Abstract
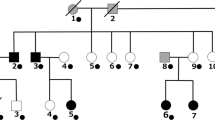
The involvement of the mesocorticolimbic dopamine system in behaviors that are compromised in patients with mood disorder has led to the investigation of dopamine system genes as candidates for bipolar disorder. In particular, the functional VNTRs in the exon III of the dopamine D4 (DRD4) and in intron I of the tyrosine hydroxylase (TH) genes have been investigated in numerous association studies that have produced contrasting results. Likewise, linkage studies in multiplex bipolar families have shown both positive and negative results for markers in close proximity to DRD4 and TH on 11p15.5. We performed a linkage disequilibrium analysis of the DRD4 and TH VNTRs in a sample of 145 nuclear families comprised of DSM-IV bipolar probands and their biological parents. An excess of transmissions and non transmissions was observed for the DRD4 4- and 2-repeat alleles respectively. The biased transmission showed a parent of origin effect (POE) since it was derived almost exclusively from the maternal meiosis (4-repeat allele maternally transmitted 40 times vs 20 times non-transmitted; χ2 = 6.667; df = 1; P = 0.009; while paternally transmitted 26 times vs 21 times non-transmitted; χ2 = 0.531; df = 1; P = 0.46). The analysis of TH did not reveal biased transmission of intron I VNTR alleles. Although replication of our study is necessary, the fact that DRD4 exhibit POE and is located on 11p15.5, in close proximity to a cluster of imprinted genes, suggests that genomic imprinting may be operating in bipolar disorder.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Gershon ES, Martinez M, Goldin LR, Gejman PV . Genetic mapping of common diseases: the challenges of manic-depressive illness and schizophrenia Trends Genet 1990; 6: 282–287
Risch N, Botstein D . A manic depressive history [news] Nat Genet 1996; 12: 351–353
Berrettini WH . On the future of genetic research in bipolar and schizophrenic syndromes Neuropsychopharmacology 1999; 21: 1–2
Egeland JA, Gerhard DS, Pauls DL, Sussex JN, Kidd KK, Allen CR et al. Bipolar affective disorders linked to DNA markers on chromosome 11 Nature 1987; 325: 783–787
Kelsoe JR, Ginns EI, Egeland JA, Gerhard DS, Goldstein AM, Bale SJ et al. Re-evaluation of the linkage relationship between chromosome 11p loci and the gene for bipolar affective disorder in the Old Order Amish Nature 1989; 342: 238–243
Nothen M, Korner J, Lanczik M, Fritze J, Propping P . Tyrosine hydroxylase polymorphisms and manic-depressive illness [letter; comment] Lancet 1990; 336: 575
Detera-Wadleigh SD, Berrettini WH, Goldin LR, Boorman D, Anderson S, Gershon ES . Close linkage of c-Harvey-ras-1 and the insulin gene to affective disorder is ruled out in three North American pedigrees Nature 1987; 325: 806–808
Byerley W, Plaetke R, Hoff M, Jensen S, Holik J, Reimherr F et al. Tyrosine hydroxylase gene not linked to manic-depression in seven of eight pedigrees Hum Hered 1992; 42: 259–263
Sidenberg DG, King N, Kennedy JL . Analysis of new D4 dopamine receptor (DRD4) coding region variants and TH microsatellite in the Old Order Amish family (OOA110) Psychiatr Genet 1994; 4: 95–99
Smyth C, Kalsi G, Brynjolfsson J, O'Neill J, Curtis D, Rifkin L et al. Further tests for linkage of bipolar affective disorder to the tyrosine hydroxylase gene locus on chromosome 11p15 in a new series of multiplex British affective disorder pedigrees [published erratum appears in Am J Psychiatry 1997; 154: 139] Am J Psychiatry 1996; 153: 271–274
