Abstract
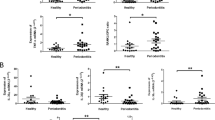
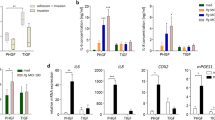
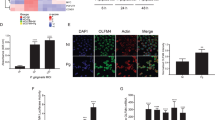
The Toll-like receptor (TLR)4 is the major sensor for bacterial lipopolysaccharide and its two common co-segregating polymorphisms, Asp299Gly and Thr399Ile, which occur at a frequency of between 6 and 10%, have been associated with infectious diseases, LPS hypo-responsiveness and cardiovascular disease. Porphyromonas gingivalis is a Gram-negative bacterium implicated in chronic periodontitis and is a known TLR4 and TLR2 agonist. We obtained two gingival epithelial cell primary cultures from subjects heterozygous for the TLR4 polymorphism Asp299Gly and compared response characteristics with similar cells from patients (four) with the wild-type TLR4 genes. Cytokine responses and transcriptome profiles of gingival epithelial cell primary culture cells to TNFα challenge were similar for all primary epithelial cell cultures. P. gingivalis challenge, however, gave markedly different responses for Asp299Gly heterozygous and wild-type epithelial cell cultures. The epithelial cells heterozygous for the TLR4 polymorphism Asp299Gly were functionally hypo-responsive, evidenced by differences in BD-2 mRNA expression, mRNA response profile by microarray analysis and by pro-inflammatory and chemokine cytokines at the protein and mRNA level. These findings emphasize variance in human epithelial cell TLRs, linked with Asp299Gly carriage, which results in a hypo-responsive epithelial cell phenotype less susceptible to Gram-negative diseases and associated systemic conditions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 digital issues and online access to articles
$119.00 per year
only $19.83 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Takeda K, Akira S . Microbial recognition by Toll-like receptors. J Dermatol Sci 2004; 34: 73–82.
Beutler B . Innate immunity: an overview. Mol Immunol 2004; 40: 845–859.
Akira S, Takeda K, Kaisho T . Toll-like receptors: critical proteins linking innate and acquired immunity. Nat Immunol 2001; 2: 675–680.
Pivarcsi A, Bodai L, Rethi B, Kenderessy-Szabo A, Koreck A, Szell M et al. Expression and function of Toll-like receptors 2 and 4 in human keratinocytes. Int Immunol 2003; 15: 721–730.
Guillot L, Medjane S, Le-Barillec K, Balloy V, Danel C, Chignard M et al. Response of human pulmonary epithelial cells to lipopolysaccharide involves Toll-like receptor 4 (TLR4)-dependent signaling pathways: evidence for an intracellular compartmentalization of TLR4. J Biol Chem 2004; 279: 2712–2718.
Droemann D, Goldmann T, Branscheid D, Clark R, Dalhoff K, Zabel P et al. Toll-like receptor 2 is expressed by alveolar epithelial cells type II and macrophages in the human lung. Histochem Cell Biol 2003; 119: 103–108.
Cario E, Rosenberg IM, Brandwein SL, Beck PL, Reinecker HC, Podolsky DK . Lipopolysaccharide activates distinct signaling pathways in intestinal epithelial cell lines expressing Toll-like receptors. J Immunol 2000; 164: 966–972.
Wolfs TG, Buurman WA, van Schadewijk A, de Vries B, Daemen MA, Hiemstra PS et al. In vivo expression of Toll-like receptor 2 and 4 by renal epithelial cells: IFN-gamma and TNF-alpha mediated up-regulation during inflammation. J Immunol 2002; 168: 1286–1293.
Zhang D, Zhang G, Hayden MS, Greenblatt MB, Bussey C, Flavell RA et al. A toll-like receptor that prevents infection by uropathogenic bacteria. Science 2004; 303: 1522–1526.
Lorenz E, Mira JP, Frees KL, Schwartz DA . Relevance of mutations in the TLR4 receptor in patients with gram-negative septic shock. Arch Intern Med 2002; 162: 1028–1032.
Arbour NC, Lorenz E, Schutte BC, Zabner J, Kline JN, Jones M et al. TLR4 mutations are associated with endotoxin hyporesponsiveness in humans. Nat Genet 2000; 25: 187–191.
Hawn TR, Verbon A, Janer M, Zhao LP, Beutler B, Aderem A . Toll-like receptor 4 polymorphisms are associated with resistance to Legionnaires' disease. Proc Natl Acad Sci USA 2005; 102: 2487–2489.
