011) from the other 10 OCAC studies. We investigated possible explanations for the heterogeneity we observed in the NECC study, but the reason could not be elucidated. Genotyping error is the most likely reason for experimental bias towards the null. Therefore, we regenotyped the PROGINS allele in the NECC case–control study. The results were 98% concordant with the original genotyping data, ruling this out as an explanation. Also, standard epidemiological risk and protective factors are observed with the NECC study suggesting no coding errors in the data with respect to case–control status. Further stratification of White race by Jewish ancestry was done and the results were consistent across Jewish and non-Jewish Whites (data not shown). The age distribution and participation rates are consistent with the other OCAC studies (Table 1). This heterogeneity may simply be due to chance.
Heterogeneity was also present with the +331 variant and endometrioid ovarian cancer, however no single study accounted for this heterogeneity. The minor allele frequency of this SNP is approximately 5% and the fluctuations in the data may simply represent chance; further follow-up is needed.
If these are true results and variation at the PGR locus is associated with endometrioid ovarian cancer only, then it has implications for the identification of moderate risk genes for ovarian cancer. In the past, ovarian cancer has frequently been treated as a single-disease entity for genetic association studies, mainly because studies have been too small to perform subtype analyses that are substantially powered. However, there is a large body of evidence that indicates different germline and somatic genetic factors contribute to different histological subtypes of ovarian cancer. For example, BRCA1 mutation carriers appear to predispose to serous ovarian cancers (Pal et al, 2005); mutations in the PTEN tumour suppressor gene are more associated with endometrioid ovarian cancers (Obata et al, 1998); and K-ras mutations are more common in mucinous tumours than in either serous of endometrioid subtypes (Gemignani et al, 2003).
In conclusion, in the present analysis, we were able to exclude an overall effect of these variants in the PGR with risk of ovarian cancer. However, our evidence suggests histology-specific effects, demonstrating the necessity of data pooling to examine subgroup effects for this cancer. Although the PROGINS is unlikely to represent appreciable susceptibility risk factor, given the restriction of the association to endometrioid histology, the magnitude of the observed odds ratio, and the modest allele frequency of this variant, further analysis of this gene with regard to the endometrioid subtype is warranted to provide insight into the mechanisms underlying disease aetiology.
Change history
16 November 2011
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Agoulnik IU, Tong XW, Fischer DC, Korner K, Atkinson NE, Edwards DP, Headon DR, Weigel NL, Kieback DG (2004) A germline variation in the progesterone receptor gene increases transcriptional activity and may modify ovarian cancer risk. J Clin Endocrinol Metab 89: 6340–6347
Akahira J, Inoue T, Suzuki T, Ito K, Konno R, Sato S, Moriya T, Okamura K, Yajima A, Sasano H (2000) Progesterone receptor isoforms A and B in human epithelial ovarian carcinoma: immunohistochemical and RT-PCR studies. Br J Cancer 83: 1488–1494
Auranen A, Song H, Waterfall C, Dicioccio RA, Kuschel B, Kjaer SK, Hogdall E, Hogdall C, Stratton J, Whittemore AS, Easton DF, Ponder BA, Novik KL, Dunning AM, Gayther S, Pharoah PD (2005) Polymorphisms in DNA repair genes and epithelial ovarian cancer risk. Int J Cancer 117: 611–618
Berchuck A, Schildkraut JM, Wenham RM, Calingaert B, Ali S, Henriott A, Halabi S, Rodriguez GC, Gertig D, Purdie DM, Kelemen L, Spurdle AB, Marks J, Chenevix-Trench G (2004) Progesterone receptor promoter +331A polymorphism is associated with a reduced risk of endometrioid and clear cell ovarian cancers. Cancer Epidemiol Biomarkers Prev 13: 2141–2147
Cooper GS, Schildkraut JM, Whittemore AS, Marchbanks PA (1999) Pregnancy recency and risk of ovarian cancer. Cancer Causes Control 10: 397–402
