Abstract
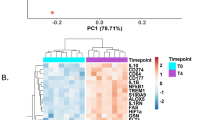
Tularemia is a febrile disease caused by the highly contagious bacterium Francisella tularensis. We undertook an analysis of the transcriptional response in peripheral blood during the course of ulceroglandular tularemia by use of Affymetrix microarrays comprising 14 500 genes. Samples were obtained from seven individuals at five occasions during 2 weeks after the first hospital visit and convalescent samples 3 months later. In total, 265 genes were differentially expressed, 95 of which at more than one time point. The differential expression was verified with real-time quantitative polymerase chain reaction for 36 genes (R2=0.590). The most prominent changes were noted in samples drawn on days 2–3 and a considerable proportion of the upregulated genes appeared to represent an interferon-γ-induced response and also a proapoptotic response. Genes involved in the generation of innate and acquired immune responses were found to be downregulated, presumably a pathogen-induced event. A logistic regression analysis revealed that seven genes were good predictors of the early phase of tularemia. This is the first description of the transcriptional host response to ulceroglandular tularemia and the study has identified gene subsets relevant to the pathogenesis of the disease and subsets that may serve as early diagnostic biomarkers.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 digital issues and online access to articles
$119.00 per year
only $19.83 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Rubins KH, Hensley LE, Jahrling PB, Whitney AR, Geisbert TW, Huggins JW et al. The host response to smallpox: analysis of the gene expression program in peripheral blood cells in a nonhuman primate model. Proc Natl Acad Sci USA 2004; 101: 15190–15195.
Yu SL, Chen HW, Yang PC, Peck K, Tsai MH, Chen JJ et al. Differential gene expression in Gram-negative and Gram-positive sepsis. Am J Respir Crit Care Med 2004; 169: 1135–1143.
Tärnvik A, Berglund L . Tularaemia. Eur Respir J 2003; 21: 361–373.
Dennis DT, Inglesby TV, Henderson DA, Bartlett JG, Ascher MS, Eitzen E et al. Tularemia as a biological weapon: medical and public health management. JAMA 2001; 285: 2763–2773.
Eliasson H, Sjöstedt A, Bäck E . Clinical use of a diagnostic PCR for Francisella tularensis in patients with suspected ulceroglandular tularaemia. Scand J Infect Dis 2005; 37: 833–837.
Syrjala H . Peripheral blood leukocyte counts, erythrocyte sedimentation rate and C-reactive protein in tularemia caused by the type B strain of Francisella tularensis. Infection 1986; 14: 51–54.
Bolstad BM, Irizarry RA, Åstrand M, Speed TP . A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 2003; 19: 185–193.
Lönnstedt IST . Replicated microarray data. Stat Sinica 2002; 12: 31–46.
Kovarova H, Halada P, Man P, Golovliov I, Krocova Z, Spacek J et al. Proteome study of Francisella tularensis live vaccine strain-containing phagosome in Bcg/Nramp1 congenic macrophages: resistant allele contributes to permissive environment and susceptibility to infection. Proteomics 2002; 2: 85–93.
Garin J, Diez R, Kieffer S, Dermine JF, Duclos S, Gagnon E et al. The phagosome proteome: insight into phagosome functions. J Cell Biol 2001; 152: 165–180.
Couillault C, Pujol N, Reboul J, Sabatier L, Guichou JF, Kohara Y et al. TLR-independent control of innate immunity in Caenorhabditis elegans by the TIR domain adaptor protein TIR-1, an ortholog of human SARM. Nat Immunol 2004; 5: 488–494.
Song C, Jin B . TRAIL (CD253), a new member of the TNF superfamily. J Biol Regul Homeost Agents 2005; 19: 73–77.
Ledig S, Wagner S, Manns MP, Beil W, Athmann C . Role of the receptor-mediated apoptosis in Helicobacter pylori in gastric epithelial cells. Digestion 2004; 70: 178–186.
