Abstract
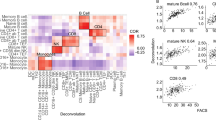
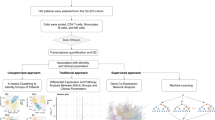
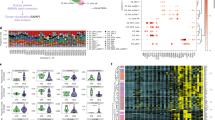
We carried out gene expression profiling of peripheral blood mononuclear cells (PBMCs) in 29 patients with active rheumatoid arthritis (RA) and 21 control subjects using Affymetrix U95Av2 arrays. Using cluster analysis, we observed a significant alteration in the expression pattern of 81 genes (P<0.001) in the PBMCs of RA patients compared with controls. Many of these genes correlated with differences in monocyte counts between the two study populations, and we show that a large fraction of these genes are specifically expressed at high levels in monocytes. In addition, a logistic regression analysis was performed to identify genes that performed best in the categorization of RA and control samples. Glutaminyl cyclase, IL1RA, S100A12 (also known as calgranulin or EN-RAGE) and Grb2-associated binding protein (GAB2) were among the top discriminators. Along with previous data, the overexpression of S100A12 in RA patients emphasizes the likely importance of RAGE pathways in disease pathogenesis. The altered expression of GAB2, an intracellular adaptor molecule involved in regulating phosphatase function, is of particular interest given the recent identification of the intracellular phosphatase PTPN22 as a risk gene for RA. These data suggest that a detailed study of gene expression patterns in peripheral blood can provide insight into disease pathogenesis. However, it is also clear that substantially larger sample sizes will be required in order to evaluate fully gene expression profiling as a means of identifying disease subsets, or defining biomarkers of outcome and response to therapy in RA.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 digital issues and online access to articles
$119.00 per year
only $19.83 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Gregersen PK . Teasing apart the complex genetics of human autoimmunity: lessons from rheumatoid arthritis. Clin Immunol 2003; 107: 1–9.
Gulko PS, Winchester RJ . Rheumatoid arthritis. In: Frank Austen K, Frank MM, Atkinson JP, Cantor H (eds). Samter's Immunologic Diseases. Lippincot Williams and Wilkins: Philadelphia, PA, 1995, pp 427–463.
Weinblatt ME, Kremer JM, Bankhurst AD et al. A trial of etanercept, a recombinant tumor necrosis factor receptor: Fc fusion protein, in patients with rheumatoid arthritis receiving methotrexate. N Engl J Med 1999; 340: 253–259.
Lipsky PE, van der Heijde DM, St Clair EW et al. Infliximab and methotrexate in the treatment of rheumatoid arthritis. Anti-Tumor Necrosis Factor Trial in Rheumatoid Arthritis with Concomitant Therapy Study Group. N Engl J Med 2000; 343: 1594–1602.
Staudt LM . Gene expression profiling of lymphoid malignancies. Annu Rev Med 2002; 53: 303–318.
Baechler EC, Batliwalla FM, Karypis G et al. Interferon-inducible gene expression signature in peripheral blood cells of patients with severe lupus. Proc Natl Acad Sci USA 2003; 100: 2610–2615.
Finnin M, Hamilton JA, Moss ST . Characterization of a CSF-induced proliferating subpopulation of human peripheral blood monocytes by surface marker expression and cytokine production. J Leukoc Biol 1999; 66: 953–960.
Lohn M, Mueller C, Langner J . Cell cycle retardation in monocytoid cells induced by aminopeptidase N (CD13). Leuk Lymphoma 2002; 43: 407–413.
Jawaheer D, Seldin MF, Amos CI et al. A genomewide screen in multiplex rheumatoid arthritis families suggests genetic overlap with other autoimmune diseases. Am J Hum Genet 2001; 68: 927–936.
Jawaheer D, Seldin MF, Amos CI, et al, North American Rheumatoid Arthritis Consortium. Screening the genome for rheumatoid arthritis susceptibility genes: a replication study and combined analysis of 512 multicase families. Arthritis Rheum 2003; 48: 906–916.
Harris ED . Clinical features of rheumatoid arthritis. In: Kelley WN, Ruddy S, Harris Jr ED, Sledge CB (eds). Textbook of Rheumatology, 5th edn. WB Saunders Company: Philadelphia, 1997, pp 898–932.
