Abstract
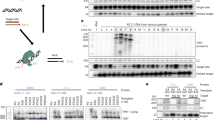
Plasmid vectors have been widely used for DNA vaccines and gene therapy. Following intramuscular injection, the plasmid that persists is extrachromosomal and integration into host DNA, if it occurs at all, is negligible. However, new technologies for improving DNA delivery could increase the frequency of integration. In the present study, we tested the effect of electroporation on plasmid uptake and potential integration following intramuscular injection in mice, using a plasmid containing the mouse erythropoietin gene. Electroporation increased plasmid tissue levels by approximately six- to 34-fold. Using a quantitative gel-purification assay for integration, electroporation was found to markedly increase the level of plasmid associated with high-molecular-weight genomic DNA. To confirm integration and identify the insertion sites, we developed a new assay – referred to as repeat-anchored integration capture (RAIC) PCR – that is capable of detecting rare integration events in a complex mixture in vivo. Using this assay, we identified four independent integration events. Sequencing of the insertion sites suggested a random integration process, but with short segments of homology between the vector breakpoint and the insertion site in three of the four cases. This is the first definitive demonstration of integration of plasmid DNA into genomic DNA following injection in vivo.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Herweijer H, Wolff JA . Progress and prospects: naked DNA gene transfer and therapy. Gene Therapy 2003; 10: 453–458.
Donnelly JJ, Ulmer JB, Shiver JW, Liu MA . DNA vaccines. Annu Rev Immunol 1997; 15: 617–648.
Nishikawa M, Huang L . Nonviral vectors in the new millennium: delivery barriers in gene transfer. Hum Gene Ther 2001; 12: 861–870.
Bigey P, Bureau MF, Scherman D . In vivo plasmid DNA electrotransfer. Curr Opin Biotechnol 2002; 13: 443–447.
Gehl J . Electroporation: theory and methods, perspectives for drug delivery, gene therapy and research. Acta Physiol Scand 2003; 177: 437–447.
Trezise AEO . In vivo DNA electrotransfer. DNA Cell Biol 2002; 21: 869–877.
Fattori E, La-Monica N, Ciliberto G, Toniatti C . Electro-gene-transfer: a new approach for muscle gene delivery. Somat Cell Mol Genet 2002; 27: 75–83.
Nichols WW, Ledwith BJ, Manam SV, Troilo PJ . Potential DNA vaccine integration into host cell genome. Ann NY Acad Sci 1995; 772: 30–39.
Ledwith BJ et al. Plasmid DNA vaccines: assay for integration into host genomic DNA. Dev Biol (Basel) 2000; 104: 33–43.
Ledwith BJ et al. Plasmid DNA vaccines: investigation of integration into host cellular DNA following intramuscular injection in mice. Intervirology 2000; 43: 258–272.
Manam S et al. Plasmid DNA vaccines: tissue distribution and effects of DNA sequence, adjuvants and delivery method on integration into host DNA. Intervirology 2000; 43: 273–281.
Martin T et al. Plasmid DNA malaria vaccine: the potential for genomic integration after intramuscular injection. Hum Gene Ther 1999; 10: 759–768.
Rizzuto G et al. Efficient and regulated erythropoietin production by naked DNA injection and muscle electroporation. Proc Natl Acad Sci USA 1999; 96: 6417–6422.
Minami M, Poussin K, Brechot C, Paterlini P . A novel PCR technique using Alu-specific primers to identify unknown flanking sequences from the human genome. Genomics 1995; 29: 403–408.
Sorensen AB, Duch M, Jorgensen P, Pedersen FS . Amplification and sequence analysis of DNA flanking integrated proviruses by a simple two-step polymerase chain reaction method. J Virol 1993; 67: 7118–7124.
Longo MC, Berninger MS, Hartley JL . Use of uracil DNA glycosylase to control carry-over contamination in polymerase chain reactions. Gene 1990; 93: 125–128.
Krayev AS et al. The nucleotide sequence of the ubiquitous repetitive DNA sequence B1 complementary to the most abundant class of mouse fold-back RNA. Nucleic Acids Res 1980; 8: 1201–1215.
Waterston RH, et al. Mouse Genome Sequencing Consortium. Initial sequencing and comparative analysis of the mouse genome. Nature 2002; 420: 520–562.
Zhang G et al. Efficient expression of naked DNA delivered intraarterially to limb muscles of nonhuman primates. Hum Gene Ther 2001; 12: 427–438.
Hui EK-W, Wang P-C, Lu SJ . Strategies for cloning unknown cellular flanking DNA sequences from foreign integrants. Cell Mol Life Sci 1998; 54: 1403–1411.
Jin YF, Ishibashi T, Nomoto A, Masuda M . Isolation and analysis of retroviral integration targets by solo long terminal repeat inverse PCR. J Virol 2002; 76: 5540–5547.
Würtele H, Little KCE, Chartrand P . Illegitimate DNA integration in mammalian cells. Gene Therapy 2003; 10: 1791–1799.
Merrihew RV et al. High-frequency illegitimate integration of transfected DNA at preintegrated target sites in a mammalian genome. Mol Cell Biol 1996; 16: 10–18.
Bode J et al. Fatal connections: when DNA ends meet on the nuclear matrix. J Cell Biochem 2000; Suppl 35: 3–22.
Wu X, Li Y, Crise B, Burgess SM . Transcription start regions in the human genome are favored targets for MLV integration. Science 2003; 300: 1749–1751.
Nakai H et al. AAV serotype 2 vectors preferentially integrate into active genes in mice. Nat Genet 2003; 34: 297–302.
Zhang L, Nolan E, Kreitschitz S, Rabussay DP . Enhanced delivery of naked DNA to the skin by non-invasive in vivo electroporation. Biochim Biophy Acta 2002; 1572: 1–9.
Gilbert RA, Jaroszeski MJ, Heller R . Novel electrode designs for electrochemotherapy. Biochim Biophys Acta 1997; 1334: 9–14.
Acknowledgements
We thank Carolann Beare, Steven Hall, Donna Lynch, and Brenda Givler for assistance with the antemortem portion of the study, and Yuan Liu for assistance with analysis of the integration sites using Bioinformatics. We also thank Warren Nichols for his continued interest and ever insightful discussions.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Wang, Z., Troilo, P., Wang, X. et al. Detection of integration of plasmid DNA into host genomic DNA following intramuscular injection and electroporation. Gene Ther 11, 711–721 (2004). https://doi.org/10.1038/sj.gt.3302213
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.gt.3302213
Keywords
This article is cited by
-
Harnessing Rift Valley fever virus NSs gene for cancer gene therapy
Cancer Gene Therapy (2022)
-
Characterization of integration frequency and insertion sites of adenovirus DNA into mouse liver genomic DNA following intravenous injection
Gene Therapy (2022)
-
Simulation study for the pulling translocation of a polymer globule
Polymer Journal (2021)
-
AIV polyantigen epitope expressed by recombinant baculovirus induces a systemic immune response in chicken and mouse models
Virology Journal (2020)
-
Recombinant PAL/PilE/FlaA DNA vaccine provides protective immunity against Legionella pneumophila in BALB/c mice
BMC Biotechnology (2020)