Abstract
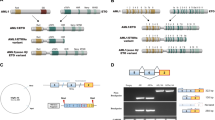
Chromosomal band 1p34–36 is a commonly rearranged locus in many types of cancers. We cloned the breakpoint region of a chromosomal translocation, t(1;14)(p34;q32), found in the human multiple myeloma (MM) cell line, ODA. This rearrangement occurred between the nearby switch region of the immunoglobulin heavy chain (IgH) gene (Sγ3) at 14q32 and the first intron of the human retinoic acid-inducible E3 protein (E3)/lysosome-associated protein, transmembrane-5 (LAPTm5) gene at the 1p34 locus. Consequently, the E3 gene, which is a hematopoietic cell-specific transcript induced by retinoic acid and located at the rearranged allele, was interrupted within its coding region and was not expressed in the ODA cell line in spite of the other allele still being intact. The expression derived from the remaining intact allele in ODA cells was silenced by DNA methylation at sequences within the first intron around a GC-rich EagI site. Interestingly, the silenced expression of E3 mRNA due to DNA methylation of intron 1 sequences was frequently encountered in MM cells [6/10 (60%) of MM cell lines tested], while E3 is expressed in normal plasma cells and in most other hematopoietic cell lines including those of B-cell lineage. Thus, as the E3 protein has been suggested to be involved in cellular differentiation and apoptotic pathways in certain cell types, our results suggest that loss of E3 gene expression might be a crucial event during the progression of human MM.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Wlodarskia I, Mecucci C, Stul M, Michaux L, Pittaluga S, Hernandez JM et al. Fluorescence in situ hybridization identifies new chromosomal changes involving 3q27 in non-Hodgkin's lymphomas with BCL6/LAZ3 rearrangement. Genes Chromosom Cancer 1995; 1: 1–7.
Dierlamm J, Pittaluga S, Wlodarska I, Stul M, Thomas J, Boogaerts M et al. Marginal zone B-cell lymphomas of different sites share similar cytogenetic and morphologic features. Blood 1996; 87: 299–307.
Emi M, Yoshimoto M, Sato T, Matsumoto S, Utada Y, Ito I et al. Allelic loss at 1p34, 13q12, 17p13.3, and 17q21.1 correlates with poor postoperative prognosis in breast cancer. Genes Chromosom Cancer 1999; 26: 134–141.
Kim GJ, Park SY, Kim H, Chun YH, Park SH . Chromosomal aberrations in neuroblastoma cell lines identified by cross species color banding and chromosome painting. Cancer Genet Cytogenet 2001; 129: 10–16.
Derre J, Lagace R, Nicolas A, Mairal A, Chibon F, Coindre JM et al. Leiomyosarcomas and most malignant fibrous histiocytomas share very similar comparative genomic hybridization imbalances: an analysis of a series of 27 leiomyosarcomas. Lab Invest 2001; 81: 211–215
Burnett RC, Thirman MJ, Rowley JD, Diaz MO . Molecular analysis of the T-cell acute lymphoblastic leukemia-associated t(1;7) (p34;q34) that fuses LCK and TCRB. Blood 1994; 84: 1232–1236.
Chesi M, Bergsagel PL, Brents LA, Smith CM, Gerhard DS, Kuehl WM . Dysregulation of cyclin D1 by translocation into an IgH gamma switch region in two multiple myeloma cell lines. Blood 1996; 88: 674–681.
Iida S, Rao PH, Butler M, Corradini P, Boccadoro M, Klein B et al. Deregulation of MUM1/IRF4 by chromosomal translocation in multiple myeloma. Nat Genet 1997; 17: 226–230.
Yoshida S, Nakazawa N, Iida S, Hayami Y, Sato S, Wakita A et al. Detection of MUM1/IRF4-IgH fusion in multiple myeloma. Leukemia 1999; 13: 1812–1816.
Chesi M, Nardini E, Brents LA, Schrock E, Ried T, Kuehl WM, Bergsagel PL . Frequent translocation t(4;14)(p16.3;q32.3) in multiple myeloma is associated with increased expression and activating mutations of fibroblast growth factor receptor 3. Nat Genet 1997; 16: 260–264.
Chesi M, Bergsagel PL, Shonukan OO, Martelli ML, Brents LA, Chen T et al. Frequent dysregulation of the c-maf proto-oncogene at 16q23 by translocation to an Ig locus in multiple myeloma. Blood 1998; 91: 4457–4463.
Avet-Loiseau H, Gerson F, Magrangeas F, Minvielle S, Harousseau JL, Bataille R, Intergroupe Francophone du Myelome. Rearrangements of the c-myc oncogene are present in 15% of primary human multiple myeloma tumors. Blood 2001; 98: 2853–2855.
Hanamura I, Iida S, Akano Y, Hayami Y, Kato M, Miura K et al. Ectopic expression of MAFB gene in human myeloma cells carrying (14;20)(q32;q11) chromosomal translocations. Jpn J Cancer Res 2001; 92: 638–644.
Iida S, Hanamura I, Suzuki T, Kamiya T, Kato M, Hayami Y et al. A novel human multiple myeloma-derived cell line, NCU-MM-1, carrying t(2;11)(q11;q23) and t(8;22)(q24;q11) chromosomal translocations with overexpression of c-Myc protein. Int J Hematol 2000; 72: 85–91.
