Abstract
The variability and mutational changes of the CAG microsatellite in the TATA-box binding protein gene (TBP) were studied. We sequenced the microsatellite of the TBP gene of 25 unrelated individuals from northern Germany (10 SCA17 patients and 15 unaffected control individuals). In addition, the microsatellites were sequenced from individuals of 10 northern German families with at least one family member affected by SCA17. To study also the evolutionary history of this CAG/CAA microsatellite in nonhuman primates, the homologous regions were analysed from Pan troglodytes, Gorilla gorilla, Pongo pygmaeus, P. abellii, Hylobates lar, Nomascus leucogenys, Symphalangus syndactylus, Macaca mulatta, Papio hamadryas, Colobus polykomos and Callithrix jacchus. Three major conclusions were drawn: (i) Patterns of synonymous CAA interruptions in the microsatellite are characteristic and likely to result from selection for stabilizing the repetitive region; (ii) Interspecific comparisons indicate that SCA17 is likely to be a human trait. The most common allele in humans (37 repeats) is close to the threshold value upon which neurodegenerative changes can occur and may act as a repository for expanded, pathogenic alleles; (iii) The cassette-like structure of five out of 17 expanded alleles can be attributed to unequal crossing over. This can explain the rare and sporadic de novo generation of SCA17 alleles.
Similar content being viewed by others
Introduction
Autosomal-dominant cerebellar ataxias are neurodegenerative diseases that strike through progressive degeneration of the cerebellum, brain stem and spinal cord.1 Twenty-eight forms have been identified genetically and six of these disorders are closely associated with an unstable expansion of tandemly repeated CAG (or CTG) trinucleotides that code for polyglutamine tracts in the functional protein of the respective gene.2 This holds also true for the spinocerebellar ataxia type 17 (SCA17) for which an abnormal CAG expansion in the TATA-box binding protein (TBP) gene was first described by Nakamura et al.3 Increased numbers of tandemly repeated CAG repeats are seen as the primary cause for the pathogenesis of dominant ataxias because they correlate to increasingly instable structures and malfunction of the encoded proteins.3, 4 However, the primary biochemical mechanisms of such trinucleotide diseases5 as well as the evolutionary significance of simple repeats and particularly of polyamino acids in proteins is still not completely understood.6
Evolutionary geneticists consider two major processes that can alter the length of microsatellites, in human genetics also referred to as simple tandem repeats (STR):
-
1)
Slippage replication as a result from the mispairing of the matrix and replicated strands during DNA replication,7 and
-
2)
recombinational changes such as unequal crossing over or gene conversion.8
Slippage replication is considered the dominant process for length alterations of STRs and will generate over time highly variable microsatellites. Unfortunately, the evolutionary dynamics of microsatellites is far from being fully understood. The limited knowledge is mirrored in theoretical models developed to simulate allelic variability of microsatellite loci. Starting from simple stepwise mutation models,9 more realistic scenarios for microsatellite evolution including parameters such as, for example, length dependent mutation rates, restriction in length of microsatellites have been put forward.10, 11, 12 Nevertheless, no model can currently satisfactorily predict the distribution of microsatellite alleles in populations. Obviously, allelic variation of microsatellites cannot exclusively be explained by slippage replication.13, 14
Evolutionary genetic studies can contribute to a better understanding of dynamic processes of microsatellites. The comparison of the genetic structures within and between closely related species may be of help to disentangle the mechanisms that determine genetic variability of microsatellites. Rubinsztein et al15 applied such an evolutionary approach to three microsatellites of humans and nonhuman primates that are related to the human polyglutamine diseases SCA1, Machado–Joseph disease and the instability of the androgen receptor gene. In the analysed genes, there was no obvious selection against an increase of microsatellites below a critical length of 38–40 repeats. Other studies of Rubinsztein et al16, 17 suggest that microsatellites can evolve directionally and in closely related species at different rates.