Ghayur T, Banerjee S, Hugunin M, Butler D, Herzog L, Carter A et al. Caspase-1 processes IFN-gamma-inducing factor and regulates LPS-induced IFN-gamma production. Nature 1997; 386: 619–623.
Hersh D, Monack DM, Smith MR, Ghori N, Falkow S, Zychlinsky A . The Salmonella invasin SipB induces macrophage apoptosis by binding to caspase-1. Proc Natl Acad Sci USA 1999; 96: 2396–2401.
Zhang L, Cui R, Cheng X, Du J . Antiapoptotic effect of serum and glucocorticoid-inducible protein kinase is mediated by novel mechanism ac. Cancer Res 2005; 65: 457–464.
Hitoshi Y, Lorens J, Kitada SI, Fisher J, LaBarge M, Ring HZ et al. Toso, a cell surface, specific regulator of Fas-induced apoptosis in T cells. Immunity 1998; 8: 461–471.
Kim CH . The greater chemotactic network for lymphocyte trafficking: chemokines and beyond. Curr Opin Hematol 2005; 12: 298–304.
Seitzer U, Kayser K, Hohn H, Entzian P, Wacker HH, Ploetz S et al. Reduced T-cell receptor CD3zeta-chain protein and sustained CD3epsilon expression at the site of mycobacterial infection. Immunology 2001; 104: 269–277.
Wisniewski HG, Vilcek J . Cytokine-induced gene expression at the crossroads of innate immunity, inflammation and fertility: TSG-6 and PTX3/TSG-14. Cytokine Growth Factor Rev 2004; 15: 129–146.
Hu SP, Harrison C, Xu K, Cornish CJ, Geczy CL . Induction of the chemotactic S100 protein, CP-10, in monocyte/macrophages by lipopolysaccharide. Blood 1996; 87: 3919–3928.
Fitzgerald KA, Rowe DC, Golenbock DT . Endotoxin recognition and signal transduction by the TLR4/MD2-complex. Microbes Infect 2004; 6: 1361–1367.
Ritchie KJ, Hahn CS, Kim KI, Yan M, Rosario D, Li L et al. Role of ISG15 protease UBP43 (USP18) in innate immunity to viral infection. Nat Med 2004; 10: 1374–1378.
Werner ER, Werner-Felmayer G, Fuchs D, Hausen A, Reibnegger G, Yim JJ et al. Tetrahydrobiopterin biosynthetic activities in human macrophages, fibroblasts, THP-1, and T 24 cells. GTP-cyclohydrolase I is stimulated by interferon-gamma, and 6-pyruvoyl tetrahydropterin synthase and sepiapterin reductase are constitutively present. J Biol Chem 1990; 265: 3189–3192.
Gross SS, Levi R, Madera A, Park KH, Vane J, Hattori Y . Tetrahydrobiopterin synthesis is induced by LPS in vascular smooth muscle and is rate-limiting for nitric oxide production. Adv Exp Med Biol 1993; 338: 295–300.
Ihle JN . STATs: signal transducers and activators of transcription. Cell 1996; 84: 331–334.
Rouyez MC, Lestingi M, Charon M, Fichelson S, Buzyn A, Dusanter-Fourt I . IFN regulatory factor-2 cooperates with STAT1 to regulate transporter associated with antigen processing-1 promoter activity. J Immunol 2005; 174: 3948–3958.
Asmis R, Wang Y, Xu L, Kisgati M, Begley JG, Mieyal JJ . A novel thiol oxidation-based mechanism for adriamycin-induced cell injury in human macrophages. FASEB J 2005; 13: 1866–1868.
Lähteenmaki K, Edelman S, Korhonen TK . Bacterial metastasis: the host plasminogen system in bacterial invasion. Trends Microbiol 2005; 13: 79–85.
Nakagawa I, Nakata M, Kawabata S, Hamada S . Transcriptome analysis and gene expression profiles of early apoptosis-related genes in Streptococcus pyogenes-infected epithelial cells. Cell Microbiol 2004; 6: 939–952.
