Abstract
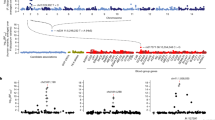
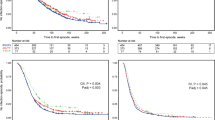
We describe the haplotypic structure of the interferon regulatory factor-1 (IRF-1) locus in two West African ethnic groups, Fulani and Mossi, that differ in their susceptibility and immune response to Plasmodium falciparum malaria. Both populations showed significant associations between IRF-1 polymorphisms and carriage of P. falciparum infection, with different patterns of association that may reflect their different haplotypic architecture. Genetic variation at this locus does not therefore account for the Fulani-specific resistance to malaria while it could contribute to parasite clearance's ability in populations living in endemic areas. We then conducted a case–control study of three haplotype-tagging single nucleotide polymorphisms (htSNPs) in 370 hospitalised malaria patients (160 severe and 210 uncomplicated) and 410 healthy population controls, all from the Mossi ethnic group. All three htSNPs showed correlation with blood infection levels in malaria patients, and the rs10065633 polymorphism was associated with severe disease (P=0.02). These findings provide the first evidence of the involvement in malaria susceptibility of a specific locus within the 5q31 region, previously shown to be linked with P. falciparum infection levels.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 digital issues and online access to articles
$119.00 per year
only $19.83 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Kroger A, Koster M, Schroeder K, Hauser H, Mueller PP . Activities of IRF-1. J Interferon Cytokine Res 2002; 22: 5–14.
Imanishi D, Yamamoto K, Tsushima H, Miyazaki Y, Kuriyama K, Tomonaga M et al. Identification of a novel cytokine response element in the human IFN regulatory factor-1 gene promoter. J Immunol 2000; 165: 3907–3916.
Guo Z, Garg S, Hill KM, Jayashankar L, Mooney MR, Hoelscher M et al. A distal regulatory region is required for constitutive and IFN-beta-induced expression of murine TLR9 gene. J Immunol 2005; 175: 7407–7418.
Foss GS, Prydz H . Interferon regulatory factor 1 mediates the interferon-gamma induction of the human immunoproteasome subunit multicatalytic endopeptidase complex-like 1. J Biol Chem 1999; 274: 35196–35202.
Manzella L, Conte E, Cocchiaro G, Guarniera E, Sciacca B, Bonaiuto C et al. Role of interferon regulatory factor 1 in monocyte/macrophage differentiation. Eur J Immunol 1999; 29: 3009–3016.
Huang Y, Krein PM, Winston BW . Characterization of IFN-gamma regulation of the complement factor B gene in macrophages. Eur J Immunol 2001; 31: 3676–3686.
Yamada G, Ogawa M, Akagi K, Miyamoto H, Nakano N, Itoh S et al. Specific depletion of the B-cell population induced by aberrant expression of human interferon regulatory factor 1 gene in transgenic mice. Proc Natl Acad Sci USA 1991; 88: 532–536.
Elser B, Lohoff M, Kock S, Giaisi M, Kirchhoff S, Krammer PH et al. IFN-gamma represses IL-4 expression via IRF-1 and IRF-2. Immunity 2002; 17: 703–712.
Senaldi G, Shaklee CL, Guo J, Martin L, Boone T, Mak TW et al. Protection against the mortality associated with disease models mediated by TNF and IFN-gamma in mice lacking IFN regulatory factor-1. J Immunol 1999; 163: 6820–6826.
Tan RS, Feng C, Asano Y, Kara AU . Altered immune response of interferon regulatory factor 1-deficient mice against Plasmodium berghei blood-stage malaria infection. Infect Immun 1999; 67: 2277–2283.
Rihet P, Traore Y, Abel L, Aucan C, Traore-Leroux T, Fumoux F . Malaria in humans: Plasmodium falciparum blood infection levels are linked to chromosome 5q31-q33. Am J Hum Genet 1998; 63: 498–505.
Marquet S, Abel L, Hillaire D, Dessein H, Kalil J, Feingold J et al. Genetic localization of a locus controlling the intensity of infection by Schistosoma mansoni on chromosome 5q31-q33. Nat Genet 1996; 14: 181–184.
Modiano D, Petrarca V, Sirima BS, Nebie I, Diallo D, Esposito F et al. Different response to Plasmodium falciparum malaria in West African sympatric ethnic groups. Proc Natl Acad Sci USA 1996; 93: 13206–13211.
Modiano D, Chiucchiuini A, Petrarca V, Sirima BS, Luoni G, Perlmann H et al. Humoral response to Plasmodium falciparum Pf155/ring-infected erythrocyte surface antigen and Pf332 in three sympatric ethnic groups of Burkina Faso. Am J Trop Med Hyg 1998; 58: 220–224.
Modiano D, Chiucchiuini A, Petrarca V, Sirima BS, Luoni G, Roggero MA et al. Interethnic differences in the humoral response to non-repetitive regions of the Plasmodium falciparum circumsporozoite protein. Am J Trop Med Hyg 1999; 61: 663–667.
Dolo A, Modiano D, Maiga B, Daou M, Dolo G, Guindo H et al. Difference in susceptibility to malaria between two sympatric ethnic groups in Mali. Am J Trop Med Hyg 2005; 72: 243–248.