Smyth C, Kalsi G, Curtis D, Brynjolfsson J, O'Neill J, Rifkin L et al. Two-locus admixture linkage analysis of bipolar and unipolar affective disorder supports the presence of susceptibility loci on chromosomes 11p15 and 21q22 Genomics 1997; 39: 271–278
Malafosse A, Leboyer M, d'Amato T, Amadeo S, Abbar M, Campion D et al. Manic depressive illness and tyrosine hydroxylase gene: linkage heterogeneity and association Neurobiol Dis 1997; 4: 337–349
De bruyn A, Mendelbaum K, Sandkuijl LA, Delvenne V, Hirsch D, Staner L et al. Nonlinkage of bipolar illness to tyrosine hydroxylase, tyrosinase, and D2 and D4 dopamine receptor genes on chromosome 11 Am J Psychiatry 1994; 151: 102–106
Serretti A, Macciardi F, Cusin C, Lattuada E, Souery D, Lipp O et al. Linkage of mood disorders with D2, D3 and TH genes: a multicenter study J Affect Disord 2000; 58: 51–61
Willner P . Dopaminergic Mechanisms in Depression and Mania Raven Press: New York 1995
Bocchetta A, Piccardi MP, Palmas MA, Chillotti C, Oi A, Del Zompo M . Family-based association study between bipolar disorder and DRD2, DRD4, DAT, and SERT in Sardinia Am J Med Genet 1999; 88: 522–526
Furlong RA, Rubinsztein JS, Ho L, Walsh C, Coleman TA, Muir WJ et al. Analysis and metaanalysis of two polymorphisms within the tyrosine hydroxylase gene in bipolar and unipolar affective disorders Am J Med Genet 1999; 88: 88–94
Li T, Liu X, Sham PC, Aitchison KJ, Cai G, Arranz MJ et al. Association analysis between dopamine receptor genes and bipolar affective disorder Psychiatry Res 1999; 86: 193–201
Lim LC, Nothen MM, Korner J, Rietschel M, Castle D, Hunt N et al. No evidence of association between dopamine D4 receptor variants and bipolar affective disorder Am J Med Genet 1994; 54: 259–263
Oruc L, Verheyen GR, Furac I, Jakovljevic M, Ivezic S, Raeymaekers P et al. Analysis of the tyrosine hydroxylase and dopamine D4 receptor genes in a Croatian sample of bipolar I and unipolar patients Am J Med Genet 1997; 74: 176–178
Perez de Castro I, Torres P, Fernandez-Piqueras J, Saiz-Ruiz J, Llinares C . No association between dopamine D4 receptor polymorphism and manic depressive illness J Med Genet 1994; 31: 897–898
Serretti A, Lilli R, Di Bella D, Bertelli S, Nobile M, Novelli E et al. Dopamine receptor D4 gene is not associated with major psychoses Am J Med Genet 1999; 88: 486–491
Serretti A, Lilli R, Lorenzi C, Lattuada E, Smeraldi E . DRD4 exon 3 variants associated with delusional symptomatology in major psychoses: a study on 2011 affected subjects Am J Med Genet 2001; 105: 283–290
Todd RD, Lobos EA, Parsian A, Simpson S, DePaulo JR . Manic-depressive illness and tyrosine hydroxylase markers. Bipolar Disorder Working Group Lancet 1996; 347: 1634
Weiss J, Magert HJ, Cieslak A, Forssmann WG . Association between different psychotic disorders and the DRD4 polymorphism, but no differences in the main ligand binding region of the DRD4 receptor protein compared to controls Eur J Med Res 1996; 1: 439–445
Asghari V, Sanyal S, Buchwaldt S, Paterson A, Jovanovic V, Van Tol HH . Modulation of intracellular cyclic AMP levels by different human dopamine D4 receptor variants J Neurochem 1995; 65: 1157–1165
Van Tol HH, Wu CM, Guan HC, Ohara K, Bunzow JR, Civelli O et al. Multiple dopamine D4 receptor variants in the human population Nature 1992; 358: 149–152
Lichter JB, Barr CL, Kennedy JL, Van Tol HH, Kidd KK, Livak KJ . A hypervariable segment in the human dopamine receptor D4 (DRD4) gene Hum Mol Genet 1993; 2: 767–773
Jovanovic V, Guan HC, Van Tol HH . Comparative pharmacological and functional analysis of the human dopamine D4.2 and D4.10 receptor variants Pharmacogenetics 1999; 9: 561–568
