Abstract
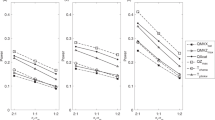
Multipoint interval mapping (MIM) and the MAPMAKER/SIBS program (M/S) are two methods of mapping quantitative loci by examining identity by descent (IBD) sharing in a region spanned by multiple microsatellite DNA markers. For the purpose of comparison, we simulated a quantitative trait controlled by a two-locus model, and evaluated the power and genome-wide false positive rate of both approaches. Based on our simulation, we examined the effects of marker density (5 cM, 10 cM and 20 cM) and sibship size (2, 3, 4 and 5) on the power to detect linkage. Our results indicate that a 10 cM map provides the optimal trade-off between power and type I error, and that the power of MIM increases with sibship size and, in general, performs better than MAPMAKER/SIBS. Furthermore, we conclude that using a reasonable sample of randomly ascertained sibships, it is possible to map a quantitative trait locus (QTL) which accounts for 25% of the phenotypic variance.
Similar content being viewed by others
Article PDF
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Shugart, Y., Goldgar, D. Multipoint genomic scanning for quantitative loci: effects of map density, sibship size and computational approach. Eur J Hum Genet 7, 103–109 (1999). https://doi.org/10.1038/sj.ejhg.5200280
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5200280
Keywords
This article is cited by
-
An evaluation of the variance components approach: type I error, power and size of the estimated effect
European Journal of Human Genetics (2002)