Abstract
Genetic testing of the cystic fibrosis transmembrane conductance (CFTR) gene is currently performed in couples undergoing assisted reproduction techniques (ART), because of the high prevalence of healthy carriers in the population and the pathogenic relationship with congenital bilateral absence of vas deferens (CBAVD). However, discordant data have been reported concerning the usefulness of this genetic test in couples with no family history of cystic fibrosis (CF). In this study, we report the results of CFTR molecular screening in 1195 couples entering ART. Genetic testing was initially carried out in a single partner of each couple. CFTR mutations were detected in 55 subjects (4.6%), a percentage that overlaps with the one reported in the general population. However, significantly higher frequencies of were found in CBAVD individuals (37.5%) and in males with nonobstructive azoospermia (6.6%). The 5T allele was found in 78 patients (6.5%). This figure was again significantly different in males with nonobstructive-azoospermia (9.9%) and in those with CBAVD (100%). All together, 139 subjects (11.6%) had either a CFTR mutation or the 5T allele. Subsequent molecular analysis of their partners disclosed a CFTR mutation or 5T allele in nine cases (6.5%). However, none of these couples had CFTR alterations in both members, a CFTR mutation being invariably present in one partner and the 5T allele in the other. In order to improve genetic counselling of these couples, the TG-M470V-5T association was analyzed, and a statistically significant relationship between 12TG-V470 and CBAVD was detected.
Similar content being viewed by others
Introduction
Cystic fibrosis (CF) is the most common autosomal recessive disease in Caucasians, with a newborn prevalence of about one in 2500 and a carrier frequency of one in 25. This disease is caused by over 900 different mutations in the cystic fibrosis transmembrane regulator (CFTR) gene, located at 7q31.1, 2 Based on the CFTR protein molecular defect, the pathogenic alterations have been divided into five classes.3 Resulting phenotypes are classified as severe or mild CF. The only feature of the least severe form of CF is obstructive azoospermia, due to congenital bilateral absence of vas deferens (CBAVD).4, 5 Approximately 75–80% of patients with CBAVD carry mutations in at least one CF allele. Homozygous CBAVD individuals show one severe and one mild pathogenic mutation or two mild mutations, but never two severe mutations.6, 7, 8, 9, 10 In addition to classic CFTR mutations, up to 40% of CBAVD patients show the 5T allele, that is, five thymidines within intron 8,7, 8, 11, 12 which results in a reduction of the splicing efficiency of the CFTR gene.13 This polymorphism is usually found with a 5–10% prevalence in the general population. Compound heterozygotes for the 5T allele and a CFTR mutation display a variable phenotype, ranging from normal to male infertility or nonclassic CF.14 Recently, it has also been reported that most of CBAVD individuals carrying the 5T allele show in cis an adjacent 12–13 TG repeat and a V470 allele within exon 10,14 suggesting that these polymorphisms do affect the penetrance of the 5T allele.
Since spermatogenesis is normal in approximately 90% of CBAVD men,15 these individuals become good candidates for assisted reproduction techniques (ART), but at risk of transmitting a mutated CFTR allele to their offspring. This possibility has suggested that all CBAVD individuals and their partners should undergo genetic testing of the most common CFTR mutations before ART.16, 17 However, the utility of screening the couples candidates for an ART procedure for problems not related to obstructive azoospermia in the male partner and without a family history of CF is still debated.18
In this study we report the results of a multicentric molecular screening of the most common CFTR gene mutations in 1195 couples undergoing ART, due to different causes of infertility. These findings provide large-scale data on the prevalence of CFTR mutations and 5T allele in these couples, and raise some general issues related to current genetic counselling practices.
Materials and methods
Patients
In total, 1195 consecutive couples underwent genetic counselling for in vitro fertilization from January 2000 to May 2004 in four centers in Italy (one in Chieti, two in Rome and one in Ferrara). All couples were carefully evaluated in order to identify the cause of infertility. A detailed sexual, personal and family history was collected for each couple. All male partners underwent semen analysis according to WHO recommendations.19 In the female partners, the presence of ovulation was determined by measurements of serum progesterone in the mid-luteal phase. Evaluation of female endocrine status also included analysis of FSH, LH, prolactin, androstenedione, testosterone and thyroid hormones. Tubal patency was investigated by hysterosalpingography. When indicated, diagnostic laparoscopy was performed.
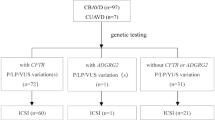
Infertility was due to nonobstructive azoospermia or oligozoospermia in 316 couples (26.4%), obstructive azoospermia (CBAVD) in 16 couples (1.3%), and disorders of the female partner in 93 couples (7.8%). In the remaining 782 couples (65.4%), the cause of infertility was not identified (unexplained infertility). Couples with a family history of CF were not included in this study. All patients gave informed consent to the study, which was approved by local ethic committees.
