Abstract
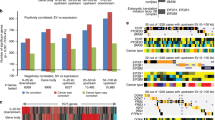
Single-nucleotide polymorphism (SNP) array analysis was performed using the 10K GeneChip array on a series of 26 paired follicular lymphoma (FL) and transformed-FL (t-FL) biopsies and the lymphoma cell lines SCI-1, DoHH2 and RL2261. Regions of acquired homozygosity were detected in 43/52 (83%) primary specimens with a mean of 1.7 and 3.0 aberrations in the FL and t-FL, respectively. A notable feature was the occurrence of recurring sites of acquired uniparental disomy (aUDP) on 6p, 9p, 12q and 17p in cell lines and primary samples. Homozygosity of 9p and 17p arose predominantly in t-FL and in three cases rendered the cell homozygous for a pre-existing mutation of either CDKN2A or TP53. These data suggest that mutation precedes mitotic recombination, which leads to the removal of the remaining wild-type allele. In all, 18 cases exhibited abnormalities in both FL and t-FL samples. In 10 cases blocks of homozygosity were detected in FL that were absent in the subsequent t-FL sample. These differences support the notion that FL and t-FL may arise in a proportion of patients by divergence from a common malignant ancestor cell rather than by clonal evolution from an antecedent FL.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Horning SJ . Natural history of and therapy for the indolent non-Hodgkin's lymphomas. Semin Oncol 1993; 20: 75–88.
Gallagher CJ, Gregory WM, Jones AE, Stansfeld AG, Richards MA, Dhaliwal HS et al. Follicular lymphoma: prognostic factors for response and survival. J Clin Oncol 1986; 4: 1470–1480.
Cullen MH, Lister TA, Brearley RI, Shand WS, Stansfeld AG . Histological transformation of non-Hodgkin's lymphoma: a prospective study. Cancer 1979; 44: 645–651.
Hubbard SM, Chabner BA, DeVita VT, Simon R, Berard CW, Jones RB et al. Histologic progression in non-Hodgkin's lymphoma. Blood 1982; 59: 258–264.
Acker B, Hoppe RT, Colby TV, Cox RS, Kaplan HS, Rosenberg SA . Histologic conversion in the non-Hodgkin's lymphomas. J Clin Oncol 1983; 1: 11–16.
Bastion Y, Sebban C, Berger F, Felman P, Salles G, Dumontet C et al. Incidence, predictive factors, and outcome of lymphoma transformation in follicular lymphoma patients. J Clin Oncol 1997; 15: 1587–1594.
Lossos IS . Higher-grade transformation of follicular lymphoma – a continuous enigma. Leukemia 2005; 19: 1331–1333.
Lossos IS, Levy R . Higher grade transformation of follicular lymphoma: phenotypic tumor progression associated with diverse genetic lesions. Semin Cancer Biol 2003; 13: 191–202.
Hoglund M, Sehn L, Connors JM, Gascoyne RD, Siebert R, Sall T et al. Identification of cytogenetic subgroups and karyotypic pathways of clonal evolution in follicular lymphomas. Genes Chromosomes Cancer 2004; 39: 195–204.
Martinez-Climent JA, Alizadeh AA, Segraves R, Blesa D, Rubio-Moscardo F, Albertson DG et al. Transformation of follicular lymphoma to diffuse large cell lymphoma is associated with a heterogeneous set of DNA copy number and gene expression alterations. Blood 2003; 101: 3109–3117.
Goff LK, Neat MJ, Crawley CR, Jones L, Jones E, Lister TA et al. The use of real-time quantitative polymerase chain reaction and comparative genomic hybridization to identify amplification of the REL gene in follicular lymphoma. Br J Haematol 2000; 111: 618–625.
Nagy M, Balazs M, Adam Z, Petko Z, Timar B, Szereday Z et al. Genetic instability is associated with histological transformation of follicle center lymphoma. Leukemia 2000; 14: 2142–2148.
Hough RE, Goepel JR, Alcock HE, Hancock BW, Lorigan PC, Hammond DW . Copy number gain at 12q12–14 may be important in the transformation from follicular lymphoma to diffuse large B cell lymphoma. Br J Cancer 2001; 84: 499–503.
Calhoun ES, Hucl T, Gallmeier E, West KM, Arking DE, Maitra A et al. Identifying allelic loss and homozygous deletions in pancreatic cancer without matched normals using high-density single-nucleotide polymorphism arrays. Cancer Res 2006; 66: 7920–7928.