Modiano D, Luoni G, Petrarca V, Sodiomon Sirima B, De Luca M, Simpore J et al. HLA class I in three West African ethnic groups: genetic distances from sub-Saharan and Caucasoid populations. Tissue Antigens 2001; 57: 128–137.
Modiano D, Luoni G, Sirima BS, Lanfrancotti A, Petrarca V, Cruciani F et al. The lower susceptibility to Plasmodium falciparum malaria of Fulani of Burkina Faso (West Africa) is associated with low frequencies of classic malaria-resistance genes. Trans R Soc Trop Med Hyg 2001; 95: 149–152.
Kwiatkowski DP . How malaria has affected the human genome and what human genetics can teach us about malaria. Am J Hum Genet 2005; 77: 171–192.
Mackinnon MJ, Mwangi TW, Snow RW, Marsh K, Williams TN . Heritability of malaria in Africa. PLoS Med 2005; 2: e340.
Timmann C, Evans JA, König IR, Kleensang A, Rüschendorf F, Lenzen J et al. Genome-wide linkage analysis of malaria infection intensity and mild disease. PLoS Genet 2007; 3: e48.
Saito H, Tada S, Ebinuma H, Wakabayashi K, Takagi T, Saito Y et al. Interferon regulatory factor 1 promoter polymorphism and response to type 1 interferon. J Cell Biochem 2001; 81: 191–200.
Saito H, Tada S, Wakabayashi K, Nakamoto N, Takahashi M, Nakamura M et al. The detection of IRF-1 promoter polymorphisms and their possible contribution to T helper 1 response in chronic hepatitis C. J Interferon Cytokine Res 2002; 22: 693–700.
Ball TB, Ji H, Kimani J, McLaren P, Marlin C, Hill AV et al. Polymorphisms in IRF-1 associated with resistance to HIV-1 infection in highly exposed uninfected Kenyan sex workers. AIDS 2007; 21: 1091–1101.
Forton JT, Udalova IA, Campino S, Rockett KA, Hull J, Kwiatkowski DP . Localization of a long-range cis-regulatory element of IL13 by allelic transcript ratio mapping. Genome Res 2007; 17: 82–87.
Modiano D, Luoni G, Sirima BS, Simpore J, Verra F, Konate A et al. Haemoglobin C protects against clinical Plasmodium falciparum malaria. Nature 2001; 414: 305–308.
Jurinke C, van den Boom D, Cantor CR, Koster H . Automated genotyping using the DNA MassArray technology. Methods Mol Biol 2001; 170: 103–116.
Ye S, Dhillon S, Ke X, Collins AR, Day INM . An efficient procedure for genotyping single nucleotide polymorphisms. Nucleic Acids Res 2001; 29: e88-8.
Stephens M, Smith NJ, Donnelly P . A new statistical method for haplotype reconstruction from population data. Am J Hum Genet 2001; 68: 978–989.
Ackerman H, Usen S, Mott R, Richardson A, Sisay-Joof F, Katundu P et al. Haplotypic analysis of the TNF locus by association efficiency and entropy. Genome Biol 2003; 4: R24.
Zhang K, Jin L . HaploBlockFinder: haplotype block analyses. Bioinformatics 2003; 19: 1300–1301.
Nyholt DR . A simple correction for multiple testing for single-nucleotide polymorphisms in linkage disequilibrium with each other. Am J Hum Genet 2004; 74: 765–769.
Acknowledgements
We dedicate this work to the memory of our dear friend and colleague Dr G Luoni, whom we really miss and we would love to keep discussing the data with. We are grateful to the villagers of Barkoumbilen and Barkoundouba, as well as to the hospitalised children and their families, for their very kind collaboration and participation to this study. We are deeply indebted to the personnel of the Immuno-parasitology Unit of the Centre National del Rechèrche et Formation sur le Paludisme of Ouagadougou, Burkina Faso. We thank Dr A Morris for statistical support, Dr Federica Verra for critical review of the manuscript and Professor Marita Troye-Blomberg for useful discussions and supervision of VD Mangano's PhD work. The study was supported by the EU, Sixth Framework Programme, BIOMALPAR Network of Excellence, contract number LSHP-CT-2004-503578, by the Italian Ministry for University and Research (MIUR COFIN, 2003, 2006), by the Istituto Pasteur—Fondazione Cenci Bolognetti of the University of Rome ‘La Sapienza’ and by the UK Medical Research Council.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mangano, V., Luoni, G., Rockett, K. et al. Interferon regulatory factor-1 polymorphisms are associated with the control of Plasmodium falciparum infection. Genes Immun 9, 122–129 (2008). https://doi.org/10.1038/sj.gene.6364456
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.gene.6364456
Keywords
This article is cited by
-
A genome scan for Plasmodium falciparum malaria identifies quantitative trait loci on chromosomes 5q31, 6p21.3, 17p12, and 19p13
Malaria Journal (2014)
-
Genetic polymorphisms linked to susceptibility to malaria
Malaria Journal (2011)
-
Host candidate gene polymorphisms and clearance of drug-resistant Plasmodium falciparum parasites
Malaria Journal (2011)
-
IL12B polymorphisms are linked but not associated with Plasmodium falciparum parasitemia: a familial study in Burkina Faso
Genes & Immunity (2008)