Screening of CFTR mutations
Analysis of the CFTR gene was carried out on genomic DNA isolated from peripheral blood of one of the two partners in each couple. As a rule, the male partner was selected when CBAVD, nonobstructive azoospermia or oligozoospermia were diagnosed. The female partner was chosen when the infertility of the couple was due to disorders affecting the women. In couples with unexplained infertility, either the male or the female partner was randomly selected. Out of 1195 analyzed partners, 721 were females (60.3%), while 474 were males (39.7%). Molecular screening of CFTR sequence alterations was carried out using a reverse dot blot-based commercial kit, which identifies the 29 most common mutations in Italy and the polyT polymorphism (Inno-Lipa CFTR 12 and 17+Tn, Innogenetics, Ghent, Belgium). When a CFTR mutation or the 5T allele was detected in the first examined partner, the analysis was extended to the second partner of the couple.
TG-M470V-5T combination
In 67 males carrying the 5T allele, the TG repeat adjacent to the polyT sequence and the M470V polymorphism of exon 10 were also analyzed. In total, 13 of these individuals had CBAVD, six nonobstructive azoospermia, 24 oligozoospermia, and 24 normal sperm count. Analysis of the TG repeat was carried out by PCR amplification and direct sequencing of intron 8 of CFTR gene, using the following primers: 5′ GCCAAATATCTTAGTTTTAGA 3′ (forward) and 5′ CAAATAATTCCCCAAATC 3′ (reverse). Detection of the M470V polymorphism was carried out by PCR amplification of exon 10 followed by HpaI restriction endonuclease digestion, according to Cuppens et al.20
Statistical analysis
The frequencies of CFTR mutations and 5T allele were compared in the four subgroups of infertile couples using the χ2 test or Fisher's exact test, as appropriate. The percentage differences between genotype and phenotype subgroups were analyzed in 2 × 2 tables and their statistical significances were calculated by applying Yates’ correction for continuity and Bonferroni's correction for multiple comparisons. The relative risk was estimated using odds ratio (OR) and 95% confidence intervals (95% CI). The OR and 95% CI for the presence of 12TG-V470-5T association in CBAVD patients compared to non-CBAVD patients for the subgroup of infertile couples was calculated. Statistical analysis was performed using SPSS® Advanced Statistical™ 10.0 software (SPSS, Chicago, IL, USA).
Results
CFTR mutations
Analyzed and detected CFTR mutations are summarized in Table 1. Mutational screening of the CFTR gene showed that 55 of the 1195 partners (4.6%) had a CFTR mutation, while 78 subjects (6.5%) had the 5T allele. Six CBAVD individuals (0.5%) were compound heterozygotes for a CFTR mutation and the 5T polymorphism. The phenotype of subjects carrying CFTR mutations is reported in Table 2 (values reported in this table were used for statistical analysis).
An heterozygous CFTR mutation was detected in 20/304 (6.6%) males with nonobstructive infertility, 5/93 (5.4%) infertile females and 30/782 (3.8%) subjects with unexplained infertility. The 5T allele was found in 30 males with nonobstructive infertility (9.9%), 10 subjects (62.5%) with CBAVD, seven infertile females (7.5%) and 31 (4.0%) partners of couples with unexplained infertility. In three males (one with CBAVD and two with normal spermatogenesis) homozygosity for the 5T polymorphism was found. Six CBAVD individuals (37.5%) were compound heterozygotes for a CFTR mutation and the 5T allele. A wild-type genotype was found in 254 (83.6%) partners with nonobstructive infertility, 81 (87.1%) infertile females and 721 (92.2%) partners of couples with unexplained infertility. No CBAVD male resulted to be homozygous for wild-type CF alleles.
Statistical analysis showed significant differences in the prevalence of CFTR mutations between couples in which the male partners had nonobstructive infertility and those with unexplained infertility (χ2=4.72, P<0.05; OR=1.77, 95% CI=0.99–31.6). Similarly, the different distribution of the 5T allele among these two groups was statistically significant (χ2=13.3, P<0.001; OR=2.65, 95% CI=1.58–4.47).
In summary, after the first molecular screening, 139 out of 1195 partners (11.6%) were either carriers of a CFTR mutation or the 5T allele, or compound heterozygotes. Subsequent to a positive finding in one subject, molecular screening of the CFTR gene was also carried out in the respective partner. In nine out of 139 subjects (6.4%), a CFTR alteration was detected, including a mutation in three cases and the 5T polymorphism in the remaining six. The genotypes of these nine couples are summarized in Table 3. No couple in which both partners were carriers of pathogenic CFTR mutations was found.