In this study, we analyse the genetic variability of the microsatellite of the TBP gene in human populations and estimate the stability of long, abnormal alleles within families. Furthermore, the variability of microsatellite sequences of the human TBP gene is compared with the homologous genes of 11 nonhuman primate species. Computer simulations and inter- and intraspecific comparisons support that length variation and expansion of the TBP microsatellite cannot simply be explained by slippage replication alone. We found that extremely long microsatellites most likely occurred through sporadic mutation events such as for example, unequal crossing over rather than by stepwise slippage replication processes.
Materials and methods
Samples
Thirty individuals from 10 northern German families with at least one family member affected by SCA17 were genotyped by sequencing the microsatellite region of the TBP gene. Our data complement 10 family analyses that have been documented in the literature.3, 18, 19, 20, 21, 22, 23 The same region of the TBP gene plus 116 bp of the 5′-flanking and a 119 bp long fragment of the 3′-flanking region was also sequenced for another 25 unrelated persons (10 affected and 15 unaffected) from northern Germany. All probands consented in genotyping the TBP locus sequences from GenBank (NM_003194; NT_007583) and fragment length data concerning 1394 from 11 Austrian (Innsbruck), German (Rostock, Hamburg, Berlin, Göttingen, Bochum, Würzburg, Munich, Tübingen, Freiburg) and Swiss (Zürich) cities24 were retrieved for comparison. The microsatellites regions homologous to the human TBP gene were also analysed for Pan troglodytes (N=4), Gorilla gorilla (N=1), Pongo pygmaeus (N=3), P. abellii (N=1), Hylobates lar (N=2), Nomascus leucogenys (N=1), Symphalangus syndactylus (N=1), Macaca mulatta (N=1), Papio hamadryas (N=1), Colobus polykomos (N=1) and Callithrix jacchus (N=1).
DNA isolation, PCR and sequencing of microsatellites
DNA was isolated from blood samples according to standard procedures. The microsatellite and flanking regions of the TBP gene were amplified according to Bauer et al24 using HotStar Taq™ (Qiagen, Hilden, Germany) and the primers SCA17-F, 5′-TTCTCCTTGCTTTCCACAGG -3′ and SCA17-R, 5′-GGGGAGGGATACAGTGGAGT-3′. Purified PCR-products (QIAquick PCR purification kit (Qiagen, Hilden, Germany) were subsequently sequenced using Big Dye (Applied Biosystems, Foster City, CA, USA) or DTCS chemistry (Beckman Coulter, Fullerton, CA, USA). Sequencing was performed on automatic sequencers (ABI 310, Applied Biosystems, Foster City, USA; CEQ8000, Beckman Coulter, Fullerton, CA, USA).
Statistical analyses
Population genetic analyses were based on the dataset of Bauer et al.24 The program GENEPOP25 was used for genetic data analyses. A significance level of 5% was accepted for all tests. The heterozygosity of the microsatellite of the TBP gene was estimated in order to assess the order of magnitude of mutational changes. Measures of genetic variation, however, critically depend upon the parameter θ=4Neμ, where Ne is the effective population size and μ the mutation rate. The parameter θ was estimated from the degree of heterozygosity derived from the infinite allele model26 and the stepwise mutation model.9 Assuming single step mutations, the estimator θF that has been proposed by Xu and Fu27 for microsatellites, was also determined.
Simulations
A computer program (C+ source code) was written in order to simulate stepwise mutations. This deterministic model considers only length variation and neglects point mutations and random effects. The selection scenario was based on clinical observations: In phenotypically healthy persons, the polygutamine stretch is not longer than 42 repeats, that is, amino acids. Incomplete penetrance is observed in persons with a microsatellite region comprising of 43–49 glutamine residues.18, 28 Longer stretches of glutamine will lead to the manifestation of the disease, typically in the third life decade. The model was simplified by defining eleven allele classes: class 1 contains alleles with 22–24 repeats, class 2 contains 25–27 repeats, and so on, and class 11 more than 49 repeats. Alleles with 21–42 repeats are considered selectively neutral. Allele sizes between 43 and 49 repeats show partial dominance h and experience negative selection s, carriers of alleles with more than 49 repeats have also a reduced reproductive success. Finally, zygotes with less than 22 repeats are not viable. Thus, the model restricts variation within a limited range of repeats. The model also considers correlations between allele length and mutation rate as well as mutational length biases, that is, mutational increase of repeat number is favoured. The simulation process transmits new mutations into neighbouring classes. Thus, we simulate mutational events that change microsatellite length through three-repeat-steps neglecting interruptions of microsatellites.
