Abstract
Cleft lip with or without cleft palate is the most frequent craniofacial malformation in humans (∼1/700). Its etiology is multifactorial; some are a result of a genetic mutation, while others may be due to environmental factors, with genetic predisposition playing an important role. The prevalence varies widely between populations and the mode of inheritance remains controversial. The interferon regulatory factor-6 (IRF6) gene has been shown to harbor mutations in patients with van der Woude syndrome, a dominant form of clefts associated with small pits of the lower lip. Moreover IRF6 has been associated with nonsyndromic cleft of the palate (CL/P) in two separate studies. We investigated the role of IRF6 in a set of 195 trios from Belgium. Cleft occurred as an isolated feature. We studied association of the IRF6 locus using two variants: one in the IRF6 gene and the other 100 kpb 3′ of the gene. Our independent study group confirms that the IRF6 locus is associated with nonsyndromic cleft lip with or without palate. This result, with previous studies performed in the United States and Italy, shows for the first time the implication of IRF6 in isolated CL/P in northern Europe. It is likely that association to this locus can be identified in various populations and that the IRF6 locus thus represents an important genetic modifier for this multifactorial malformation.
Similar content being viewed by others
Introduction
Normal development of lips and palate is a highly regulated and complex process.1 On the basis of murine models, human syndromes, association and expression studies, a wide range of transcription factors, growth factors and signaling molecules have been shown to be involved in this process.2, 3, 4, 5 When the structure or expression of one or more of these genes is disrupted, a cleft of the lip with or without a cleft of the palate (CL/P) may occur. Despite recent investigations on the mode of inheritance and interaction of different factors, genetics of orofacial clefts remains controversial.
Approximately 70% of CL/P cases are nonsyndromic, occurring as an isolated condition without any other associated recognizable anomaly. The remaining 30% are syndromic, associated with other abnormalities.6, 7 The nonsyndromic forms of orofacial clefts are likely due to gene–environment interactions, whereas the syndromic forms are mainly due to genetic alterations.
Inconsistent results have accumulated from genetic studies of complex traits, such as nonsyndromic CL/P, over the past decade. Numerous linkage and association studies performed to search for genes predisposing to CL/P could not be replicated.7 This is not unexpected, given the complicated heterogeneous nature of isolated CL/P and the number of confounding factors. In addition, the resolution of genetic scans has not been sufficient. An interesting approach to study genetics of clefts is to evaluate genes known to contribute to syndromic forms of CL/P and especially to those phenotypically closest to nonsyndromic CL/P. They may serve as a starting point to unravel genetic variants for nonsyndromic CL/P.
The best phenocopy for isolated clefts is probably the van der Woude syndrome (VWS, OMIM# 119300). It is a dominantly inherited disorder in which pits of the lower lip (85% of cases) and hypodontia (25% of cases) are the only features distinguishing it from isolated CL/P.8, 9 Linkage studies pointed to a locus on 1q32–q41.10 Recently, the interferon regulatory factor-6 (IRF6) gene on 1q32.2 was shown to harbor mutations in patients with van der Woude and/or popliteal pterygium syndrome,11 and this has been since reproduced in other studies.12 Moreover, studies investigating the role of IRF6 in the occurrence of isolated clefts showed strong evidence of overtransmission of the V allele of an intragenic IRF6 polymorphism (V274I) in Asian and South American populations.13 Moreover, linkage disequilibrium with CL/P was detected from 40 kbp 5′ of IRF6 to 135 kbp 3′ of IRF6 in the Iowan and Danish populations.13 This result was confirmed by a study supporting the involvement of IRF6 in nonsyndromic CL/P in an Italian population.14 Here, we show that IRF6 gene is likely implicated in nonsyndromic CL/P across different populations as we show study association between the intragenic SNP and CL/P in the Belgian population.
Materials and methods
Family assessment
Families were collected in collaboration with the Cleft Lip and Palate Center, Cliniques universitaires Saint Luc, Brussels, Belgium. Families are from Belgium, which has a heterogeneous population with immigrants from various origins.
