Abstract
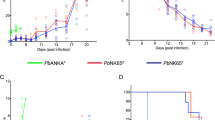
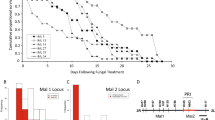
Unprecedented cure after infection with the lethal Plasmodium berghei ANKA was observed in an F2 progeny generated by intercrossing the wild-derived WLA and the laboratory C57BL/6 mouse strains. Resistant mice were able to clear parasitaemia and establish immunity. The observed resistance was disclosed as a combinatorial effect of genetic factors derived from the two parental strains. Genetic mapping of survival time showed that the WLA allele at a locus on chromosome 1 (colocalizing with Berghei resistance 1 (Berr1), a locus associated with resistance to experimental cerebral malaria) increases the probability to resist early death. Also, the C57Bl/6 allele at a novel locus on chromosome 9 (Berr3) confers overall resistance to this lethal Plasmodium infection. This report underlines the value of using wild-derived mouse strains to identify novel genetic factors in the aetiology of disease phenotypes, and provides a unique model for studying parasite clearance and immunity associated with malaria.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 digital issues and online access to articles
$119.00 per year
only $19.83 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Weatherall DJ, Clegg JB . Genetic variability in response to infection: malaria and after. Genes Immun 2002; 3: 331–337.
Weatherall DJ . Host genetics and infectious disease. Parasitology 1996; 112 (Suppl): S23–S29.
Kwiatkowski D . Genetic susceptibility to malaria getting complex. Curr Opin Genet Dev 2000; 10: 320–324.
Fortin A, Belouchi A, Fong Tam M . Genetic control of blood parasitaemia in mouse malaria maps to chromosome 8. Nat Genet 1997; 17: 382–383.
Foote SJ et al. Mouse loci for malaria-induced mortality and the control of parasitaemia. Nat Genet; 1997; 17: 380–381.
Burt RA, Baldwin TM, Marshall VM, Foote SJ . Temporal expression of an H2-linked locus in host response to mouse malaria. Immunogenetics 1999; 50: 278–285.
Fortin A, Cardon LR, Tam M, Skamene E, Stevenson MM, Gros P . Identification of a new malaria susceptibility locus (Char4) in recombinant congenic strains of mice. Proc Natl Acad Sci USA 2001; 98: 10793–19798.
Hernandez-Valladares M, Naessens J, Gibson JP . Confirmation and dissection of QTL controlling resistance to malaria in mice. Mamm Genome 2004; 15: 390–398.
Hernandez-Valladares M, Rihet P, ole-MoiYoi OK, Iraqi FA . Mapping of a new quantitative trait locus for resistance to malaria in mice by a comparative mapping approach with human chromosome 5q31–q33. Immunogenetics 2004; 56: 115–117.
Ohno T, Ishih A, Kohara Y, Yonekawa H, Terada M, Nishimura M . Chromosomal mapping of the host resistance locus to rodent malaria (Plasmodium yoelii) infection in mice. Immunogenetics 2001; 53: 736–740.
Nagayasu E, Nagakura K, Akaki M . Association of a determinant on mouse chromosome 18 with experimental severe Plasmodium berghei malaria. Infect Immun 2002; 70: 512–516.
Bagot S, Campino S, Penha-Goncalves C, Pied S, Cazenave PA, Holmberg D . Identification of two cerebral malaria resistance loci using an inbred wild-derived mouse strain. Proc Natl Acad Sci USA 2002; 99: 9919–9923.
Lander E, Kruglyak L . Genetic dissection of complex traits: guidelines for interpreting and reporting linkage results. Nat Genet 1995; 11: 241–247.
Broman KW, Wu H, Sen S, Churchill GA . R/qtl: QTL mapping in experimental crosses. Bioinformatics 2003; 19: 889–890.
Broman KW . Mapping quantitative trait loci in the case of a spike in the phenotype distribution. Genetics 2003; 163: 1169–1175.
Campino S, Behrschmidt C, Bagot S . Unique genetic variation revealed by a microsatellite polymorphism survey in ten wild-derived inbred strains. Genomics 2002; 79: 618–620.
Guenet JL . Wild mice as a source of genetic polymorphism. Pathol Biol (Paris) 1998; 46: 685–688.
Guenet JL, Bonhomme F . Wild mice: an ever-increasing contribution to a popular mammalian model. Trends Genet 2003; 19: 24–31.
Bagot S, Idrissa Boubou M, Campino S . Susceptibility to experimental cerebral malaria induced by Plasmodium berghei ANKA in inbred mouse strains recently derived from wild stock. Infect Immun 2002; 70: 2049–2056.
Belnoue E, Kayibanda M, Deschemin JC . CCR5 deficiency decreases susceptibility to experimental cerebral malaria. Blood 2003; 101: 4253–4259.
Adachi K, Tsutsui H, Kashiwamura SI . Plasmodium berghei infection in mice induces liver injury by an IL-12- and toll-like receptor/myeloid differentiation factor 88-dependent mechanism. J Immunol 2001; 167: 5928–5934.
Janse CJ, Van Vianen PH . Flow cytometry in malaria detection. Methods Cell Biol 1994; 42: 295–318.
Acknowledgements
We are very grateful to Isabelle Lanctin and Danièle Voegtlé for technical assistance. We are most indebted to António Coutinho, Maria Mota, Sarah Auburn and Taane Clark for continuous support during the development of this work and for critical reading of the manuscript. This work was supported by the Fundação para a Ciência e Tecnologia, Portugal (36392/99), and by the CNRS LEA ‘Génétique et dévelopement de la tolérance naturelle’. It was also supported by the Fundação Calouste Gulbenkian, ICCTI, French Embassy in Lisbon, by the ‘Programme de recherche fondamentale en microbiologie, maladies infectieuses et parasitaires’ of the French Ministry of Research, by INSERM U511 and by the Swedish Research Council.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Campino, S., Bagot, S., Bergman, ML. et al. Genetic control of parasite clearance leads to resistance to Plasmodium berghei ANKA infection and confers immunity. Genes Immun 6, 416–421 (2005). https://doi.org/10.1038/sj.gene.6364219
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.gene.6364219
Keywords
This article is cited by
-
Expression of CD300lf by microglia contributes to resistance to cerebral malaria by impeding the neuroinflammation
Genes & Immunity (2020)
-
Genetic analysis of cerebral malaria in the mouse model infected with Plasmodium berghei
Mammalian Genome (2018)
-
Identification of the Plasmodium berghei resistance locus 9 linked to survival on chromosome 9
Malaria Journal (2013)
-
Susceptibility to lethal cerebral malaria is regulated by epistatic interaction between chromosome 4 (Berr6) and chromosome 1 (Berr7) loci in mice
Genes & Immunity (2013)
-
Host resistance to malaria: using mouse models to explore the host response
Mammalian Genome (2011)