Results
Mutation rate and population size
In Bauer et al's24 collective 22 TBP alleles were detected. The length differences were assigned to different numbers of CAG/CAA repeats. The number of repeats varies between 26 and 51. The allele frequencies show a trimodal distribution with local maxima for 37, 31 and 29 repeats, respectively, and a mean length of 36.1±1.8 repeats. The observed degree of heterozygosity (Hobs=0.785) is close to the expectation (Hexp=0.758). The genotype distributions in the collective revealed no significant differences between the 11 localities (exact test, P=0.604).
Applying the Fisher–Wright model with H=4Neμ/(1+4Neμ), the unbiased estimate of genetic variation θ equals 3.139 with H=0.758. Estimates based on the stepwise mutation model put forward by Ohta and Kimura,9 however, are 2–3 times higher than that given by the infinite allele model: θ̃=8.066. The unbiased estimate of genetic variation θ proposed by Xu and Fu27 was considerably large θF=10.281. Once θ is known, an estimate of the mutation rate depending on the population size is μ=θ/4 Ne, where 0.785⩽θ/4N⩽2.570. Assuming mutation-selection balance, the equilibrium frequency of a harmful allele with partial dominance is q̂≈μ/hs and thus q̂≈a/sNe, where h>0 is the degree of dominance and s the selection coefficient. Using recent demographic parameters of the German population (N≈8 × 107, 720.000 births/year, 1.2 children/woman) and assuming a generation time of 20–25 years, an estimate of the effective population size ranges between 106 and 107. For 0.1<s<0.2 and h=1, the frequency of harmful SCA17 alleles is calculated as between 4 × 10−4 and 10−6 in the German population.
Family analyses
Members of 10 northern German families with at least one family member affected by SCA17 were genotyped at the TBP locus and the resulting data were complemented with literature data on another 10 families. In total, the compiled data set reveals 10 mutations in a total of 47 generational steps within families affected by SCA17. The 10 mutations detected in the family analyses are summarized in Table 1. Three mutations cause drastic increases of the number of repeats within the TBP microsatellite region (families 2, 4 and 5). The characteristic CAA interruptions of the perfect repeats indicate in two instances duplication events that can be identified as de novo paternal expansions (families 2 and 5). Both families were affected for the first time by SCA17. On the other hand, seven of the observed mutations (within three families described by Zühlke et al,18, 23 and one family of this study) can readily be explained by stepwise slippage processes.
Computer simulations
Computer simulations were run in order to identify parameter settings for a stepwise model of slippage replication that may explain a distribution of alleles as observed in 1384 Austrian, German and Swiss probands by Bauer et al.24 In short, no parameter settings were found that generated an allele distribution as the observed one. However, the simulations yielded some interesting results: A mutational bias in favour for long alleles (eg 70% of mutations cause an increase) and no selection on intermediate allele sizes (43–49 repeats) leads to far too high frequencies of intermediate alleles, exceeding 30%. Assuming increasing selection (0.1<s<0.2; q<10−5) decreases the frequency of intermediate alleles and a negative kurtosis of the frequency distribution around allele length of about 35 repeats is found. A positive correlation between mutation rate and repeat length leads to positively skewed distributions of allele frequency with fairly high frequencies of intermediate alleles.