TG-M470V-5T combination
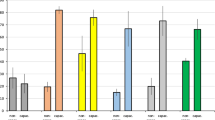
In total, 67 males carrying the 5T allele were analyzed for the TG-M470V-5T combination: 30 (44.8%) had 12 TG and V470 allele, 19 (28.3%) showed 11 TG and M470 allele, 16 (23.9%) showed 11 TG and V470 allele, and two (3.0%) showed 12 TG and M470 allele. The distribution of different TG-M470V associations in relation to the phenotype is summarized in Table 4. Statistical analysis showed that the 12 TG-V470-5T genotype was significantly associated with CBAVD (11/13 males or 84.6%) as compared to non-CBAVD subjects (19/54 or 35.2%) (χ2=8.45, P<0.01; OR=10.13, 95% CI=2.03–50.53).
Discussion
The increasing use of ART in the treatment of infertile couples has raised concerns about the risk of transmitting defective genes underlying the condition of infertility. Molecular screening to detect CFTR mutations is routinely performed before ART, in couples in which the male partner is affected by CBAVD. However, the usefulness of systematic CFTR testing in couples undergoing ART has not been demonstrated. In the present study, we analyzed the CFTR gene in one partner of 1195 infertile couples enrolled into an ART protocol. To our knowledge, this is the largest series reported so far. Using a commercial kit capable of identifying the 29 most common CFTR mutations in Italy and the 5T polymorphism, we found one CF mutation in 55 subjects (4.6%). This result well correlates with the frequency expected in the general Italian population, indicating that the prevalence of CF heterozygotes is similar in the population at large and in infertile couples. However, some differences become quite obvious by comparing the frequency of CFTR mutations in the different subgroups. As expected, a higher frequency of CFTR mutations (37.5%) has been found in the CBAVD individuals, although this percentage is lower when compared to the one reported by others.6, 7, 8, 9, 10 This discrepancy could be due to the presence in Italian CBAVD patients of specific CFTR mutations not included among those investigated, as previously reported in other studies.21 Unexpectedly, we also found a significant difference in the prevalence of CFTR mutations between males with nonobstructive infertility and the male partners of couples with unexplained infertility. An increased prevalence of CFTR mutations in male subjects with nonobstructive infertility has been reported in previous studies,22, 23 but this association has not been invariably confirmed.18, 24, 25 Thus, the relation between CFTR mutations and male infertility requires further assessment.
The prevalence of the 5T polymorphism in the total series of analyzed individuals (6.5%) was not different from the expected (5–10%), but again a significant difference was found within different couples’ subgroups. The 5T allele was invariably found in the 16 CBAVD males: in 6 males, this allele was present in association with a CFTR mutation and in 10 males as the unique alteration. It is likely that this second group of individuals do carry a second CFTR mutation, which was undetectable in our study, because absent in the used diagnostic kit.21, 26 The prevalence of the 5T allele in the present series of CBAVD males was even higher than previously reported, supporting the pathogenic role of this polymorphism for CBAVD. The frequency of the 5T allele was also significantly higher in non-CBAVD infertile males, pointing to a possible relationship between CFTR mutations and nonobstructive male infertility. However, a clear explanation for this relation is still not available. Altogether, we detected a CFTR mutation or the 5T allele in 139 (11.6%) single partners of our couples, in agreement with the figure expected in the general Caucasian population. Molecular analysis of the second partner showed the presence of a CFTR mutation or 5T allele in 6.4% of the cases, a figure that is slightly lower compared to the one detected in the first round of the screening. This result could be explained assuming that the index infertile partners (including CBAVD individuals) who display the highest frequency of CFTR mutations had been selected and analyzed in the first step of the screening. In none of our couples, both partners carried a CFTR mutation, in contrast to the expected prevalence in the general population of one in 625 couples, assuming a carrier prevalence of one in 25. This result is likely to be attributable to two main factors: (1) the exclusion from the study of couples with a positive CF family history, and (2) the 75% mutation detection rate of the tested commercial kit. This latter factor gives rise to a residual risk of 1/100 for analyzed subjects to be carriers of a less common CF mutation. As a consequence, among the 1056 subjects who were negative after the molecular screening, approximately 10 CF carriers might have been missed in this study. However, the aim of the present study was to evaluate the residual risk of couples undergoing ART for generating a CF child after molecular screening of one single partner. Our data indicate that the residual risk of CF in the offspring is not higher in couples undergoing ART, compared to the one expected in the general population. In fact, since a negative test reduces the risk of being a CF carrier to 1/100, the residual risk of generating a CF affected child for a couple in which only one partner had been analyzed, can be estimated in 1/10 000 (risk of the tested partner: 1/100; risk of the not tested partner: 1/25; risk that both partners are CF carrier: 1/2500; risk of an affected child: 1/10 000). Thus, the analysis of the most common CFTR pathogenic mutations in only one partner of the couple may be considered an useful tool in clinical practice, when the use of mutations scanning techniques on both partners would be too expensive or time-consuming.