Raghavan M, Lillington DM, Skoulakis S, Debernardi S, Chaplin T, Foot NJ et al. Genome-wide single nucleotide polymorphism analysis reveals frequent partial uniparental disomy due to somatic recombination in acute myeloid leukemias. Cancer Res 2005; 65: 375–378.
Gaasenbeek M, Howarth K, Rowan AJ, Gorman PA, Jones A, Chaplin T et al. Combined array-comparative genomic hybridization and single-nucleotide polymorphism-loss of heterozygosity analysis reveals complex changes and multiple forms of chromosomal instability in colorectal cancers. Cancer Res 2006; 66: 3471–3479.
Midorikawa Y, Yamamoto S, Ishikawa S, Kamimura N, Igarashi H, Sugimura H et al. Molecular karyotyping of human hepatocellular carcinoma using single-nucleotide polymorphism arrays. Oncogene 2006; 25: 5581–5590.
Nielaender I, Martin-Subero JI, Wagner F, Martinez-Climent JA, Siebert R . Partial uniparental disomy: a recurrent genetic mechanism alternative to chromosomal deletion in malignant lymphoma. Leukemia 2006; 20: 904–905.
Robinson WP . Mechanisms leading to uniparental disomy and their clinical consequences. Bioessays 2000; 22: 452–459.
James C, Ugo V, Casadevall N, Constantinescu SN, Vainchenker W . A JAK2 mutation in myeloproliferative disorders. Pathogenesis and therapeutic and scientific prospects. Trends Mol Med 2005; 11: 546–554.
Kralovics R, Passamonti F, Buser AS, Teo SS, Tiedt R, Passweg JR et al. A gain-of-function mutation of JAK2 in myeloproliferative disorders. N Engl J Med 2005; 352: 1779–1790.
Baxter EJ, Scott LM, Campbell PJ, East C, Fourouclas N, Swanton S et al. Acquired mutation of the tyrosine kinase JAK2 in human myeloproliferative disorders. Lancet 2005; 365: 1054–1061.
Levine RL, Wadleigh M, Cools J, Ebert BL, Wernig G, Huntly BJ et al. Activating mutation in the tyrosine kinase JAK2 in polycythemia vera, essential thrombocythemia, and myeloid metaplasia with myelofibrosis. Cancer Cell 2005; 7: 387–397.
Fitzgibbon J, Smith LL, Raghavan M, Smith ML, Debernardi S, Skoulakis S et al. Association between acquired uniparental disomy and homozygous gene mutation in acute myeloid leukemias. Cancer Res 2005; 65: 9152–9154.
Davies AJ, Lee AM, Taylor C, Clear AJ, Goff LK, Iqbal S et al. A limited role for TP53 mutation in the transformation of follicular lymphoma to diffuse large B-cell lymphoma. Leukemia 2005; 19: 1459–1465.
Tartaglia M, Kalidas K, Shaw A, Song X, Musat DL, van der Burgt I et al. PTPN11 mutations in Noonan syndrome: molecular spectrum, genotype–phenotype correlation, and phenotypic heterogeneity. Am J Hum Genet 2002; 70: 1555–1563.
Galmarini CM, Bouchet BP, Audoynaud C, Lamblot C, Falette N, Bertholon J et al. A p21/WAF1 mutation favors the appearance of drug resistance to paclitaxel in human noncancerous epithelial mammary cells. Int J Cancer 2006; 119: 60–66.
Engel E . A fascination with chromosome rescue in uniparental disomy: Mendelian recessive outlaws and imprinting copyrights infringements. Eur J Hum Genet 2006; 14: 1158–1169.
Murthy SK, DiFrancesco LM, Ogilvie RT, Demetrick DJ . Loss of heterozygosity associated with uniparental disomy in breast carcinoma. Mod Pathol 2002; 15: 1241–1250.
Teh MT, Blaydon D, Chaplin T, Foot NJ, Skoulakis S, Raghavan M et al. Genomewide single nucleotide polymorphism microarray mapping in basal cell carcinomas unveils uniparental disomy as a key somatic event. Cancer Res 2005; 65: 8597–8603.
Dave SS, Wright G, Tan B, Rosenwald A, Gascoyne RD, Chan WC et al. Prediction of survival in follicular lymphoma based on molecular features of tumor-infiltrating immune cells. N Engl J Med 2004; 351: 2159–2169.
