Abstract
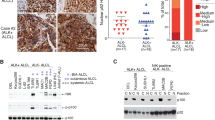
Using a cDNA microarray, we found that suppressor of cytokine signaling 3 (SOCS3) is highly expressed in anaplastic lymphoma kinase (ALK)+ anaplastic large cell lymphoma (ALCL) cell lines. As SOCS3 is induced by activated signal transducer and activator of transcription 3 (STAT3), and ALK activates STAT3, we hypothesized that SOCS3 may play a role in ALK+ ALCL pathogenesis via the Janus kinase 3 (JAK3)-STAT3 pathway. Using ALCL cell lines, we show by coimmunoprecipitation experiments that SOCS3 physically binds with JAK3 in vitro, and that JAK3 inhibition by WHI-P154 downregulates SOCS3 expression. Western blot analysis confirmed expression of SOCS3 and also showed coexpression of phosphorylated (activated) STAT3 (pSTAT3). Direct sequencing of the SOCS3 gene showed no mutations or alternative splicing. In ALCL tumors that were assessed by immunohistochemistry, nine of 12 (75%) ALK+ tumors were SOCS3 positive and eight (67%) coexpressed pSTAT3. In comparison, 18 of 25 (72%) ALK-- tumors were SOCS3 positive and seven (28%) coexpressed pSTAT3. These results show that SOCS3 is overexpressed in ALCL, attributable to JAK3-STAT3 activation and likely related to ALK in ALK+ tumors. However, SOCS3 is also expressed in tumors that lack STAT3 and ALK suggesting alternative mechanisms of upregulation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Starr R, Willson TA, Viney EM, Murray LJ, Rayner JR, Jenkins BJ et al. A family of cytokine-inducible inhibitors of signalling. Nature 1997; 387: 917–921.
Zhang JG, Farley A, Nicholson SE, Willson TA, Zugaro LM, Simpson RJ et al. The conserved SOCS box motif in suppressors of cytokine signaling binds to elongins B and C and may couple bound proteins to proteasomal degradation. Proc Natl Acad Sci USA 1999; 96: 2071–2076.
Auernhammer CJ, Bousquet C, Melmed S . Autoregulation of pituitary corticotroph SOCS-3 expression: characterization of the murine SOCS-3 promoter. Proc Natl Acad Sci USA 1999; 96: 6964–6969.
Alexander WS . Suppressors of cytokine signalling (SOCS) in the immune system. Nat Rev Immunol 2002; 2: 410–416.
Chen XP, Losman JA, Rothman P . SOCS proteins, regulators of intracellular signaling. Immunity 2000; 13: 287–290.
Shouda T, Yoshida T, Hanada T, Wakioka T, Oishi M, Miyoshi K et al. Induction of the cytokine signal regulator SOCS3/CIS3 as a therapeutic strategy for treating inflammatory arthritis. J Clin Invest 2001; 108: 1781–1788.
Suzuki A, Hanada T, Mitsuyama K, Yoshida T, Kamizono S, Hoshino T et al. CIS3/SOCS3/SSI3 plays a negative regulatory role in STAT3 activation and intestinal inflammation. J Exp Med 2001; 193: 471–481.
Delsol G, Ralfkiaer E, Stein H, Wright D, Jaffe ES . Anaplastic large cell lymphoma. In: Jaffe ES, Harris NL, Stein H, Vardiman JW (eds). World Health Organization Classification of Tumours. Pathology and Genetics of Tumors of Haematopoietic and Lymphoid Tissues. Lyon, France: IARC Press, 2001, pp 230–235.
Drexler HG, Gignac SM, von Wasielewski R, Werner M, Dirks WG . Pathobiology of NPM-ALK and variant fusion genes in anaplastic large cell lymphoma and other lymphomas. Leukemia 2000; 14: 1533–1559.
Morris SW, Kirstein MN, Valentine MB, Dittmer KG, Shapiro DN, Saltman DL et al. Fusion of a kinase gene, ALK, to a nucleolar protein gene, NPM, in non-Hodgkin's lymphoma. Science 1994; 263: 1281–1284.
Falini B, Pileri S, Zinzani PL, Carbone A, Zagonel V, Wolf-Peeters C et al. ALK+ lymphoma: clinico-pathological findings and outcome. Blood 1999; 93: 2697–2706.
Shiota M, Nakamura S, Ichinohasama R, Abe M, Akagi T, Takeshita M et al. Anaplastic large cell lymphomas expressing the novel chimeric protein p80NPM/ALK: a distinct clinicopathologic entity. Blood 1995; 86: 1954–1960.
Suzuki R, Kagami Y, Takeuchi K, Kami M, Okamoto M, Ichinohasama R et al. Prognostic significance of CD56 expression for ALK-positive and ALK-negative anaplastic large-cell lymphoma of T/null cell phenotype. Blood 2000; 96: 2993–3000.