Sequence analyses of three microsatellite region of the TBP gene in primates
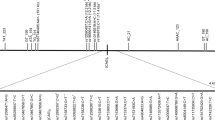
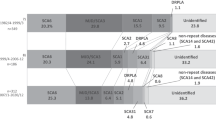
Despite some variation, the organization of the microsatellite region of the TBP gene has a common structure in all analysed primate species (Figures 1 and 2). Typically, the region starts with a (CAG)3(CAA)3 (humans and chimpanzees) or (CAG)3(CAA)2 motif (all other species) and ends with a CAA-CAG hexanucleotide. Deviations from this pattern have so far only been observed in two Italian human individuals affected by SCA17.20 The remaining internal part of the TBP microsatellite can be considered an interrupted microsatellite. Interruptions of perfect tandem repeats have been shown to stabilize the microsatellite.11 Accordingly, we used a typical CAA-CAG-CAA interruption to define a downstream and an upstream section of the internal part of the TBP microsatellite. However, nine alleles were found without the CAA-CAG-CAA interruption that has obviously been lost by recombination; one in a chimpanzee and eight in humans.
The length of the TBP microsatellite, that is, the number of tandemly repeated CAG/CAA motifs, varies substantially between primate species. As can be seen from Figures 1 and 2, the length of the TBP microsatellite increases within primates from new world monkeys to humans roughly by a factor of two. It is noteworthy that only the length of the TBP microsatellite in gorillas, chimpanzees and humans comes close to 40 repeats, the critical length for the development of SCA17. The increase in length occurs first of all in the internal part of the TBP microsatellite downstream of the CAA-CAG-CAA interruption.
Discussion
Comparisons among primate species show that the structure of the microsatellite region of the TBP gene is highly conserved. Few CAA repeats stabilize the CAG microsatellite at particular positions (Figures 1 and 2). As both the CAA and CAG triplets encode for glutamine, the rather conserved position of the CAA interruptions can hardly be explained by selection on the functional TBP. It is more likely that the CAA repeats are located at positions that best stabilize the microsatellite. A comparison of the microsatellites of humans and chimpanzees supports this assumption. The characteristic CAA-CAG-CAA interruption pattern was not altered at least during the last 6 million years. The significance of interruptions for microsatellite stability has also been recognized in studies of ataxia 1 and ataxia 24, 29, 30 and other trinucleotide diseases such as Huntington's disease, Machado–Joseph disease and spinal and bulbar muscular atrophy.31
It has been suggested that a microsatellite region needs a minimum number of at least 4–8 repeats before slippage replication can operate efficiently.12, 14, 32, 33 In most TBP microsatellite sequences from primates except chimpanzees and humans, the CAA interruptions were found at positions of the microsatellite that will prevent long homogeneous CAG tracts. In humans and chimpanzees, however, long homogeneous CAG tracts occur in the internal part of the TBP microsatellite downstream of the CAA-CAG-CAA interruption. Slippage replication and other processes may operate efficiently within these tracts. It has also been proposed that the instability of microsatellites correlates positively with increasing number of repeats.34 Accordingly, one expects more length variation of the TBP microsatellite of chimpanzees and humans than in other primate species. However, the data available do not allow for testing this hypothesis for the TBP microsatellite. If the assumption of a stabilizing effect of CAA repeats on the microsatellite is correct, the loss of these interruptions will have the opposite effect, namely a destabilization of the microsatellite (see eg Choudry et al,4 Imbert et al30). In fact, alleles without the characteristic CAA-CAG-CAA interruption were found (Figure 1), and Zühlke et al18 noticed in two families an increased instability of such alleles. The occurrence of alleles without the characteristic CAA-CAG-CAA interruption further support the assumption of unequal crossing over as an important mechanism for the generation of new TBP alleles because such alleles will result from the same events that duplicate the CAA-CAG-CAA interruptions (five out of 17 expanded alleles, see Table 1 and Figure 1). Nevertheless, the alleles described by Zühlke et al23 and Maltecca et al20 without the characteristic CAA-CAG-CAA interruption cannot be explained just by one unequal crossing over event alone. The length of the microsatellite points to further mutational events that secondarily increased the number of CAG repeats.