In the present series, no couple showed a CF mutation in both partners; the only couples at risk of generating an affected child after the molecular screening were those in which both partners were carriers of a CFTR mutation and the 5T allele, respectively. Indeed, these couples are not at risk of generate a child with full-blown CF, but the compound CF-5T genotype can result in variable phenotypes, ranging from normal to CBAVD or mild CF. Therefore, genetic counselling for these couples can be quite a difficult issue. It has been shown that penetrance of the 5T allele is modified by adjacent coexisting TG repeat and M470V polymorphism in exon 10.14, 20, 21, 26, 27, 28 Accordingly, we examined the 67 subjects carrying the 5T allele, in order to assess the relationship between the flanking polymorphisms and clinical outcome. A significant association was established between 12 TG-V470-5T and CBAVD, in agreement with previous observations.14 This result suggests that analysis of TG repeat and M470V polymorphism should be carried out in subjects with the 5T allele undergoing ART, when the other partner is heterozygous for a classic CFTR mutation. This analysis improves genetic counselling and assessment of the risk of mild CF or CBAVD to the offspring.
In conclusion, our data suggest that the prevalence of the most common CFTR mutations in patients entering ART is not increased as compared to the general population, and that the risk for couples of generating a CF child is very low, provided that they have no familiar history for the disease. We suggest that in the common practice it is sufficient to analyze only one partner (preferably the infertile one), and to perform the screening in the second one only when a CFTR mutation or a 5T allele are detected. Finally, the analysis of the 5T-TG-M470V association can provide useful information for calculating the risk of having a child with mild CF or CBAVD in couples in which one partner is a carrier of a CFTR mutation and the second carries the 5T allele.
References
Kerem B, Rommens JM, Buchanan JA et al: Identification of the cystic fibrosis gene: genetic analysis. Science 1989; 245: 1073–1080.
Cystic Fibrosis Genetic Analysis Consortium: Population variation of common cystic fibrosis mutations. Hum Mutat 1994; 4: 167–177.
Estivill X : Complexity in a monogenic disease. Nat Genet 1996; 12: 348–350.
Anguiano A, Oates RD, Amos JA et al: Congenital bilateral absence of the vas deferens. A primarily genital form of cystic fibrosis. JAMA 1992; 267: 1794–1797.
Dumur V, Gervais R, Rigot JM et al: Congenital bilateral absence of the vas deferens (CBAVD) and cystic fibrosis transmembrane regulator (CFTR): correlation between genotype and phenotype. Hum Genet 1996; 97: 7–10.
Casals T, Bassas L, Ruiz-Romero J et al: Extensive analysis of 40 infertile patients with congenital absence of the vas deferens: in 50% of cases only one CFTR allele could be detected. Hum Genet 1995; 95: 205–211.
Chillon M, Casals T, Mercier B et al: Mutations in the cystic fibrosis gene in patients with congenital absence of the vas deferens. N Engl J Med 1995; 332: 1475–1480.
Dork T, Dworniczak B, Aulehla-Scholz C et al: Distinct spectrum of CFTR gene mutations in congenital absence of vas deferens. Hum Genet 1997; 100: 365–377.
Claustres M, Guittard C, Bozon D et al: Spectrum of CFTR mutations in cystic fibrosis and in congenital absence of the vas deferens in France. Hum Mutat 2000; 16: 143–156.
Quinzii C, Castellani C : The cystic fibrosis transmembrane regulator gene and male infertility. J Endocrinol Invest 2000; 23: 684–689.
Jarvi K, McCallum S, Zielenski J et al: Heterogeneity of reproductive tract abnormalities in men with absence of the vas deferens: role of cystic fibrosis transmembrane conductance regulator gene mutations. Fertil Steril 1998; 70: 724–728.
Mak V, Zielenski J, Tsui LC et al: Proportion of cystic fibrosis gene mutations not detected by routine testing in men with obstructive azoospermia. JAMA 1999; 281: 2217–2224.