Haralambieva E, Pulford KA, Lamant L, Pileri S, Roncador G, Gatter KC et al. Anaplastic large-cell lymphomas of B-cell phenotype are anaplastic lymphoma kinase (ALK) negative and belong to the spectrum of diffuse large B-cell lymphomas. Br J Haematol 2000; 109: 584–591.
Zamo A, Chiarle R, Piva R, Howes J, Fan Y, Chilosi M et al. Anaplastic lymphoma kinase (ALK) activates Stat3 and protects hematopoietic cells from cell death. Oncogene 2002; 21: 1038–1047.
Bromberg J . Stat proteins and oncogenesis. J Clin Invest 2002; 109: 1139–1142.
Khoury JD, Medeiros LJ, Rassidakis GZ, Yared MA, Tsioli P, Leventaki V et al. Differential expression and clinical significance of tyrosine-phosphorylated STAT3 in ALK+ and ALK− anaplastic large cell lymphoma. Clin Cancer Res 2003; 9: 3692–3699.
Amin HM, Medeiros LJ, Ma Y, Feretzaki M, Das P, Leventaki V et al. Inhibition of JAK3 induces apoptosis and decreases anaplastic lymphoma kinase activity in anaplastic large cell lymphoma. Oncogene 2003; 22: 5399–5407.
Alas S, Bonavida B . Rituximab inactivates signal transducer and activation of transcription 3 (STAT3) activity in B-non-Hodgkin's lymphoma through inhibition of the interleukin 10 autocrine/paracrine loop and results in down-regulation of Bcl-2 and sensitization to cytotoxic drugs. Cancer Res 2001; 61: 5137–5144.
Bowman T, Broome MA, Sinibaldi D, Wharton W, Pledger WJ, Sedivy JM et al. Stat3-mediated Myc expression is required for Src transformation and PDGF-induced mitogenesis. Proc Natl Acad Sci USA 2001; 98: 7319–7324.
Coqueret O, Gascan H . Functional interaction of STAT3 transcription factor with the cell cycle inhibitor p21WAF1/CIP1/SDI1. J Biol Chem 2000; 275: 18794–18800.
Epling-Burnette PK, Liu JH, Catlett-Falcone R, Turkson J, Oshiro M, Kothapalli Jr R et al. Inhibition of STAT3 signaling leads to apoptosis of leukemic large granular lymphocytes and decreased Mcl-1 expression. J Clin Invest 2001; 107: 351–362.
Grad JM, Zeng XR, Boise LH . Regulation of Bcl-xL: a little bit of this and a little bit of STAT. Curr Opin Oncol 2000; 12: 543–549.
Naka T, Matsumoto T, Narazaki M, Fujimoto M, Morita Y, Ohsawa Y et al. Accelerated apoptosis of lymphocytes by augmented induction of Bax in SSI-1 (STAT-induced STAT inhibitor-1) deficient mice. Proc Natl Acad Sci USA 1998; 95: 15577–15582.
Ning ZQ, Li J, McGuinness M, Arceci RJ . STAT3 activation is required for Asp(816) mutant c-Kit induced tumorigenicity. Oncogene 2001; 20: 4528–4536.
Baggerly KA, Coombes KR, Hess KR, Stivers DN, Abruzzo LV, Zhang W . Identifying differentially expressed genes in cDNA microarray experiments. J Comput Biol 2001; 8: 639–659.
Fischer P, Nacheva E, Mason DY, Sherrington PD, Hoyle C, Hayhoe FG et al. A Ki-1 (CD30)-positive human cell line (Karpas 299) established from a high-grade non-Hodgkin's lymphoma, showing a 2;5 translocation and rearrangement of the T-cell receptor beta-chain gene. Blood 1988; 72: 234–240.
Sudbeck EA, Liu XP, Narla RK, Mahajan S, Ghosh S, Mao C et al. Structure-based design of specific inhibitors of Janus kinase 3 as apoptosis-inducing antileukemic agents. Clin Cancer Res 1999; 5: 1569–1582.
Sasaki A, Yasukawa H, Suzuki A, Kamizono S, Syoda T, Kinjyo I et al. Cytokine-inducible SH2 protein-3 (CIS3/SOCS3) inhibits Janus tyrosine kinase by binding through the N-terminal kinase inhibitory region as well as SH2 domain. Genes Cells 1999; 4: 339–351.
Chiarle R, Gong JZ, Guasparri I, Pesci A, Cai J, Liu J et al. NPM-ALK transgenic mice spontaneously develop T-cell lymphomas and plasma cell tumors. Blood 2003; 101: 1919–1927.
Amin HM, McDonnell TJ, Ma Y, Lin Q, Fujio Y, Kunisada K et al. Selective inhibition of STAT3 induces apoptosis and G(1) cell cycle arrest in ALK-positive anaplastic large cell lymphoma. Oncogene 2004; 23: 5426–5431.