Spontaneous occurrences of the disease suggest mutations in the respective microsatellite region that can be traced through family analyses. A compilation of sequences from the literature18, 19, 20, 21, 22, 23 and own data includes seven families with 10 mutational events. Two of these mutations can hardly be explained by slippage replication. The mutations also affected the characteristic CAA-CAG-CAA interruptions (Table 1, families 2 and 5). The cassette-like duplication pattern points to unequal crossing over as the underlying mutational process. In contrast, Choudry et al4 concluded that slippage is the dominant process responsible for length variation of SCA2 alleles because they noticed a lack of cassette-like patterns in their sample (see below). Nevertheless, as unequal crossing over has been identified as an important mechanism for the generation of new TBP alleles in family analyses, it is not surprising that computer simulations with parameter settings according to a stepwise model of slippage replication cannot generate similar allele distributions as were observed in Austrian, German and Swiss populations.
The comparison of the microsatellite region of the TBP gene among primates suggests that first of all humans will be affected by SCA17. Apes show smaller number of repeats (shorter than 21 repeats) whereas most human alleles have sizes reaching up to 40 repeats. Indeed, the most frequent human TBP allele has 37 repeats in the microsatellite region that is only slightly below the critical value for developing the SCA17 phenotype. Only one unequal crossing over event that increases the microsatellite region by just a few repeats will generate TBP alleles with at least incomplete penetrance. In other primate species, however, only unequal crossing over events involving many repeats may generate TBP alleles that cause the SCA17 phenotype. When comparing humans with nonhuman primates, Andrés et al35 suggested long, normal alleles as the repository for expanded, pathogenic SCA8 alleles in humans. Various comparative studies of primates have demonstrated consistently that homologous microsatellite regions have increased in size during human evolution and their liability to diseases is a human-specific trait.17, 31, 35, 36
Andrés et al,35 have also described homogeneous allelic distributions among human populations that rule out ethnicity as a genetic risk for allele expansion. Correspondingly, we found similar allele frequency distributions in different local populations in central Europe. Therefore, the presence of disease alleles in populations rather indicates rare founder events through de novo mutation. If these assumptions are correct, the very low frequency of the SCA17 phenotype may also indicate that unequal crossing over events that considerably expand the number of CAG repeats are rare at the TBP locus. The population genetic analyses provide additional support for this conclusion, for example, assuming a mutation rate between 10−4 and 10−5 and an effective population size of 10−7 the disease has a frequency of about 10−5 in the German population.
References
Harding AE : Clinical features and classification of inherited ataxias. Adv Neurol 1993; 61: 1–14.
Margolis RL : The spinocerebellar ataxias: order emerges from chaos. Curr Neurol Neurosci Rep 2002; 2: 447–456.
Nakamura K, Jeong S-Y, Uchihara T et al: SCA17, a novel autosomal dominant cerebellar ataxia caused by an expanded polyglutamine in TATA-binding protein. Hum Mol Genet 2001; 10: 1441–1448.
Choudry S, Mukerji M, Srivastava AK et al: CAG repeat instability at SCA2 locus: anchoring CAA interruptions and linked single nucleotide polymorphisms. Hum Mol Genet 2001; 10: 2437–2446.
La Spada AR, Richards RI, Wieringa B : Dynamic mutations on the move in Banff. Nat Genet 2004; 36: 667–670.
Djian P : Evolution of simple repeats in DNA and their relation to human disease. Cell 1998; 94: 155–160.
Levinson G, Gutman GA : Slipped-strand mispairing: a major mechanism for DNA sequence evolution. Mol Biol Evol 1987; 4: 203–221.
Garza JC, Freimer M, Freimer NB : Microsatellite allele frequencies in humans and chimpanzees, with implications for constraints on allele size. Mol Biol Evol 1995; 12: 594–603.