Fuchs HA, Sergent JS . Rheumatoid arthritis: the clinical picture. In: Koopman WJ (ed). Arthritis and Allied Conditions, 13th edn. Williams and Wilkins: Baltimore, 1997, pp 1041–1070.
Bovin LF, Rieneck K, Workman C et al. Blood cell gene expression profiling in rheumatoid arthritis. Discriminative genes and effect of rheumatoid factor. Immunol Lett 2004; 93: 217–226.
Kawanaka N, Yamamura M, Aita T et al. CD14+, CD16+ blood monocytes and joint inflammation in rheumatoid arthritis. Arthritis Rheum 2002; 46: 2578–2586.
Cairns AP, Crockard AD, Bell AL . The CD14+ CD16+ monocyte subset in rheumatoid arthritis and systemic lupus erythematosus. Rheumatol Int 2002; 21: 189–192.
Geissmann F, Jung S, Littman DR . Blood monocytes consist of two principal subsets with distinct migratory properties. Immunity 2003; 19: 71–82.
Abrahams VM, Cambridge G, Lydyard PM, Edwards JC . Induction of tumor necrosis factor alpha production by adhered human monocytes: a key role for Fcgamma receptor type IIIa in rheumatoid arthritis. Arthritis Rheum 2000; 43: 608–616.
Stuhlmuller B, Ungethum U, Scholze S et al. Identification of known and novel genes in activated monocytes from patients with rheumatoid arthritis. Arthritis Rheum 2000; 43: 775–790.
Hirohata S, Yanagida T, Hashimoto H, Tomita T, Ochi T . Suppressive influences of methotrexate on the generation of CD14(+) monocyte-lineage cells from bone marrow of patients with rheumatoid arthritis. Clin Immunol 1999; 91: 84–89.
van der Pouw Kraan TC, van Gaalen FA, Huizinga TW, Pieterman E, Breedveld FC, Verweij CL . Discovery of distinctive gene expression profiles in rheumatoid synovium using cDNA microarray technology: evidence for the existence of multiple pathways of tissue destruction and repair. Genes Immun 2003; 4: 187–196.
Bennett L, Palucka AK, Arce E et al. Interferon and granulopoiesis signatures in systemic lupus erythematosus blood. J Exp Med 2003; 97: 711–723.
Crow MK, Wohlgemuth J . Microarray analysis of gene expression in lupus. Arthritis Res Ther 2003; 5: 279–287.
Maas K, Chan S, Parker J et al. Cutting edge: molecular portrait of human autoimmune disease. J Immunol 2002; 169: 5–9.
Bomprezzi R, Ringner M, Kim S et al. Gene expression profile in multiple sclerosis patients and healthy controls: identifying pathways relevant to disease. Hum Mol Genet 2003; 12: 2191–2199.
Foell D, Roth J . Proinflammatory S100 proteins in arthritis and autoimmune disease. Arthritis Rheum 2004; 50: 3762–3771.
Hofmann MA, Drury S, Hudson BI et al. RAGE and arthritis: the G82S polymorphism amplifies the inflammatory response. Genes Immun 2002; 3: 123–135.
Rouleau P, Vandal K, Ryckman C et al. The calcium-binding protein S100A12 induces neutrophil adhesion, migration, and release from bone marrow in mouse at concentrations similar to those found in human inflammatory arthritis. Clin Immunol 2003; 107: 46–54.
Foell D, Kucharzik T, Kraft M et al. Neutrophil derived human S100A12 (EN-RAGE) is strongly expressed during chronic active inflammatory bowel disease. Gut 2003; 52: 847–853.
Schilling S, Niestroj AJ, Rahfeld JU et al. Identification of human glutaminyl cyclase as a metalloenzyme. Potent inhibition by imidazole derivatives and heterocyclic chelators. J Biol Chem 2003; 278: 49773–49779.
Nishida K, Hirano T . The role of Gab family scaffolding adapter proteins in the signal transduction of cytokine and growth factor receptors. Cancer Sci 2003; 94: 1029–1033.
Nishida K, Wang L, Morii E et al. Requirement of Gab2 for mast cell development and KitL/c-Kit signaling. Blood 2002; 99: 1866–1869.
Arnaud M, Crouin C, Deon C, Loyaux D, Bertoglio J . Phosphorylation of Grb2-associated binder 2 on serine 623 by ERK MAPK regulates its association with the phosphatase SHP-2 and decreases STAT5 activation. J Immunol 2004; 173: 3962–3971.