Poquet Y, Kroca M, Halary F, Stenmark S, Peyrat MA, Bonneville M et al. Expansion of Vgamma9 Vdelta2 T cells is triggered by Francisella tularensis-derived phosphoantigens in tularemia but not after tularemia vaccination. Infect Immun 1998; 66: 2107–2114.
Kroca M, Tärnvik A, Sjöstedt A . The proportion of circulating gammadelta T cells increases after the first week of onset of tularaemia and remains elevated for more than a year. Clin Exp Immunol 2000; 120: 280–284.
Zhang K, Zhao H . Assessing reliability of gene clusters from gene expression data. Funct Integr Genomics 2000; 1: 156–173.
Jenner RG, Young RA . Insights into host responses against pathogens from transcriptional profiling. Nat Rev Microbiol 2005; 3: 281–294.
Nagelkerke NJD . A note on a general definition of the coefficient of determination. Biometrika 1991; 78: 691–692.
Andersson H, Hartmanová B, Bäck E, Eliasson H, Landfors M, Rydén P et al. Transcriptional profiling of the peripheral blood response during tularemia. J Med Microbiol 2006; 55: 263–271.
Conlan JW, Chen W, Shen H, Webb A, KuoLee R . Experimental tularemia in mice challenged by aerosol or intradermally with virulent strains of Francisella tularensis: bacteriologic and histopathologic studies. Microb Pathog 2003; 34: 239–248.
Tärnvik A, Sandström G, Löfgren S . Time of lymphocyte response after onset of tularemia and after tularemia vaccination. J Clin Microbiol 1979; 10: 854–860.
Sjöstedt A . Intracellular survival mechanisms of Francisella tularensis, a stealth pathogen. Microbes Infect 2005; 8: 561–567.
Lai XH, Sjöstedt A . Delineation of the molecular mechanisms of Francisella tularensis-induced apoptosis in murine macrophages. Infect Immun 2003; 71: 4642–4646.
Lai XH, Golovliov I, Sjöstedt A . Francisella tularensis induces cytopathogenicity and apoptosis in murine macrophages via a mechanism that requires intracellular bacterial multiplication. Infect Immun 2001; 69: 4691–4694.
Telepnev M, Golovliov I, Grundström T, Tärnvik A, Sjöstedt A . Francisella tularensis inhibits Toll-like receptor-mediated activation of intracellular signalling and secretion of TNF-alpha and IL-1 from murine macrophages. Cell Microbiol 2003; 5: 41–51.
Whitney AR, Diehn M, Popper SJ, Alizadeh AA, Boldrick JC, Relman DA et al. Individuality and variation in gene expression patterns in human blood. Proc Natl Acad Sci USA 2003; 100: 1896–1901.
Thach DC, Lin B, Walter E, Kruzelock R, Rowley RK, Tibbetts C et al. Assessment of two methods for handling blood in collection tubes with RNA stabilizing agent for surveillance of gene expression profiles with high density microarrays. J Immunol Methods 2003; 283: 269–279.
Acknowledgements
This study was supported by grants from the Swedish Medical Research Council, Samverkansnämnden, Västerbottens läns landsting, and the Medical Faculty, Umeå University, Umeå, Sweden, DARPA, Grant LN00A033 from Ministry of Education, Sport and Youth, Czech Republic.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Andersson, H., Hartmanová, B., Bäck, E. et al. Transcriptional profiling of the peripheral blood response during tularemia. Genes Immun 7, 503–513 (2006). https://doi.org/10.1038/sj.gene.6364321
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.gene.6364321
Keywords
This article is cited by
-
Tularemia vaccines
Folia Microbiologica (2016)
-
Prior infection with Type A Francisella tularensis antagonizes the pulmonary transcriptional response to an aerosolized Toll-like receptor 4 agonist
BMC Genomics (2015)
-
Expression profiling of lymph nodes in tuberculosis patients reveal inflammatory milieu at site of infection
Scientific Reports (2015)
-
Host response transcriptional profiling reveals extracellular components and ABC (ATP-binding cassette) transporters gene enrichment in typhoid fever-infected Nigerian children
BMC Infectious Diseases (2011)