Ohta T, Kimura M : A model of mutation appropriate to estimate the number of electrophoretically detectable alleles in finite populations. Genet Res 1973; 22: 201–204.
Zhivotovsky LA, Feldman MW, Grishechkin SA : Biased mutations and microsatellite variation. Mol Biol Evol 1997; 14: 926–933.
Kruglyak S, Durrett RT, Schug MD et al: Equilibrium distributions of microsatellite repeat length resulting from a balance between slippage events and point mutations. Proc Natl Acad Sci USA 1998; 95: 10774–10778.
Sibly RM, Whittaker JC, Talbot M : A maximum-likelihood approach to fitting equilibrium models of microsatellite evolution. Mol Biol Evol 2001; 18: 413–417.
Colson I, Goldstein DB : Evidence for complex mutations at microsatellite loci in Drosophila. Genetics 1999; 152: 617–627.
Bachmann L, Bareiß P, Tomiuk J : Allelic variation, fragment length analyses and population genetic models: a case study of Drosophila microsatellites. J Zool Syst Evol Res 2004; 42: 215–222.
Rubinsztein DC, Leggo J, Coetzee GA et al: Sequence variation and size ranges of CAG repeats in the Machado–Joseph disease, spinocerebellar ataxia type 1 and androgen receptor genes. Hum Mol Genet 1995; 4: 1585–1590.
Rubinsztein DC, Leggo J, Goodburn S et al: Microsatellite evolution - evidence for directionality and variation in rate between species. Nat Genet 1995; 10: 337–343.
Rubinsztein DC, Amos B, Cooper G : Microsatellite and trinucleotide-repeat evolution: evidence for mutational bias and different rates of evolution in different lineages. Phil Trans Royal Soc B 1999; 354: 1095–1099.
Zühlke C, Dalski A, Schwinger E et al: Spinocerebellar ataxia type 17: Report of a family with reduced penetrance of an unstable Gln 49 TBP allele, haplotype analysis supporting a founder effect for unstable alleles and comparative analysis of SCA17 genotypes. BMC Med Genet 2005; 6: 27.
Koide R, Kobayashi S, Shimohata T et al: A neurological disease caused by an expanded CAG trinucleotide repeat in the TATA-binding protein gene: a new polyglutamine disease? Hum Mol Genet 1999; 8: 2047–2053.
Maltecca F, Filla A, Castaldo I et al: Intergenerational instability and marked anticipation in SCA-17. Neurology 2001; 61: 1441–1443.
Rolfs A, Koeppen AH, Bauer I et al: Clinical features and neuropathology of autosomal dominant spinocerebellar ataxia (SCA17). Ann Neurol 2003; 54: 367–375.
Shatunov A, Fridman EA, Pagan FOL et al: Small de novo duplication in the repeat region of the TATA-box-binding protein gene manifest with a phenotype similar to variant Creutzfeldt–Jakob disease. Clin Genet 2004; 66: 496–501.
Zühlke C, Hellenbroich Y, Dalski A et al: Different types of repeat expansion in the TATA-binding protein gene are associated with a new form of inherited ataxia. Eur J Hum Genet 2001; 9: 160–164.
Bauer P, Laccone F, Rolfs A et al: Trinucleotide repeat expansion in SCA17/TBP in white patients with Huntington's disease-like phenotype. J Med Genet 2004; 41: 230–232.
Raymond M, Rousset F : GENEPOP (version 1.2): a population genetics software for exact tests and ecumenism. J Hered 1995; 86: 248–249.
Wright S : Evolution in Mendelian populations. Genetics 1931; 16: 97–159.
Xu H, Fu YX : Estimating effective population size or mutation rate with microsatellites. Genetics 2004; 166: 555–563.
Zühlke C, Spranger M, Spranger S : SCA17 caused by homozygous repeat expansion in TBP due to partial isodisomy 6. Eur J Hum Genet 2003; 11: 629–632.