Polymeropoulos MH, Xiao H, Rath DS, Merril CR . Tetranucleotide repeat polymorphism at the human tyrosine hydroxylase gene (TH) Nucleic Acids Res 1991; 19: 3753
Puers C, Hammond HA, Jin L, Caskey CT, Schumm JW . Identification of repeat sequence heterogeneity at the polymorphic short tandem repeat locus HUMTH01[AATG]n and reassignment of alleles in population analysis by using a locus-specific allelic ladder Am J Hum Genet 1993; 53: 953–958
Meloni R, Albanese V, Ravassard P, Treilhou F, Mallet J . A tetranucleotide polymorphic microsatellite, located in the first intron of the tyrosine hydroxylase gene, acts as a transcription regulatory element in vitro Hum Mol Genet 1998; 7: 423–428
Greenberg DA . Linkage analysis of ‘necessary’ disease loci versus ‘susceptibility’ loci Am J Hum Genet 1993; 52: 135–143
Kidd KK . Associations of disease with genetic markers: deja vu all over again [editorial] Am J Med Genet 1993; 48: 71–73
Chang FM, Kidd JR, Livak KJ, Pakstis AJ, Kidd KK . The world-wide distribution of allele frequencies at the human dopamine D4 receptor locus Hum Genet 1996; 98: 91–101
Ott J . Analysis of Human Genetic Linkage, 3rd edn The John Hopkins University Press: Baltimore and London 1999
Petronis A . The genes for major psychosis: aberrant sequence or regulation? Neuropsychopharmacology 2000; 23: 1–12
First MB, Spitzer RL, Gibbon M, Williams JBW . Structured Clinical Interview for DSM-IV Axis I Disorders, Research Version, Non-patient Edition (SCID-I/NP) Biometrics Research, New York Stat Psychiatric Institute: NY 1997
Edwards A, Hammond HA, Jin L, Caskey Ct, Chakraborty R . Genetic variation at five trimeric and tetrameric tandem repeat loci in four human population groups Genomics 1992; 12: 241–253
Dutton CM, Paynton C, Sommer SS . General method for amplifying regions of very high G+C content Nucleic Acids Res 1993; 21: 2953–2954
Sham PC, Curtis D . An extended transmission/disequilibrium test (TDT) for multi-allele marker loci Ann Hum Genet 1995; 59: 323–336
Falk CT, Rubinstein P . Haplotype relative risks: an easy reliable way to construct a proper control sample for risk calculations Ann Hum Genet 1987; 51: 227–233
Schaid DJ, Sommer SS . Comparison of statistics for candidate-gene association studies using cases and parents Am J Hum Genet 1994; 55: 402–409
Grigoroiu-Serbanescu M, Nothen M, Propping P, Poustka F, Magureanu S, Vasilescu R et al. Clinical evidence for genomic imprinting in bipolar I disorder Acta Psychiatr Scand 1995; 92: 365–370
McMahon FJ, Stine OC, Meyers DA, Simpson SG, DePaulo JR . Patterns of maternal transmission in bipolar affective disorder Am J Hum Genet 1995; 56: 1277–1286
Gershon ES, Badner JA, Detera-Wadleigh SD, Ferraro TN, Berrettini WH . Maternal inheritance and chromosome 18 allele sharing in unilineal bipolar illness pedigrees Am J Med Genet 1996; 67: 202–207
McMahon FJ, Chen YS, Patel S, Kokoszka J, Brown MD, Torroni A et al. Mitochondrial DNA sequence diversity in bipolar affective disorder Am J Psychiatry 2000; 157: 1058–1064
Paterson AD . Sixth World Congress of Psychiatric Genetics X Chromosome Workshop Am J Med Genet 1999; 88: 279–286
Reik W, Walter J . Genomic imprinting: parental influence on the genome Nat Rev Genet 2001; 2: 21–32
Hall JG . Genomic imprinting: review and relevance to human diseases Am J Hum Genet 1990; 46: 857–873
Cassidy SB, Schwartz S . Prader-Willi and Angelman syndromes. Disorders of genomic imprinting Medicine (Baltimore) 1998; 77: 140–151