Informed consent was obtained from each participant prior to the participation in the study, as approved by the local ethics committee. All patients were ascertained by contributing physicians for the absence of other anomalies and only isolated clefts were included in this study. Patients were asked about familial cleft history. Cleft occurred as a sporadic feature in 190 children, and in five families, the affected child had one affected parent or sibling. As cleft palate is genetically different from cleft lip with or without cleft palate, no cleft palate-only patients were included in this study. Overall, 195 families (190 as trios) with altogether 498 persons were included in this study.
Venous blood samples were drawn for the majority of adult participants. Genomic DNA was extracted from the buffy coats using ‘QIAamp DNA Blood Mini Kit, Qiagen Inc., Valencia, CA, USA’ or from whole blood using ‘DNA purification kit, Gentra systems Inc., Minneapolis, MN, USA’. When whole-blood collection was not possible, DNA was extracted from a buccal-cell sample.
Genotyping
Two single-nucleotide polymorphisms (SNPs) rs2013162 in IRF6 gene and rs2235543 100 kbp 3′ of the gene were genotyped (Table 1, Figure 1). The same SNPs were used in the two previous studies,13, 14 which allows an easy comparison of the results. These SNPs were highly associated with CL/P in the two reports.13, 14 Genotyping was performed using the Taqman chemistry (Applied Biosystems) as described previously.14
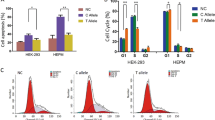
Structure of the IRF6 gene. Exons (rectangles) are drawn to scale, except for exon 9, which is longer than shown. Untranslated regions are smaller rectangles. Exons 3 and 4 encode the DNA-binding domain and exons 7 and 8 code for the protein-binding domain (both in gray). Line connecting exons represents introns and 5′-, 3′-untranslated regions. Arrows indicate the position of the two SNPs in exon 5 of IRF6 and 100 kbp 3′ of the gene, respectively.
Statistical analysis
Hardy–Weinberg equilibrium was assessed in both patients and parents. Transmission disequilibrium test (TDT) was used to measure the distortion of the inheritance of alleles from parents to affected offspring. The null hypothesis was that the transmission of alleles from parents to child was at random.15 Association between the 2 markers and CL/P was tested using Genehunter16 and the Family Based Association Test (FBAT).17, 18, 19 Haplotype transmission disequilibrium was calculated using haplotype-based association test.19 Linkage disequilibrium between the two markers was assessed using the ldmax program from the GOLD software package.20
Results
Hardy–Weinberg equilibrium was assessed for the two SNPs in the Belgian population to ascertain if there was a deviation that would bias our findings. χ2 tests were performed and Hardy–Weinberg equilibrium was observed in both patients and parents.
Results of the TDT showed a significant association of the intragenic rs2013162 SNP with clefting in both Genehunter (P=0.02; results not shown) and FBAT analysis (0.01) (Table 2).
No significant association was obtained when we genotyped the marker 100 kbp 3′ of the gene. Nevertheless, there was a trend towards borderline positive association (P=0.3) (Table 2). This marker lacked a high degree of heterozygosity in our sample (20.2%), and thus most parents were uninformative. For both markers, the common allele (G) was the one that was overtransmitted.
To increase informativeness, we also conducted haplotype-based transmission disequilibrium analysis using genotype data. The haplotype made of the two frequent alleles at each SNP had the most significant distortion (P=0.004), whereas the haplotype made of the two rare alleles showed no association with the condition (Table 3).
Linkage disequilibrium between markers genotyped in this study showed that their association was not random (Δ2=0.014; D′=0.427).
Discussion
The majority of genetic variants underlying susceptibility to isolated clefts are yet to be discovered. Several genes accounting for a sizeable fraction of the syndromic forms of CL/P occurring in a Mendelian pattern have been identified (TBX22, MSX1, P63, FGFR1, TGFBR, etc).21, 22, 23, 24, 25 Mutations in these genes, however, account for less than 15% of the total genetic susceptibility for isolated CL/P and the remaining component is likely to be polygenic in nature – the interaction of mutations in multiple genes, each with a weak effect, in combination with environmental influences.