Zhang Q, Raghunath PN, Xue L, Majewski M, Carpentieri DF, Odum N et al. Multilevel dysregulation of STAT3 activation in anaplastic lymphoma kinase-positive T/null-cell lymphoma. J Immunol 2002; 168: 466–474.
Sasaki A, Inagaki-Ohara K, Yoshida T, Yamanaka A, Sasaki M, Yasukawa H et al. The N-terminal truncated isoform of SOCS3 translated from an alternative initiation AUG codon under stress conditions is stable due to the lack of a major ubiquitination site, Lys-6. J Biol Chem 2003; 278: 2432–2436.
Morris SW, Xue L, Ma Z, Kinney MC . Alk+ CD30+ lymphomas: a distinct molecular genetic subtype of non-Hodgkin's lymphoma. Br J Haematol 2001; 113: 275–295.
Herling M, Rassidakis GZ, Viviani S, Bonfante V, Giardini R, Gianni M et al. Anaplastic lymphoma kinase (ALK) is not expressed in Hodgkin's disease: results with ALK-11 antibody in 327 untreated patients. Leuk Lymphoma 2001; 42: 969–979.
Brender C, Nielsen M, Kaltoft K, Mikkelsen G, Zhang Q, Wasik M et al. STAT3-mediated constitutive expression of SOCS-3 in cutaneous T-cell lymphoma. Blood 2001; 97: 1056–1062.
Sommer VH, Clemmensen OJ, Nielsen O, Wasik M, Lovato P, Brender C et al. In vivo activation of STAT3 in cutaneous T-cell lymphoma. Evidence for an antiapoptotic function of STAT3. Leukemia 2004; 18: 1288–1295.
Darnell Jr JE, Kerr IM, Stark GR . Jak-STAT pathways and transcriptional activation in response to IFNs and other extracellular signaling proteins. Science 1994; 264: 1415–1421.
Lin J, Tang H, Jin X, Jia G, Hsieh JT . p53 regulates Stat3 phosphorylation and DNA binding activity in human prostate cancer cells expressing constitutively active Stat3. Oncogene 2002; 21: 3082–3088.
Vignais ML, Gilman M . Distinct mechanisms of activation of Stat1 and Stat3 by platelet-derived growth factor receptor in a cell-free system. Mol Cell Biol 1999; 19: 3727–3735.
Kube D, Holtick U, Vockerodt M, Ahmadi T, Haier B, Behrmann I et al. STAT3 is constitutively activated in Hodgkin cell lines. Blood 2001; 98: 762–770.
Garcia JF, Camacho FI, Morente M, Fraga M, Montalban C, Alvaro T et al. Hodgkin and Reed–Sternberg cells harbor alterations in the major tumor suppressor pathways and cell-cycle checkpoints: analyses using tissue microarrays. Blood 2003; 101: 681–689.
Coppo P, Dusanter-Fourt I, Millot G, Nogueira MM, Dugray A, Bonnet ML et al. Constitutive and specific activation of STAT3 by BCR-ABL in embryonic stem cells. Oncogene 2003; 22: 4102–4110.
Garcia R, Yu CL, Hudnall A, Catlett R, Nelson KL, Smithgall T et al. Constitutive activation of Stat3 in fibroblasts transformed by diverse oncoproteins and in breast carcinoma cells. Cell Growth Differ 1997; 8: 1267–1276.
Nelson KL, Rogers JA, Bowman TL, Jove R, Smithgall TE . Activation of STAT3 by the c-Fes protein-tyrosine kinase. J Biol Chem 1998; 273: 7072–7077.
Smith PD, Crompton MR . Expression of v-src in mammary epithelial cells induces transcription via STAT3. Biochem J 1998; 331 (Part 2): 381–385.
Krebs DL, Hilton DJ . SOCS proteins: negative regulators of cytokine signaling. Stem Cells 2001; 19: 378–387.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Cho-Vega, J., Rassidakis, G., Amin, H. et al. Suppressor of cytokine signaling 3 expression in anaplastic large cell lymphoma. Leukemia 18, 1872–1878 (2004). https://doi.org/10.1038/sj.leu.2403495
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.leu.2403495
Keywords
This article is cited by
-
Programmed death-ligand 1 (PD-L1) characterization of circulating tumor cells (CTCs) in muscle invasive and metastatic bladder cancer patients
BMC Cancer (2016)
-
Gene expression profiling of isolated tumour cells from anaplastic large cell lymphomas: insights into its cellular origin, pathogenesis and relation to Hodgkin lymphoma
Leukemia (2009)
-
JunB expression is a common feature of CD30+ lymphomas and lymphomatoid papulosis
Modern Pathology (2005)
-
Inhibition of ALK enzymatic activity in T-cell lymphoma cells induces apoptosis and suppresses proliferation and STAT3 phosphorylation independently of Jak3
Laboratory Investigation (2005)