De Vivo I, Huggins GS, Hankinson SE, Lescault PJ, Boezen M, Colditz GA, Hunter DJ (2002) A functional polymorphism in the promoter of the progesterone receptor gene associated with endometrial cancer risk. Proc Natl Acad Sci USA 99: 12263–12268
García-Closas M, Brinton LA, Lissowska J, Sherman ME, Szeszenia-Dabrowska N, Peplonska B, Welch R, Yeager M, Bardin-Mikolajczak A, Zatonski W, Chanock SJ (2007) Ovarian cancer risk and common variation in the sex hormone-binding globulin gene: a population-based case–control study. BMC Cancer 7: 60
Gayther SA, Song H, Ramus SJ, Kjaer SK, Whittemore AS, Quaye L, Tyrer J, Shadforth D, Hogdall E, Hogdall C, Blaeker J, DiCioccio R, McGuire V, Webb PM, Beesley J, Green AC, Whiteman DC, Australian Ovarian Cancer Study Group, Goodman MT, Lurie G, Carney ME, Modugno F, Ness RB, Edwards RP, Moysich KB, Goode EL, Couch FJ, Cunningham JM, Sellers TA, Wu AH, Pike MC, Iversen ES, Marks JR, Garcia-Closas M, Brinton L, Lissowska J, Peplonska B, Easton DF, Jacobs I, Ponder BA, Schildkraut J, Pearce CL, Chenevix-Trench G, Berchuck A, Pharoah PDP, Ovarian Cancer Association Consortium (2007) Tagging single nucleotide polymorphisms in cell cycle control genes and susceptibility to invasive epithelial ovarian cancer. Cancer Res 67: 3027–3035
Gemignani ML, Schlaerth AC, Bogomolniy F, Barakat RR, Lin O, Soslow R, Venkatraman E, Boyd J (2003) Role of KRAS and BRAF gene mutations in mucinous ovarian carcinoma. Gynecol Oncol 90: 378–381
Hartge P, Schiffman MH, Hoover R, McGowan L, Lesher L, Norris HJ (1989) A case–control study of epithelial ovarian cancer. Am J Obstet Gynecol 161: 10–16
Kastner P, Krust A, Turcotte B, Stropp U, Tora L, Gronemeyer H, Chambon P (1990) Two distinct estrogen-regulated promoters generate transcripts encoding the two functionally different human progesterone receptor forms A and B. EMBO J 9: 1603–1614
Lancaster JM, Berchuck A, Carney ME, Wiseman R, Taylor JA (1998) Progesterone receptor gene polymorphism and risk for breast and ovarian cancer. Br J Cancer 78: 277
Lancaster JM, Wenham RM, Halabi S, Calingaert B, Marks JR, Moorman PG, Bentley RC, Berchuck A, Schildkraut JM (2003) No relationship between ovarian cancer risk and progesterone receptor gene polymorphism in a population-based, case–control study in North Carolina. Cancer Epidemiol Biomarkers Prev 12: 226–227
Lohmueller KE, Pearce CL, Pike M, Lander ES, Hirschhorn JN (2003) Meta-analysis of genetic association studies supports a contribution of common variants to susceptibility to common disease. Nat Genet 33: 177–182
Manolitsas TP, Englefield P, Eccles DM, Campbell IG (1997) No association of a 306-bp insertion polymorphism in the progesterone receptor gene with ovarian and breast cancer. Br J Cancer 75: 1398–1399
Merritt MA, Green AC, Nagle CM, Webb PM, Australian Cancer Study (Ovarian Cancer), Australian Ovarian Cancer Study Group (2008) Talcum powder, chronic pelvic inflammation and NSAIDs in relation to risk of epithelial ovarian cancer. Int J Cancer 122: 170–176
McKenna NJ, Kieback DG, Carney DN, Fanning M, McLinden J, Headon DR (1995) A germline TaqI restriction fragment length polymorphism in the progesterone receptor gene in ovarian carcinoma. Br J Cancer 71: 451–455
Ness RB, Grisso JA, Klapper J, Schlesselman JJ, Silberzweig S, Vergona R, Morgan M, Wheeler JE (2000) Risk of ovarian cancer in relation to estrogen and progestin dose and use characteristics of oral contraceptives. SHARE study group. Steroid Hormones and reproductions. Am J Epidemiol 152: 233–241
Norman AW, Litwack G (1997) Estrogens and progestins. In Hormones Norman AW, Litwack G (eds), pp 361–386. San Diego, CA: Academic Press
Obata K, Morland SJ, Watson RH, Hitchcock A, Chenevix-Trench G, Thomas EJ, Campbell IG (1998) Frequent PTEN/MMAC mutations in endometrioid but not serous or mucinous epithelial ovarian tumors. Cancer Res 58: 2095–2097
Pal T, Permuth-Wey J, Betts JA, Krischer JP, Fiorica J, Arango H, LaPolla J, Hoffman M, Martino MA, Wakeley K, Wilbanks G, Nicosia S, Cantor A, Sutphen R (2005) BRCA1 and BRCA2 mutations account for a large proportion of ovarian carcinoma cases. Cancer 104: 2807–2816
