Abstract
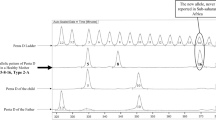
Polymorphism at the TNFd locus has been implicated in a number of disease association studies. The TNFd locus consists of three regions of (GA)n repeats separated by an imperfect repeat of two guanine bases. TNFd alleles are genotyped by the number of repeats in the first (GA)n repeat region, and until now the second repeat region had been thought to be nonpolymorphic. We report the existence of suballeles present within the TNFd microsatellite locus, detected using induced heteroduplex generator (IHG) technology. These alleles cannot be detected using conventional typing strategies as they represent altered distribution of the (GA)n repeats or sequence variation within the repeat. The suballeles affect the frequencies of the conventional d3 and d4 alleles leading to significantly altered allele frequencies. Some studies have associated the d3 and d4 alleles with disease outcome. We re-analysed one such study cohort using IHG technology and demonstrated a high proportion of incorrectly assigned TNFd3 alleles.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 digital issues and online access to articles
$119.00 per year
only $19.83 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
de Baey A, Fellerhoff B, Maier S, Martinozzi S, Weidle U, Weiss EH . Complex expression pattern of the TNF region gene LST1 through differential regulation, initiation, and alternative splicing. Genomics 1997; 45: 591–600.
Rollinger-Holzinger I, Eibl B, Pauly M et al. LST1: a gene with extensive alternative splicing and immunomodulatory function. J Immunol 2000; 15: 169–176.
Holzinger I, de Baey A, Messer G, Kick G, Zwierzina H, Weiss EH . Cloning and genomic characterisation of LST1: a new gene in the human TNF region. Immunogenetics 1995; 42: 315–322.
Turner DM, Grant SCD, Lamb WR et al. A genetic marker of high TNF-α production in heart transplant recipients. Transplantation 1995; 60: 1113–1117.
Hajeer AH, Lear JT, Ollier WE et al. Preliminary evidence of an association of tumour necrosis factor microsatellites with increased risk of multiple basal carcinomas. Br J Dermatol 2000; 142: 441–445.
Kunstmann E, Epplen C, Elitok E et al. Helicobacter pylori infection and polymorphisms in the tumor necrosis factor region. Electrophoresis 1999; 20: 1756–1761.
Mu H, Chen JJ, Jiang Y, King MC, Thomson G, Criswell LA . Tumor necrosis factor a microsatellite polymorphism is associated with rheumatoid arthritis susceptibility through an interaction with the HLA-DRB1 shared epitope. Arthritis Rheum 1999; 42: 438–442.
Honchel R, McDonell S, Schaid DJ, Thibodeau SN . Tumor necrosis factor-α allelic frequency and chromosome 6 allelic imbalance in patients with colorectal cancer. Cancer Res 1996; 56: 145–149.
Greenberg SJ, Fujihara K, Selkirk SM et al. Novel compound tetra-, dinucleotide microsatellite polymorphism in the tumor necrosis factor/lymphotoxin locus. Clin Diagn Lab Immunol 1997; 4: 79–84.
Van der Silk AR, Shing DC, Eerligh P, Giphart MJ . Subtyping for TNFa microsatellite sequence variation. Immunogenetics 2000; 52: 29–34.
Middleton PG, Taylor PR, Jackson G, Proctor SJ, Dickinson AM . Cytokine gene polymorphisms associating with severe acute graft-versus-host disease in HLA-identical sibling transplants. Blood 1998; 92: 3943–3948.
Cavet J, Middleton PG, Segall M, Noreen H, Savies SM, Dickinson AM . Recipient tumor necrosis factor-alpha and interleukin-10 gene polymorphisms associate with early mortality and graft-versus-host disease severity in HLA-matched sibling bone marrow transplants. Blood 1999; 94: 3941–3946.
Udalova IA, Nedospasov SA, Webb GC, Chaplin DD, Turetskaya RL . Highly informative typing of the human TNF locus using six adjacent polymorphic markers. Genomics 1993; 16: 180–186.
Wood N, Bidwell J . Genetic screening and testing by induced heteroduplex formation. Electrophoresis 1996; 17: 247–254.
Morse HR, Olomolaiye OO, Wood NA et al. Induced heteroduplex genotyping of TNF-alpha, IL-1 beta, IL-6 and IL-10 polymorphisms associated with transcriptional regulation. Cytokine 1999; 11: 789–795.
McDonnell GM, Kirk CW, Middleton D et al. Genetic association studies of tumour necrosis factor a and b and tumour necrosis factor receptor 1 and 2 polymorphisms across the clinical spectrum of multiple sclerosis. J Neurol 1999; 246: 1051–1058.
Schneider S, Roessli D, Excoffier L . Arlequin ver 2.000: software for population genetics data analysis. Genetics and Biometry Laboratory, University of Geneva, Switzerland.
Niizeki H, Naruse T, Hecker KH et al. Polymorphisms in the tumour necrosis factor (TNF) genes are associated with susceptibility to effects of ultraviolet-B radiation on induction of contact hypersensitivity. Tissue Antigens 2001; 58: 369–378.
Mulcahy B, Waldron-Lynch F, McDermott MF et al. Genetic variability in the tumor necrosis factor-lymphotoxin region influences susceptibility to rheumatoid arthritis. Am J Hum Genet 1996; 59: 676–683.
Hohler T, Grossmann S, Stradmann-Bellinghausan B et al. Differential association of polymorphisms in the TNFalpha region with psoriatic arthritis but not psoriasis. Ann Rheum Dis 2002; 61: 213–218.
Turner DM, Pravica V, Sinnott PJ, Hutchinson IV . Polymorphism in the promoter region of the gene encoding human allograft inflammatory factor-1. Eur J Immunogenet 2001; 28: 449–450.
Bidwell JL, Keen LJ, Gallagher G et al. Cytokine gene polymorphism in human disease: on-line databases. Genes Immun 1999; 1: 3–19.
Acknowledgements
We would like to thank Dr. Nigel Wood for synthesis of the IHG reagents, and Doris Culpan for her help with the nucleotide sequencing.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Spink, C., Keen, L., Middleton, P. et al. Discrimination of suballeles present at the TNFd microsatellite locus using induced heteroduplex analysis. Genes Immun 5, 76–79 (2004). https://doi.org/10.1038/sj.gene.6364039
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.gene.6364039