Abstract
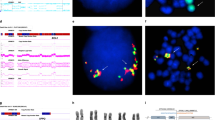
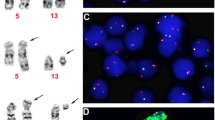
The orphan homeobox gene HOX11L2 was previously found to be transcriptionally activated as a result of the t(5;14)(q35;q32) translocation in three T-ALL cases. We now tested by RT-PCR Hox11L2 expression in 23 consecutive cases of T-ALL (15 children aged 0.8–14 years, eight adults aged 17–55 years) and as control 13 B-ALL patients from a single institution. Hox11L2 expression was undetectable in all patients with B-ALL, nor in adults with T-ALL. Nine children (60% of the cases), all boys, expressed Hox11L2. Blast cells from most of the latter patients carried surface CD1a, CD10 and not CD34 antigens, in contrast to the other children. FISH, M-FISH and IPM-FISH analysis failed to detect a t(5;14)(q35;q32) in one of them, which suggests a possible distinct genetic mechanism in Hox11L2 expression induction. Hence, Hox11L2 expression seems to be the most frequent abnormality in childhood T-ALL to date, comparable to the t(12;21) in child B-ALL.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Schneider NR, Carroll AJ, Shuster JJ, Pullen DJ, Link MP, Borowitz MJ, Camitta BM, Katz JA, Amylon MD . New recurring cytogenetic abnormalities and association of blast cell karyotypes with prognosis in childhood T-cell acute lymphoblastic leukemia: a Pediatric Oncology Group report of 343 cases Blood 2000 96: 2543–2549
Heerema NA, Sather HN, Sensel MG, Liu-Mares W, Lange BJ, Bostrom BC, Nachman JB, Steinherz PG, Hutchinson R, Gaynon PS, Arthur DC, Uckun FM . Association of chromosome arm 9p abnormalities with adverse risk in childhood acute lymphoblastic leukemia: a report from the Children's Cancer Group Blood 1999 94: 1537–1544
Delabesse E, Bernard M, Landman-Parker J, Davi F, Leboeuf D, Varet B, Valensi F, Macintyre EA . Simultaneous SIL-TAL1 RT-PCR detection of all tal(d) deletions and identification of novel tal(d) variants Br J Haematol 1997 99: 901–907
Dong WF, Billia F, Atkins HL, Iscove NN, Minden MD . Expression of rhombotin 2 in normal and leukaemic haemopoietic cells Br J Haematol 1996 93: 280–286
Lowsky R, DeCoteau JF, Reitmair AH, Ichinohasama R, Dong WF, Xu Y, Mak TW, Kadin ME, Minden MD . Defects of the mismatch repair gene MSH2 are implicated in the development of murine and human lymphoblastic lymphomas and are associated with the aberrant expression of rhombotin-2 (Lmo-2) and Tal-1 (SCL) Blood 1997 89: 2276–2282
Hatano M, Roberts CW, Minden M, Crist WM, Korsmeyer SJ . Deregulation of a homeobox gene, HOX11, by the t(10;14) in T cell leukemia Science 1991 253: 79–82
Salvati PD, Ranford PR, Ford J, Kees UR . HOX11 expression in pediatric acute lymphoblastic leukemia is associated with T-cell phenotype Oncogene 1995 11: 1333–1338
Bash RO, Crist WM, Shuster JJ, Link MP, Amylon M, Pullen J, Carroll AJ, Buchanan GR, Smith RG, Baer R . Clinical features and outcome of T-cell acute lymphoblastic leukemia in childhood with respect to alterations at the TAL1 locus: a Pediatric Oncology Group study Blood 1993 81: 2110–2117
Helias C, Leymarie V, Entz-Werle N, Falkenrodt A, Eyer D, Aurich-Costa J, Cherif D, Lutz P, Lessard M . Translocation t(5;14)(q35;q32) in 3 cases of childhood T-cell acute lymphoblastic leukemia: a new recurring and cryptic abnormality Leukemia 2002 16: 7–12
Bernard OA, Busson-LeConiat M, Ballerini P, Mauchauffe M, Della Valle V, Monni R, Nguyen Khac F, Mercher T, Penard-Lacronique V, Pasturaud P, Gressin L, Heilig R, Daniel MT, Lessard M, Berger R . A new recurrent and specific cryptic translocation, t(5;14)(q35;q32), is associated with expression of the Hox11L2 gene in T acute lymphoblastic leukemia Leukemia 2001 15: 1495–1504
Shirasawa S, Arata A, Onimaru H, Roth KA, Brown GA, Horning S, Arata S, Okumura K, Sasazuki T, Korsmeyer SJ . Rnx deficiency results in congenital central hypoventilation Nat Genet 2000 24: 287–290
Webber LM, Garson OM . Fluorodeoxyuridine synchronization of bone marrow cultures Cancer Genet Cytogenet 1983 8: 123–132
Aurich-Costa J, Vannier A, Gregoire E, Nowak F, Cherif D . IPM-FISH, a new M-FISH approach using IRS-PCR painting probes: application to the analysis of seven human prostate cell lines Genes Chromosomes Cancer 2001 30: 143–160
Speicher MR, Gwyn Ballard S, Ward DC . Karyotyping human chromosomes by combinatorial multi-fluor FISH Nat Genet 1996 12: 368–375
Eils R, Uhrig S, Saracoglu K, Satzler K, Bolzer A, Petersen I, Chassery J, Ganser M, Speicher MR . An optimized, fully automated system for fast and accurate identification of chromosomal rearrangements by multiplex-FISH (M-FISH) Cytogenet Cell Genet 1998 82: 160–171