Pearce CL, Hirschhorn JN, Wu AH, Burtt NP, Stram DO, Young S, Kolonel LN, Henderson BE, Altshuler D, Pike MC (2005) Clarifying the PROGINS allele association in ovarian and breast cancer risk: a haplotype-based analysis. J Natl Cancer Inst 97: 51–59
Pike MC, Pearce CL, Peters R, Cozen W, Wan P, Wu AH (2004) Hormonal factors and the risk of invasive ovarian cancer: a population-based case–control study. Fertil Steril 82: 186–195
Risch HA (1998) Hormonal etiology of epithelial ovarian cancer, with a hypothesis concerning the role of androgens and progesterone. J Natl Cancer Inst 90: 1774–1786
Risch HA, Bale AE, Beck PA, Zheng W (2006) PGR +331 A/G and increased risk of epithelial ovarian cancer. Cancer Epidemiol Biomarkers Prev 15: 1738–1741
Rodriguez GC, Walmer DK, Cline M, Krigman H, Lessey BA, Whitaker RS, Dodge R, Hughes CL (1998) Effect of progestin on the ovarian epithelium of macaques: cancer prevention through apoptosis? J Soc Gynecol Investig 5: 271–276
Romano A, Delvoux B, Fischer DC, Groothuis P (2007) The PROGINS polymorphism of the human progesterone receptor diminishes the response to progesterone. J Mol Endocrinol 38: 331–350
Romano A, Lindsey PJ, Fischer DC, Delvoux B, Paulussen AD, Janssen RG, Kieback DG (2006) Two functionally relevant polymorphisms in the human progesterone receptor gene (+331 G/A and progins) and the predisposition for breast and/or ovarian cancer. Gynecol Oncol 101: 287–295
Rosenberg L, Palmer JR, Zauber AG, Warshauer ME, Lewis Jr JL, Strom BL, Harlap S, Shapiro S (1994) A case–control study of oral contraceptive use and invasive epithelial ovarian cancer. Am J Epidemiol 139: 654–661
Rosenblatt KA, Thomas DB, Noonan EA (1992) High-dose and low-dose combined oral contraceptives: protection against epithelial ovarian cancer and the length of the protective effect. The WHO Collaborative Study of Neoplasia and Steroid Contraceptives. Eur J Cancer 28A: 1872–1876
Rowe SM, Coughlan SJ, McKenna NJ, Garrett E, Kieback DG, Carney DN, Headon DR (1995) Ovarian carcinoma-associated TaqI restriction fragment length polymorphism in intron G of the progesterone receptor gene is due to an Alu sequence insertion. Cancer Res 55: 2743–2745
Royar J, Becher H, Chang-Claude J (2001) Low-dose oral contraceptives: protective effect on ovarian cancer risk. Int J Cancer 95: 370–374
Schildkraut JM, Calingaert B, Marchbanks PA, Moorman PG, Rodriguez GC (2002) Impact of progestin and estrogen potency in oral contraceptives on ovarian cancer risk. J Natl Cancer Inst 94: 32–38
Sellers TA, Schildkraut JM, Pankratz VS, Vierkant RA, Fredericksen ZS, Olson JE, Cunningham J, Taylor W, Liebow M, McPherson C, Hartmann LC, Pal T, Adjei AA (2005) Estrogen bioactivation, genetic polymorphisms, and ovarian cancer. Cancer Epidemiol Biomarkers Prev 14: 2536–2543
Song H, Ramus SJ, Quaye L, DiCioccio RA, Tyrer J, Lomas E, Shadforth D, Hogdall E, Hogdall C, McGuire V, Whittemore AS, Easton DF, Ponder BA, Kjaer SK, Pharoah PD, Gayther SA (2006) Common variants in mismatch repair genes and risk of invasive ovarian cancer. Carcinogenesis 27: 2235–2242
Spurdle AB, Webb PM, Purdie DM, Chen X, Green A, Chenevix-Trench G (2001) No significant association between progesterone receptor exon 4 Val660Leu G/T polymorphism and risk of ovarian cancer. Carcinogenesis 22: 717–721
Study C (1987) The reduction in risk of ovarian cancer associated with oral-contraceptive use. The cancer and steroid hormone study of the centers for disease control and the national institute of child health and human development. N Engl J Med 316: 650–655
Terry KL, De Vivo I, Titus-Ernstoff L, Sluss PM, Cramer DW (2005) Genetic variation in the progesterone receptor gene and ovarian cancer risk. Am J Epidemiol 161: 442–451
Titus-Ernstoff L, Perez K, Cramer DW, Harlow BL, Baron JA, Greenberg ER (2001) Menstrual and reproductive factors in relation to ovarian cancer risk. Br J Cancer 84: 714–721
Tong D, Fabjani G, Heinze G, Obermair A, Leodolter S, Zeillinger R (2001) Analysis of the human progesterone receptor gene polymorphism progins in Austrian ovarian carcinoma patients. Int J Cancer 95: 394–397