Erridge C, Stewart J, Poxton IR . Monocytes heterozygous for the Asp299Gly and Thr399Ile mutations in the Toll-like receptor 4 gene show no deficit in lipopolysaccharide signalling. J Exp Med 2003; 197: 1787–1791.
Zhou Q, Desta T, Fenton M, Graves DT, Amar S . Cytokine profiling of macrophages exposed to Porphyromonas gingivalis, its lipopolysaccharide, or its FimA protein. Infect Immun 2005; 73: 935–943.
Chung WO, Dale BA . Innate immune response of oral and foreskin keratinocytes: utilization of different signaling pathways by various bacterial species. Infect Immun 2004; 72: 352–358.
Noguchi T, Shiba H, Komatsuzawa H, Mizuno N, Uchida Y, Ouhara K et al. Syntheses of prostaglandin E2 and E-cadherin and gene expression of beta-defensin-2 by human gingival epithelial cells in response to Actinobacillus actinomycetemcomitans. Inflammation 2003; 27: 341–349.
Imahara SD, Jelacic S, Junker CE, O'Keefe GE . The TLR4 +896 polymorphism is not associated with lipopolysaccharide hypo-responsiveness in leukocytes. Genes Immun 2005; 6: 37–43.
Triantafilou M, Triantafilou K . Lipopolysaccharide recognition: CD14, TLRs and the LPS-activation cluster. Trends Immunol 2002; 23: 301–304.
Dale SE, Doherty-Kirby A, Lajoie G, Heinrichs DE . Role of siderophore biosynthesis in virulence of Staphylococcus aureus: identification and characterization of genes involved in production of a siderophore. Infect Immun 2004; 72: 29–37.
Schroder NW, Hermann C, Hamann L, Gobel UB, Hartung T, Schumann RR . High frequency of polymorphism Arg753Gln of the Toll-like receptor-2 gene detected by a novel allele-specific PCR. J Mol Med 2003; 81: 368–372.
Folwaczny M, Glas J, Torok HP, Limbersky O, Folwaczny C . Toll-like receptor (TLR) 2 and 4 mutations in periodontal disease. Clin Exp Immunol 2004; 135: 330–335.
Van Rijn BB, Roest M, Franx A, Bruinse HW, Voorbij HA . Single step high-throughput determination of Toll-like receptor 4 polymorphisms. J Immunol Methods 2004; 289: 81–87.
Uchida Y, Shiba H, Komatsuzawa H, Takemoto T, Sakata M, Fujita T et al. Expression of IL-1 beta and IL-8 by human gingival epithelial cells in response to Actinobacillus actinomycetemcomitans. Cytokine 2001; 14: 152–161.
Quackenbush J . Microarray data normalization and transformation. Nat Genet 2002; 32 (Suppl): 496–501.
Yang YH, Dudoit S, Luu P, Lin DM, Peng V, Ngai J et al. Normalization for cDNA microarray data: a robust composite method addressing single and multiple slide systematic variation. Nucleic Acids Res 2002; 30: e15.
Eisen MB, Spellman PT, Brown PO, Botstein D . Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci USA 1998; 95: 14863–14868.
Acknowledgements
Dr David F Lappin was on secondment from The University of Glasgow Dental School, United Kingdom to the School of Dentistry University of Louisville. We recognize support for Dr DF Kinane from grants HRSA-C76HF01199-01, CDC-PA04189 and CDC H75/CCU424835-01.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kinane, D., Shiba, H., Stathopoulou, P. et al. Gingival epithelial cells heterozygous for Toll-like receptor 4 polymorphisms Asp299Gly and Thr399Ile are hypo-responsive to Porphyromonas gingivalis. Genes Immun 7, 190–200 (2006). https://doi.org/10.1038/sj.gene.6364282
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.gene.6364282
Keywords
This article is cited by
-
TLR4 896A/G and TLR9 1174G/A polymorphisms are associated with the risk of infectious mononucleosis
Scientific Reports (2020)
-
Orthodontic cell stress modifies proinflammatory cytokine expression in human PDL cells and induces immunomodulatory effects via TLR-4 signaling in vitro
Clinical Oral Investigations (2020)
-
Clinical update on head and neck cancer: molecular biology and ongoing challenges
Cell Death & Disease (2019)
-
MicroRNA-663 antagonizes apoptosis antagonizing transcription factor to induce apoptosis in epithelial cells
Apoptosis (2019)
-
Periodontal Infectogenomics
Inflammation and Regeneration (2018)