Bene MC, Castoldi G, Knapp W, Ludwig WD, Matutes E, Orfao A, van't Veer MB . Proposals for the immunological classification of acute leukemias. European Group for the Immunological Characterization of Leukemias (EGIL) Leukemia 1995 9: 1783–1786
Crist WM, Shuster JJ, Falletta J, Pullen DJ, Berard CW, Vietti TJ, Alvarado CS, Roper MA, Prasthofer E, Grossi CE . Clinical features and outcome in childhood T-cell leukemia-lymphoma according to stage of thymocyte differentiation: a Pediatric Oncology Group Study Blood 1988 72: 1891–1897
Ferrando AA, Neuberg DS, Staunton J, LOH ML, Huard C, Raimondi SC, Behm MG, Pui C-H, Downing JR, Gilliland DG, Lander ES, Golub TR, Look AT . Gene expression signatures define novel oncogenic pathways in T cell acute lymphoblastic leukemia Cancer Cell 2002 1: 75–87
Dear TN, Sanchez-Garcia I, Rabbitts TH . The HOX11 gene encodes a DNA-binding nuclear transcription factor belonging to a distinct family of homeobox genes Proc Natl Acad Sci USA 1993 90: 4431–4435
Kennedy MA, Gonzalez-Sarmiento R, Kees UR, Lampert F, Dear N, Boehm T, Rabbitts TH . HOX11, a homeobox-containing T-cell oncogene on human chromosome 10q24 Proc Natl Acad Sci USA 1991 88: 8900–8904
Avram D, Fields A, Pretty On Top K, Nevrivy DJ, Ishmael JE, Leid M . Isolation of a novel family of C(2)H(2) zinc finger proteins implicated in transcriptional repression mediated by chicken ovalbumin upstream promoter transcription factor (COUP-TF) orphan nuclear receptors J Biol Chem 2000 275: 10315–10322
Satterwhite E, Sonoki T, Willis TG, Harder L, Nowak R, Arriola EL, Liu H, Price HP, Gesk S, Steinemann D, Schlegelberger B, Oscier DG, Siebert R, Tucker PW, Dyer MJ . The BCL11 gene family: involvement of BCL11A in lymphoid malignancies Blood 2001 98: 3413–3420
Breit TM, Beishuizen A, Ludwig WD, Mol EJ, Adriaansen HJ, van Wering ER, van Dongen JJ . tal-1 deletions in T-cell acute lymphoblastic leukemia as PCR target for detection of minimal residual disease Leukemia 1993 7: 2004–2011
Borkhardt A, Repp R, Harbott J, Keller C, Berner F, Ritterbach J, Lampert F . Frequency and DNA sequence of tal-1 rearrangement in children with T-cell acute lymphoblastic leukemia Ann Hematol 1992 64: 305–308
Francois S, Delabesse E, Baranger L, Dautel M, Foussard C, Boasson M, Blanchet O, Bernard O, Macintyre EA, Ifrah N . Deregulated expression of the TAL1 gene by t(1;5)(p32;31) in patient with T-cell acute lymphoblastic leukemia Genes Chromosomes Cancer 1998 23: 36–43
Stock W, Westbrook CA, Sher DA, Dodge R, Sobol RE, Wurster-Hill D, Davey FR, Larson RA, LeBeau MM, Aplan PD, Frankel SR, Stewart CC, Bloomfield CD . Low incidence of TAL1 gene rearrangements in adult acute lymphoblastic leukemia: A cancer and leukemia group B study (8762) Clin Cancer Res 1995 1: 459–463
Cascavilla N, Musto P, D'Arena G, Ladogana S, Matera R, Carotenuto M . Adult and childhood acute lymphoblastic leukemia:clinico–biological differences based on CD34 antigen expression Haematologica 1997 82: 31–37
Thomas X, Archimbaud E, Charrin C, Magaud JP, Fiere D . CD34 expression is associated with major adverse prognostic factors in adult acute lymphoblastic leukemia Leukemia 1995 9: 249–253
Terstappen LW, Huang S, Picker LJ . Flow cytometric assessment of human T-cell differentiation in thymus and bone marrow Blood 1992 79: 666–677
Rowley JD, Reshmi S, Carlson K, Roulston D . Spectral karyotype analysis of T-cell acute leukemia Blood 1999 93: 2038–2042
Ramaswamy S, Tamayo P, Rifkin R, Mukherjee S, Yeang CH, Angelo M, Ladd C, Reich M, Latulippe E, Mesirov JP, Poggio T, Gerald W, Loda M, Lander ES, Golub TR . Multiclass cancer diagnosis using tumor gene expression signatures Proc Natl Acad Sci USA 2001 98: 15149–15154
Getz G, Levine E, Domany E . Coupled two-way clustering analysis of gene microarray data Proc Natl Acad Sci USA 2000 97: 12079–12084
Golub TR, Slonim DK, Tamayo P, Huard C, Gaasenbeek M, Mesirov JP, Coller H, Loh ML, Downing JR, Caligiuri MA, Bloomfield CD, Lander ES . Molecular classification of cancer: class discovery and class prediction by gene expression monitoring Science 1999 286: 531–537
Acknowledgements
We thank J-L Preud'homme for critical reading of the manuscript, M-P Gaub for providing material for B-ALL patients, J-P Bergerat for adult patients material, D Cherif and J Aurich-Costa for providing IPM-FISH chromosomal probes.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Mauvieux, L., Leymarie, V., Helias, C. et al. High incidence of Hox11L2 expression in children with T-ALL. Leukemia 16, 2417–2422 (2002). https://doi.org/10.1038/sj.leu.2402709
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.leu.2402709