Vegeto E, Shahbaz MM, Wen DX, Goldman ME, O'Malley BW, McDonnell DP (1993) Human progesterone receptor A form is a cell- and promoter-specific repressor of human progesterone receptor B function. Mol Endocrinol 7: 1244–1255
Wacholder S, Chanock S, Garcia-Closas M, El Ghormli L, Rothman N (2004) Assessing the probability that a positive report is false: an approach for molecular epidemiology studies. J Natl Cancer Inst 96: 434–442
Whiteman DC, Siskind V, Purdie DM, Green AC (2003) Timing of pregnancy and the risk of epithelial ovarian cancer. Cancer Epidemiol Biomarkers Prev 12: 42–46
Zhou H, Luo MP, Schonthal AH, Pike MC, Stallcup MR, Blumenthal M, Zheng W, Dubeau L (2002) Effect of reproductive hormones on ovarian epithelial tumors: I. Effect on cell cycle activity. Cancer Biol Ther 1: 300–306
Acknowledgements
The Mermaid Group (comprising MALOVA and University College London) additionally thank Ian Jacobs, Lydia Quaye, Claus Hogdall, and Jan Blaeker for their contribution to the study. Finally, we express our profound thanks to all the study participants who contributed to this research.
Genotyping was supported by a grant from the Ovarian Cancer Research Fund provided by the family and friends of Kathryn Sladek Smith (PI: Andrew Berchuck). Support was provided by: US Army Medical Research and Material Command under DAMD17-01-1-0729, the Cancer Council Tasmania, and Cancer Foundation of Western Australia (Australian Ovarian Cancer Study); The National Health and Medical Research Council of Australia (199600) (Australian Cancer Study); US Public Health Service Grants 5R01-CA074850 and 5R01-CA080742 (CONN), the Roswell Park Alliance, and the National Cancer Institute (CA71966 and Core Grant CA16056; Family Registry for Ovarian Cancer Study), Mermaid1 (MALOVA), The Danish Cancer Society (MALOVA), National Cancer Institute, NIH, Bethesda (RO1 CA 61107) (MALOVA), the Fraternal Order of Eagles Cancer Research Fund and the Minnesota Ovarian Cancer Alliance (Mayo Clinic Ovarian Cancer Case–Control Study), Intramural Funds from the National Cancer Institute, NIH, Bethesda, MD (Polish Ovarian Cancer Study); SEARCH is supported by a grant from Cancer Research UK; California Cancer Research Program Grants 00-01389V-20170 and 2110200, US Public Health Service Grants CA14089, CA17054, CA61132, CA63464, N01-PC-67010 and R03-CA113148, and California Department of Health Services subcontract 050-E8709 as part of its statewide cancer reporting programme (University of Southern California). GCT and DW are supported by fellowships from the NHMRC; PW is supported by a fellowship from the Queensland Cancer Fund; HS was funded by a grant from WellBeing of women; PDPP is a Senior Clinical Research Fellow of Cancer Research UK. The full AOCS Study Group is listed on http://www.AOCStudy.org/.
Author information
Authors and Affiliations
Consortia
Corresponding author
Additional information
Supplementary Information accompanies the paper on British Journal of Cancer website (http://www.nature.com/bjc)
Supplementary information
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Pearce, C., Wu, A., Gayther, S. et al. Progesterone receptor variation and risk of ovarian cancer is limited to the invasive endometrioid subtype: results from the ovarian cancer association consortium pooled analysis. Br J Cancer 98, 282–288 (2008). https://doi.org/10.1038/sj.bjc.6604170
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.bjc.6604170
Keywords
This article is cited by
-
Predictors of pretreatment CA125 at ovarian cancer diagnosis: a pooled analysis in the Ovarian Cancer Association Consortium
Cancer Causes & Control (2017)
-
Genetic polymorphisms and gene-dosage effect in ovarian cancer risk and response to paclitaxel/cisplatin chemotherapy
Journal of Experimental & Clinical Cancer Research (2015)
-
Analyze association of the progesterone receptor gene polymorphism PROGINS with ovarian cancer risk
Molecular Biology Reports (2013)
-
Cell cycle genes and ovarian cancer susceptibility: a tagSNP analysis
British Journal of Cancer (2009)
-
Tagging single-nucleotide polymorphisms in candidate oncogenes and susceptibility to ovarian cancer
British Journal of Cancer (2009)