Isles AR, Wilkinson LS . Imprinted genes, cognition and behaviour Trends Cogn Sci 2000; 4: 309–318
Stine OC, Xu J, Koskela R, McMahon FJ, Gschwend M, Friddle C et al. Evidence for linkage of bipolar disorder to chromosome 18 with a parent-of-origin effect Am J Hum Genet 1995; 57: 1384–1394
McMahon FJ, Hopkins PJ, Xu J, McInnis MG, Shaw S, Cardon L et al. Linkage of bipolar affective disorder to chromosome 18 markers in a new pedigree series Am J Hum Genet 1997; 61: 1397–1404
Van Broeckhoven C, Verheyen G . Chromosome 18 workshop Psychiatr Genet 1998; 8: 97–108
Van Broeckhoven C, Verheyen G . Report of the chromosome 18 workshop Am J Med Genet 1999; 88: 263–270
Zhang Y, Tycko B . Monoallelic expression of the human H19 gene Nat Genet 1992; 1: 40–44
Giannoukakis N, Deal C, Paquette J, Goodyer CG, Polychronakos C . Parental genomic imprinting of the human IGF2 gene Nat Genet 1993; 4: 98–101
Cichon S, Nothen MM, Wolf HK, Propping P . Lack of imprinting of the human dopamine D4 receptor (DRD4) gene Am J Med Genet 1996; 67: 229–231
Barlow DP . Gametic imprinting in mammals Science 1995; 270: 1610–1613
Bunzel R, Blumcke I, Cichon S, Normann S, Schramm J, Propping P et al. Polymorphic imprinting of the serotonin-2A (5-HT2A) receptor gene in human adult brain Brain Res Mol Brain Res 1998; 59: 90–92
McGrath J, Solter D . Completion of mouse embryogenesis requires both the maternal and paternal genomes Cell 1984; 37: 179–183
Surani MA, Barton SC, Norris ML . Development of reconstituted mouse eggs suggests imprinting of the genome during gametogenesis Nature 1984; 308: 548–550
Tilghman SM . The sins of the fathers and mothers: genomic imprinting in mammalian development Cell 1999; 96: 185–193
Bennett ST, Wilson AJ, Cucca F, Nerup J, Pociot F, McKinney PA et al. IDDM2-VNTR-encoded susceptibility to type 1 diabetes: dominant protection and parental transmission of alleles of the insulin gene-linked minisatellite locus J Autoimmun 1996; 9: 415–421
Bennett ST, Wilson AJ, Esposito L, Bouzekri N, Undlien DE, Cucca F et al. Insulin VNTR allele-specific effect in type 1 diabetes depends on identity of untransmitted paternal allele. The IMDIAB Group Nat Genet 1997; 17: 350–352
Gabriel JM, Higgins MJ, Gebuhr TC, Shows TB, Saitoh S, Nicholls RD . A model system to study genomic imprinting of human genes Proc Natl Acad Sci USA 1998; 95: 14857–14862
Kugoh H, Mitsuya K, Meguro M, Shigenami K, Schulz TC, Oshimura M . Mouse A9 cells containing single human chromosomes for analysis of genomic imprinting DNA Res 1999; 6: 165–172
Naumova AK, Greenwood CM, Morgan K . Imprinting and deviation from Mendelian transmission ratios Genome 2001; 44: 311–320
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Muglia, P., Petronis, A., Mundo, E. et al. Dopamine D4 receptor and tyrosine hydroxylase genes in bipolar disorder: evidence for a role of DRD4. Mol Psychiatry 7, 860–866 (2002). https://doi.org/10.1038/sj.mp.4001098
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.mp.4001098
Keywords
This article is cited by
-
Imprinting in the schizophrenia candidate gene GABRB2 encoding GABAA receptor β2 subunit
Molecular Psychiatry (2011)
-
The genetics of bipolar disorder: genome ‘hot regions,’ genes, new potential candidates and future directions
Molecular Psychiatry (2008)
-
Epigenetics in mood disorders
Environmental Health and Preventive Medicine (2008)
-
A linkage and family-based association analysis of a potential neurocognitive endophenotype of bipolar disorder
NeuroMolecular Medicine (2007)
-
Genetic tests of biologic systems in affective disorders
Molecular Psychiatry (2005)