The identification of all these genes will likely require large case–control association studies. Until the time when complete genome sequence information is feasible to obtain all study subjects, interim strategies involving a subset of all common genetic variations (primarily SNPs) need to be used. Assuming the ‘common disease–common variant’ theory of complex disease susceptibility, a very dense set of genetic markers is likely to identify susceptibility genes, by including either the biologically relevant variant or one closely associated with it.
VWS closely mimics isolated CL/P, with lip pits and dental abnormalities being the only features distinguishing this syndrome from the isolated condition. Moreover, previous studies have implicated IRF6, the VWS causative gene, in isolated cleft lip or palate.13, 14 We used the TDT to see if we can replicate these results in our CL/P cohort collected within the Belgian population. We found strong evidence for association supporting the role of IRF6 in nonsyndromic CL/P in different populations.
In this study, we analyzed the two SNPs in the IRF6 locus that have earlier shown in haplotype analysis the strongest association with CL/P in two different populations. Whether these SNPs are causative themselves or whether they are only in linkage disequilibrium with the disease-causing variants is yet to be elucidated. In our study population, linkage disequilibrium between the two markers was stronger than in the Filipinos, but less significant than in the Italian study. The polymorphism V274I located in exon 7 of IRF6 showed a significant association between the V allele and nonsyndromic CL/P in Asian and South American populations, but not in Europeans.13 We could not identify the rare I allele in 110 tested controls from Europe, nor in a set of European individuals with VWS.12 Based on these two observations, the role of this polymorphism was not investigated in this study, and it is likely not the disease-causing variant.
In this study, we were able to reproduce the results that implicate the IRF6 gene in a complex trait. The 195 trios was clearly a big enough sample to observe the association, which is a necessity for studies of complex traits. As IRF6 shows such a strong association with CL/P, it is interesting to speculate that it may also be a modifier to the phenotypic variability within families with a single IRF6 mutation causing VWS and PPS.
After implicating IRF6 in the complex etiology of isolated CL/P, the question remains: how does IRF6 cause CL/P? Which are the target genes? A better understanding of the function, the targets and the pathways implicating this transcription factor is essential for a better comprehension of the cleft formation as well as for unraveling other CL/P-associated factors.
References
Murray JC, Schutte BC : Cleft palate: players, pathways, and pursuits. J Clin Invest 2004; 113: 1676–1678.
Van den Boogaard MJ, Dorland M, Beemer FA, van Amstel HK : MSX1 mutation is associated with orofacial clefting and tooth agenesis in humans. Nat Genet 2000; 24: 342–343.
FitzPatrick DR, Carr IM, McLaren L et al: Identification of SATB2 as the cleft palate gene on 2q32–q33. Hum Mol Genet 2003; 12: 2491–2501.
Loeys BL, Chen J, Neptune ER et al: A syndrome of altered cardiovascular, craniofacial, neurocognitive and skeletal development caused by mutations in TGFBR1 or TGFBR2. Nat Genet 2005; 37: 275–281.
Murray JC : Gene/environment causes of cleft lip and/or palate. Clin Genet 2002; 61: 248–256.
Jones MC : Etiology of facial clefts: prospective evaluation of 428 patients. Cleft Palate J 1988; 25: 16–20.
Schutte BC, Murray JC : The many faces and factors of orofacial clefts. Hum Mol Genet 1999; 8: 1853–1859.
Van der Woude A : Fistula labii inferioris congenita and its association with cleft lip and palate. Am J Hum Genet 1954; 6: 244–256.
Lacombe D, Pedespan JM, Fontan D : Phenotypic variability in van der Woude syndrome. Genet Counsel 1995; 6: 221–226.
Houdayer C, Soupre V, Rosenberg-Bourgin M et al: Linkage analysis of 5 novel van der Woude kindreds to 1q32–q41 markers further supports locus homogeneity of the disease trait. Ann Genet 1999; 42: 69–74.
Kondo S, Schutte BC, Richardson RJ et al: Mutations in IRF6 cause van der Woude and popliteal pterygium syndromes. Nat Genet 2002; 32: 285–289.
Ghassibe M, Revencu N, Bayet B et al: Six families with van der Woude and/or popliteal pterygium syndrome: all with a mutation in the IRF6 gene. J Med Genet 2004; 41: e15.