Begovich AB, Carlton VE, Honigberg LA et al. A missense single-nucleotide polymorphism in a gene encoding a protein tyrosine phosphatase (PTPN22) is associated with rheumatoid arthritis. Am J Hum Genet 2004; 75: 330–337.
Lee AT, Li W, Liew A et al. The PTPN22 R620W polymorphism associates with RF positive rheumatoid arthritis in a dose-dependent manner but not with HLA-SE status. Genes Immun 2005; 6: 129–133.
Baechler EC, Batliwalla FM, Karypis G et al. Expression levels for many genes in human peripheral blood cells are highly sensitive to ex vivo incubation. Genes Immun 2004; 5: 347–353.
Eisen MB, Spellman PT, Brown PO, Botstein D . Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci USA 1998; 95: 14863–14868.
Dyke GV, Patterson HD . Analysis of factorial arrangement when the data are proportional. Biometrics 1952; 8: 1–12.
Eilers PH, Boer JM, van Ommen GJ, van Houwelingen HC . Classification of microarray data with penalized logistic regression. Proc SPIE 2001; 4266: 187–198.
Li W, Yang Y . How many genes are needed for a discriminant microarray data analysis. In: Lin SM, Johnson KF (eds). Methods of Microarray Data Analysis. Kluwer Academic: Boston/Dordrecht/London, 2002, pp 137–149.
van’t Veer LJ, Dai H, van de Vijver MJ et al. Gene expression profiling predicts clinical outcome of breast cancer. Nature 2002; 415: 530–536.
Walton ID, Dietz LJ, Frenzel G et al. Microvolume laser scanning cytometry platform for biological marker discovery. Proc SPIE Int Soc Opt Eng IBOS Soc Photo-Opt Instrum Eng 2000; 3926: 192–201.
Kantor AB, Alters SE, Cheal K, Dietz LJ . Immune systems biology: immunoprofiling of cells and molecules. BioTechniques 2004; 36: 520–524.
Kantor AB, Wang W, Lin H et al. Biomarker discovery by comprehensive phenotyping for 2 autoimmune diseases. Clin Immunol 2004; 111: 186–195.
Mujumdar RB, Ernst LA, Mujumdar SR, Lewis CJ, Waggoner AS . Cyanine dye labeling reagents: sulfoindocyanine succinimidyl esters. Bioconjug Chem 1993; 4: 105–111.
Roederer M, Kantor AB, Parks DR, Herzenberg LA . Cy7PE and Cy7APC: bright new probes for immunofluorescence. Cytometry 1996; 24: 191–197.
Beavis AJ, Pennline KJ . Allo-7: a new fluorescent tandem dye for use in flow cytometry. Cytometry 1996; 24: 390–395.
Acknowledgements
We thank Leslie Goodwin and Sarah Lombardi for assistance with the microarray assays, and Jen Deng, Harini Govindarajan, V Kakkanaiah and Chris Todd for support with SurroScan assays. We also thank Robert Lundsten, Jubal Dais and Ismael Rodriquez for database support by the NSLIJHS Biorepository Informatics Group. We are grateful to Cerdi Beltre for assistance with patient recruitment, and to all the RA patients who agreed to participate in this study. This work was supported by a NIAMS contract NO1-AR-1-2256 (PKG).
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary information accompanies the paper on Genes and Immunity website (http://www.nature.com/gene).
Rights and permissions
About this article
Cite this article
Batliwalla, F., Baechler, E., Xiao, X. et al. Peripheral blood gene expression profiling in rheumatoid arthritis. Genes Immun 6, 388–397 (2005). https://doi.org/10.1038/sj.gene.6364209
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.gene.6364209
Keywords
This article is cited by
-
RA-MAP, molecular immunological landscapes in early rheumatoid arthritis and healthy vaccine recipients
Scientific Data (2022)
-
Genomics, transcriptomics and proteomics to elucidate the pathogenesis of rheumatoid arthritis
Rheumatology International (2017)
-
Gene expression analysis in RA: towards personalized medicine
The Pharmacogenomics Journal (2014)
-
Distinct genetic profile in peripheral blood mononuclear cells of psoriatic arthritis patients treated with methotrexate and TNF-inhibitors
Clinical Rheumatology (2014)
-
Gene expression patterns in peripheral blood cells associated with radiographic severity in African Americans with early rheumatoid arthritis
Rheumatology International (2013)