Chung MY, Ranum LP, Duvick LA et al: Evidence for a mechanism predisposing to intergenerational CAG repeat instability in spinocerebellar ataxia type 2. Nat Genet 1993; 5: 254–258.
Imbert G, Saudou F, Yvert G et al: Cloning of the gene for spinocerebellar ataxia 2 reveals a locus with high sensitivity to expanded CAG/glutamine repeats. Nat Genet 1996; 14: 285–291.
Djian P, Hancock J, Harjinder SC : Codon repeats in genes associated with human diseases: Fewer repeats in the genes of nonhuman primates and nucleotide substitutions concentrated at the sites of reiteration. Proc Natl Acad Sci USA 1996; 93: 417–421.
Lai Y, Sun F : The relationship between microsatellite slippage mutation rate and the number of repeat units. Mol Biol Evol 2003; 20: 2123–2131.
Rose O, Falush D : A threshold size for microsatellite expansion. Mol Biol Evol 1998; 15: 613–615.
Wierdl M, Dominska M, Petes TD : Microsatellite instability in yeast: dependence on the length of the microsatellite. Genetics 1997; 146: 769–779.
Andrés AM, Soldevila M, Saitou N et al: Understanding the dynamics of spinocerebellar ataxia 8 (SCA8) locus through a comparative genetic approach in humans and apes. Neurosci Lett 2003; 336: 143–146.
Justice CM, Den Z, Nguyen SV : Phylogenetic analysis of the Friedreich ataxia GAA trinucleotide repeat. J Mol Evol 2001; 52: 232–238.
Silveira I, Miranda C, Guimaraes L et al: Trinucleotide repeats in 202 families with ataxia: a small expanded (CAG)m allele at the SCA17 locus. Arch Neurol 2002; 59: 623–629.
Stevanin G, Fujigasaki H, Lebre AS et al: Huntington's disease-like phenotype due to trinucleotide repeat expansions in the TBP and JPH3 genes. Brain 2003; 126: 1599–1603.
McNeill SM, Novelletto A, Srinidhi J et al: Reduced penetrance of the Huntington' disease mutation. Hum Mol Genet 1997; 6: 775–779.
Toyoshima Y, Yamada M, Onodera O et al: SCA17 homozygote showing Huntington's disease-like phenotype. Ann Neurol 2004; 55: 281–286.
Zühlke C, Gehlken U, Hellenbroich Y et al: Phenotypical variability of expanded alleles in the TATA-binding protein gene. Reduced penetrance in SCA17? J Neurol 2003; 250: 161–163.
Acknowledgements
JT was supported by a grant from the German Academic Exchange Service DAAD 13/PPP-N1 and by the fortüne-program of the Universitätsklinikum Tübingen, Germany (project nr. 1189-0-0). LB was supported by a grant from the Research Council of Norway (‘National Centre for Biosystematics’, 146515/420). PB and OR were supported by the European Commission Integrated project EUROSCA (LSHM-CT-2004-503304) and by the BMBF ‘Genemove’ project. HZ was supported by the EU project INPRIMAT, contract QLRT-2001-01325. SCA17 genotypes have been made available through a joint project of the German/Austrian/Swiss consortium on HD diagnostics that allowed for the testing of the HD-like 2 and HD-like 4 (SCA17/TBP) loci.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Tomiuk, J., Bachmann, L., Bauer, C. et al. Repeat expansion in spinocerebellar ataxia type 17 alleles of the TATA-box binding protein gene: an evolutionary approach. Eur J Hum Genet 15, 81–87 (2007). https://doi.org/10.1038/sj.ejhg.5201721
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201721
Keywords
This article is cited by
-
Investigation of the Influence of TBP CAG/CAA Repeats in Conjunction with HTT CAG Repeats on Huntington’s Disease Age at Onset in a Brazilian Sample
Journal of Molecular Neuroscience (2022)
-
Instability of expanded CAG/CAA repeats in spinocerebellar ataxia type 17
European Journal of Human Genetics (2008)