Zucchero TM, Cooper ME, Maher BS et al: Interferon regulatory factor 6 (IRF6) gene variants and the risk of isolated cleft lip or palate. N Engl J Med 2004; 351: 769–780.
Scapoli L, Palmieri A, Martinelli M et al: Strong evidence of linkage disequilibrium between polymorphisms at the IRF6 locus and nonsyndromic cleft lip with or without cleft palate, in an Italian population. Am J Hum Genet 2005; 76: 180–183.
Spielman RS, McGinnis RE, Ewens WJ : Transmission test for linkage disequilibrium: the insulin gene region and insulin dependent diabetes mellitus (IDDM). Am J Hum Genet 1993; 52: 506–516.
Daly MJ, Kruglyak L, Pratt S et al: Genehunter Documentation, version 2.0 beta 1998, http://linkage.rockefeller.edu/soft/gh/index.html.
Horvath S, Xu X, Laird NM : The family based association test method: strategies for studying general genotype–phenotype associations. Eur J Hum Genet 2001; 9: 301–306.
Laird NM, Horvath S, Xu X : Implementing a unified approach to family-based tests of association. Genet Epidemiol 2000; 19 (Suppl 1): S36–S42.
Access to FBAT and HaploFBAT at http://www.biostat.harvard.edu/~fbat/default.html.
Abecasis GR, Cookson WO : GOLD – graphical overview of linkage disequilibrium. Bioinformatics 2000; 16: 182–183.
Stanier P, Forbes SA, Arnason A et al: The localization of a gene causing X-linked cleft palate and ankyloglossia (CPX) in an Icelandic kindred is between DXS326 and DXYS1X. Genomics 1993; 17: 549–555.
Van den Boogaard MJ, Dorland M, Beemer FA, van Amstel HK : MSX1 mutation is associated with orofacial clefting and tooth agenesis in humans. Nat Genet 2000; 24: 342–343.
Celli J, Duijf P, Hamel BC et al: Heterozygous germline mutations in the p53 homolog p63 are the cause of EEC syndrome. Cell 1999; 99: 143–153.
Dode C, Levilliers J, Dupont JM et al: Loss-of-function mutations in FGFR1 cause autosomal dominant Kallmann syndrome. Nat Genet 2003; 33: 463–465.
Loeys BL, Chen J, Neptune ER et al: A syndrome of altered cardiovascular, craniofacial, neurocognitive and skeletal development caused by mutations in TGFBR1 or TGFBR2. Nat Genet 2005; 37: 275–281.
Acknowledgements
We thank Jeffrey C Murray and Theresa Zucchero for providing us with SNP materials and for help with study design. We are grateful for all the families for their participation in the study. The studies were supported by the Fonds Spéciaux de Recherche – University of Louvain Medical School, the Fonds national de la recherche scientifique (FNRS) (to MV, a ‘maître de recherche du FNRS’). M Ghassibé was supported by a fellowship from FRIA (Fonds pour la formation à la recherche dans l’industrie et dans l’agriculture).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ghassibé, M., Bayet, B., Revencu, N. et al. Interferon regulatory factor-6: a gene predisposing to isolated cleft lip with or without cleft palate in the Belgian population. Eur J Hum Genet 13, 1239–1242 (2005). https://doi.org/10.1038/sj.ejhg.5201486
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201486
Keywords
This article is cited by
-
Knockdown of Crispld2 in zebrafish identifies a novel network for nonsyndromic cleft lip with or without cleft palate candidate genes
European Journal of Human Genetics (2018)
-
Sequencing of the interferon regulatory factor 6 (IRF6) gene and correlation to its phenotypes in familial non-syndromic cleft lip and palate in North Indian population
European Journal of Plastic Surgery (2014)
-
A combined targeted mutation analysis of IRF6 gene would be useful in the first screening of oral facial clefts
BMC Medical Genetics (2013)
-
Cleft lip and palate: understanding genetic and environmental influences
Nature Reviews Genetics (2011)
-
Evidence of gene–environment interaction for the IRF6 gene and maternal multivitamin supplementation in controlling the risk of cleft lip with/without cleft palate
Human Genetics (2010)