Chu CS, Trapnell BC, Curristin S, Cutting GR, Crystal RG : Genetic basis of variable exon-9 skipping in cystic fibrosis transmembrane conductance regulator messenger RNA. Nat Genet 1993; 3: 151–156.
Groman JD, Hefferon TW, Casals T et al: Variation in a repeat sequence determines whether a common variant of the cystic fibrosis transmembrane conductance regulator gene is pathogenic or benign. Am J Hum Genet 2004; 74: 176–179.
Meng MV, Black LD, Cha I et al: Impaired spermatogenesis in men with congenital absence of the vas deferens. Hum Reprod 2001; 16: 529–533.
Dequeker E, Cuppens H, Dodge J et al: Recommendations for quality improvement in genetic testing for cystic fibrosis. European concerted action on cystic fibrosis. Eur J Hum Genet 2000; 8 (Suppl 2): S2–S24.
Foresta C, Ferlin A, Gianaroli L, Dallapiccola B : Guidelines for the appropriate use of genetic tests in infertile couples. Eur J Hum Genet 2002; 10: 303–312.
Cruger DG, Agerholm I, Byriel L, Fedder J, Bruun-Petersen G : analysis of males from intracytoplasmic sperm injection couples. Clin Genet 2003; 64: 198–203.
World Health Organization: WHO Laboratory Manual for the Examination of Human Semen and Sperm–Cervical Mucus Interaction, 3rd edn. Cambridge, UK: Cambridge University Press, 1992.
Cuppens H, Lin W, Jaspers M et al: Polyvariant mutant cystic fibrosis transmembrane conductance regulator genes. The polymorphic (Tg)m locus explains the partial penetrance of the T5 polymorphism as a disease mutation. J Clin Invest 1998; 101: 487–496.
Dayangac D, Erdem H, Yilmaz E et al: Mutations of the CFTR gene in Turkish patients with congenital bilateral absence of the vas deferens. Hum Reprod 2004; 19: 1094–1100.
Van der Ven K, Messer L, van der Ven H, Jeyendran RS, Ober C : Cystic fibrosis mutation screening in healthy men with reduced sperm quality. Hum Reprod 1996; 11: 513–517.
Dohle GR, Halley DJ, Van Hemel JO et al: Genetic risk factors in infertile men with severe oligozoospermia and azoospermia. Hum Reprod 2002; 17: 13–16.
Tuerlings JH, Mol B, Kremer JA et al: Mutation frequency of cystic fibrosis transmembrane regulator is not increased in oligozoospermic male candidates for intracytoplasmic sperm injection. Fertil Steril 1998; 69: 899–903.
Pallares-Ruiz N, Carles S, Des Georges M et al: Complete mutational screening of the cystic fibrosis transmembrane conductance regulator gene: cystic fibrosis mutations are not involved in healthy men with reduced sperm quality. Hum Reprod 1999; 14: 3035–3040.
Danziger KL, Black LD, Keiles SB, Kammesheidt A, Turek PJ : Improved detection of cystic fibrosis mutations in infertility patients with DNA sequence analysis. Hum Reprod 2004; 19: 540–546.
de Meeus A, Guittard C, Desgeorges M, Carles S, Demaille J, Claustres M : Linkage disequilibrium between the M470V variant and the IVS8 polyT alleles of the CFTR gene in CBAVD. J Med Genet 1998; 35: 594–596.
Hefferon TW, Groman JD, Yurk CE, Cutting GR : A variable dinucleotide repeat in the CFTR gene contributes to phenotype diversity by forming RNA secondary structures that alter splicing. Proc Natl Acad Sci USA 2004; 101: 3504–3509.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Stuppia, L., Antonucci, I., Binni, F. et al. Screening of mutations in the CFTR gene in 1195 couples entering assisted reproduction technique programs. Eur J Hum Genet 13, 959–964 (2005). https://doi.org/10.1038/sj.ejhg.5201437
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201437
Keywords
This article is cited by
-
Considerations on the use of carrier screening testing in human reproduction: comparison between recommendations from the Italian Society of Human Genetics and other international societies
Journal of Assisted Reproduction and Genetics (2022)
-
Genetic defects in human azoospermia
Basic and Clinical Andrology (2019)
-
Meta-analyses of 4 CFTR variants associated with the risk of the congenital bilateral absence of the vas deferens
Journal of Clinical Bioinformatics (2014)
-
The CFTR polymorphisms poly-T, TG-repeats and M470V in Chinese males with congenital bilateral absence of the vas deferens
Asian Journal of Andrology (2012)
-
Identification of the second CFTR mutation in patients with congenital bilateral absence of vas deferens undergoing ART protocols
Asian Journal of Andrology (2010)