Seto M, Yamamoto K, Iida S, Akao Y, Utsumi KR, Kubonishi I et al. Gene rearrangement and overexpression of PRAD1 in lymphoid malignancy with t(11;14)(q13;q32) translocation. Oncogene 1992; 7: 1401–1406.
Tagawa S, Doi S, Taniwaki M, Abe T, Kanayama Y, Nojima J et al. Amylase-producing plasmacytoma cell lines, AD3 and FR4, with der(14)t(8;14) and dic(8)t(1;8) established from ascites. Leukemia 1990; 4: 600–605.
Zhang XG, Gaillard JP, Robillard N, Lu ZY, Gu ZJ, Jourdan M et al. Reproducible obtaining of human myeloma cell lines as a model for tumor stem cell study in human multiple myeloma. Blood 1994; 83: 3654–3663.
Tsuboi K, Iida S, Inagaki H, Kato M, Hayami Y, Hanamura I et al. MUM1/IRF4 expression as a frequent event in mature lymphoid malignancies. Leukemia 2000; 14: 449–456.
Iida S, Rao PH, Nallasivam P, Hibshoosh H, Butler M, Louie DC et al. The t(9;14)(p13;q32) chromosomal translocation associated with lymphoplasmacytoid lymphoma involves the PAX-5 gene. Blood 1996; 88: 4110–4117.
Taniwaki M, Nishida K, Ueda Y, Misawa S, Nagai M, Tagawa S et al. Interphase and metaphase detection of the breakpoint of 14q32 translocations in B-cell malignancies by double-color fluorescence in situ hybridization. Blood 1995; 85: 3223–3228.
Adra CN, Zhu S, Ko JL, Guillemot JC, Cuervo AM, Kobayashi H et al. LAPTM5: a novel lysosomal-associated multispanning membrane protein preferentially expressed in hematopoietic cells. Genomics 1996; 35: 328–337.
Scott LM, Mueller L, Collins SJ . E3, a hematopoietic-specific transcript directly regulated by the retinoic acid receptor alpha. Blood 1996; 88: 2517–2530.
Origasa M, Tanaka S, Suzuki K, Tone S, Lim B, Koike T . Activation of a novel microglial gene encoding a lysosomal membrane protein in response to neuronal apoptosis. Mol Brain Res 2001; 88: 1–13.
Hangaishi A, Ogawa S, Imamura N, Miyawaki S, Miura Y, Uike N et al. Inactivation of multiple tumor-suppressor genes involved in negative regulation of the cell cycle, MTS1/p16INK4A/CDKN2, MTS2/p15INK4B, p53, and Rb genes in primary lymphoid malignancies. Blood 1996; 87: 4949–4958.
Roman-Gomez J, Castillejo JA, Jimenez A, Gonzalez MG, Moreno F, Rodriguez MDJ et al. 5′ CpG island hypermethylation is associated with transcriptional silencing of the p21(CIP1/WAF1/SDI1) gene and confers poor prognosis in acute lymphoblastic leukemia. Blood 2002; 99: 2291–2296.
Ng MHL, Chung YF, Lo KW, Wickham NWR, Lee JCK, Huang DP . Frequent hypermethylation of p16 and p15 genes in multiple myeloma. Blood 1997; 89: 2500–2506.
Hallek M, Bergsagel PL, Anderson KC . Multiple myeloma: increasing evidence for a multistep transformation process. Blood 1998; 91: 3–21.
Dave BJ, Hess MM, Pickering DL, Zaleski DH, Pfeifer AL, Weisenburger DD et al. Rearrangements of chromosome band Ip36 in non-Hodgkin's lymphoma. Clin Cancer Res 1999; 5: 1401–1409.
Duro D, Bernard O, Della Valle V, Leblanc T, Berger R, Larsen CJ . Inactivation of the P16INK4/MTS1 gene by a chromosome translocation t(9;14)(p21-22;q11) in an acute lymphoblastic leukemia of B-cell type. Cancer Res 1996; 56: 848–854.
Acknowledgements
We thank Ms C Fukuyama and C Nakagawa for their skillful technical assistance. This work was supported in part by a Grant for S Iida and R Ueda from the Ministry of Education, Science, Sports and Culture, Japan and a Research Grant of the Princess Takamatsu Cancer Research Fund for S Iida.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Hayami, Y., Iida, S., Nakazawa, N. et al. Inactivation of the E3/LAPTm5 gene by chromosomal rearrangement and DNA methylation in human multiple myeloma. Leukemia 17, 1650–1657 (2003). https://doi.org/10.1038/sj.leu.2403026
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.leu.2403026
Keywords
This article is cited by
-
c-Myc inhibits LAPTM5 expression in B-cell lymphomas
Annals of Hematology (2023)
-
Consistent inverse correlation between DNA methylation of the first intron and gene expression across tissues and species
Epigenetics & Chromatin (2018)
-
TINAGL1 and B3GALNT1 are potential therapy target genes to suppress metastasis in non-small cell lung cancer
BMC Genomics (2014)
-
High-resolution genome-wide array-based comparative genome hybridization reveals cryptic chromosome changes in AML and MDS cases with trisomy 8 as the sole cytogenetic aberration
Leukemia (2006)
-
Antimyeloma effects of a novel synthetic retinoid Am80 (Tamibarotene) through inhibition of angiogenesis
Leukemia (2005)