Lindblad-Toh K, Tanenbaum DM, Daly MJ, Winchester E, Lui WO, Villapakkam A et al. Loss-of-heterozygosity analysis of small-cell lung carcinomas using single-nucleotide polymorphism arrays. Nat Biotechnol 2000; 18: 1001–1005.
Zhao X, Li C, Paez JG, Chin K, Janne PA, Chen TH et al. An integrated view of copy number and allelic alterations in the cancer genome using single nucleotide polymorphism arrays. Cancer Res 2004; 64: 3060–3071.
Peiffer DA, Le JM, Steemers FJ, Chang W, Jenniges T, Garcia F et al. High-resolution genomic profiling of chromosomal aberrations using Infinium whole-genome genotyping. Genome Res 2006; 16: 1136–1148.
Pinyol M, Cobo F, Bea S, Jares P, Nayach I, Fernandez PL et al. p16(INK4a) gene inactivation by deletions, mutations, and hypermethylation is associated with transformed and aggressive variants of non-Hodgkin's lymphomas. Blood 1998; 91: 2977–2984.
Elenitoba-Johnson KS, Gascoyne RD, Lim MS, Chhanabai M, Jaffe ES, Raffeld M . Homozygous deletions at chromosome 9p21 involving p16 and p15 are associated with histologic progression in follicle center lymphoma. Blood 1998; 91: 4677–4685.
Kurata M, Maesako Y, Ueda C, Nishikori M, Akasaka T, Uchiyama T et al. Characterization of t(3;6)(q27;p21) breakpoints in B-cell non-Hodgkin's lymphoma and construction of the histone H4/BCL6 fusion gene, leading to altered expression of Bcl-6. Cancer Res 2002; 62: 6224–6230.
Jordanova ES, Philippo K, Giphart MJ, Schuuring E, Kluin PM . Mutations in the HLA class II genes leading to loss of expression of HLA-DR and HLA-DQ in diffuse large B-cell lymphoma. Immunogenetics 2003; 55: 203–209.
Greenwald RJ, Tumang JR, Sinha A, Currier N, Cardiff RD, Rothstein E et al. mu-BRD2 transgenic mice develop B-cell lymphoma and leukemia. Blood 2004; 103: 1475–1484.
Chen W, Itoyama T, Chaganti RS . Splicing factor SRP20 is a novel partner of BCL6 in a t(3;6)(q27;p21) translocation in transformed follicular lymphoma. Genes Chromosomes Cancer 2001; 32: 281–284.
Shaughnessy J, Gabrea A, Qi Y, Brents L, Zhan F, Tian E et al. Cyclin D3 at 6p21 is dysregulated by recurrent chromosomal translocations to immunoglobulin loci in multiple myeloma. Blood 2001; 98: 217–223.
Smith ML, Arch R, Smith LL, Bainton N, Neat M, Taylor C et al. Development of a human acute myeloid leukaemia screening panel and consequent identification of novel gene mutation in FLT3 and CCND3. Br J Haematol 2005; 128: 318–323.
Song L, Zlobin A, Ghoshal P, Zhang Q, Houde C, Weijzen S et al. Alteration of SMRT tumor suppressor function in transformed non-Hodgkin lymphomas. Cancer Res 2005; 65: 4554–4561.
Matolcsy A, Schattner EJ, Knowles DM, Casali P . Clonal evolution of B cells in transformation from low- to high-grade lymphoma. Eur J Immunol 1999; 29: 1253–1264.
Acknowledgements
We thank Randy Gascoyne for sharing his unpublished flow data in FL, Susan Weiss for assistance in preparing the article, Iram Gull and Victoria Miller for sample collection and storage, and Jean-Baptiste Cazier and Manu Gupta for helpful comments on the article. Derville O’Shea is a MRC training fellow and Emanuela Carlotti is a Bart's and the Royal London Charitable Foundation fellow. The work was also supported by the Cancer Research UK and Kay Kendall Leukaemia Fund.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the National Centre for Biotechnology Information website (http://www.ncbi.nlm.nih.gov/projects/geo/query/acc.cgi?acc=GSE7425)
Rights and permissions
About this article
Cite this article
Fitzgibbon, J., Iqbal, S., Davies, A. et al. Genome-wide detection of recurring sites of uniparental disomy in follicular and transformed follicular lymphoma. Leukemia 21, 1514–1520 (2007). https://doi.org/10.1038/sj.leu.2404696
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.leu.2404696