Abstract
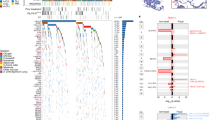
Chronic lymphocytic leukemia (CLL) is a heterogeneous disease with regard to its clinical course. The limitations of the methods currently available for prognostic assessment in CLL do not allow accurate prediction of the risk of disease progression in individual patients. The recently developed cDNA array technique provides a unique opportunity to study gene expression in various malignancies. To identify new molecular markers for prognostication of CLL patients, we analyzed cDNA arrays by using hierarchical clustering and standard statistic t-test on 34 CLL patients. We found significant expression differences in 78 genes compared to the reference tonsillar B lymphocytes. A cluster of genes, LCP1, PARP, BLR1, DEK, NPM, MCL1, SLP76, STAM, HIVEP1, EVI2B, CD25, HTLF, HIVEP2, BCL2, MNDA,PBX3, EBI2, TCF1, CGRP, CD14, IL8,GZMK, GPR17 and CD79B, was associated (P < 0.05) with the unfavorable 11q deletion and also with the unfavorable Binet stages B and C. We present here gene expression profiling that is associated with CLL patients with the 11q23 deletion. Many of the genes in the cluster have not previously been shown to be related to the initiation or progression of CLL. These novel findings provide fundamental information for further attempts to understand the interaction of the clustered genes in the leukomogenesis of CLL in order to better design treatments aimed at specific molecular target(s).
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Binet JL, Auquier A, Dighiero G, Chastang C, Piguet H, Goasguen J, Vaugier G, Potron G, Colona P, Oberling F, Thomas M, Tchernia G, Jacquillat C, Boivin P, Lesty C, Duault MT, Monconduit M, Belabbes S, Gremy F . A new prognostic classification of chronic lymphocytic leukemia derived from a multivariate survival analysis Cancer 1981 48: 198–206
Knuutila S, Elonen E, Teerenhovi L, Rossi L, Leskinen R, Bloomfield CD, de la Chapelle A . Trisomy 12 in B cells of patients with B-cell chronic lymphocytic leukemia N Engl J Med 1986 314: 865–869
Juliusson G, Oscier DG, Fitchett M, Ross FM, Stockdill G, Mackie MJ, Parker AC, Castoldi GL, Guneo A, Knuutila S, Elonen E, Gahrton G . Prognostic subgroups in B cell chronic lymphocytic leukemia defined by specific chromosomal abnormalities N Engl J Med 1990 323: 720–724
Karhu R, Knuutila S, Kallioniemi O-P, Siitonen S, Aine R, Vilpo L, Vilpo J . Frequent loss of the 11q14–24 region in chronic lymphocytic leukemia: a study by comparative genomic hybridization Genes Chromos Cancer 1997 19: 286–290
Larramendy ML, Siitonen SM, Zhu Y, Hurme M, Vilpo L, Vilpo JA, Knuutila S . Optimized mitogen stimulation induces proliferation of neoplastic B cells in chronic lymphocytic leukemia: significance for cytogenetic analysis Cytogenet Cell Genet 1998 82: 215–221
Pérez Losada A, Wessman M, Tiainen M, Hopman AHN, Willard HF, Solé F, Caballín MR, Woessner S, Knuutila S . Trisomy 12 in chronic lymphocytic leukemia: an interphase cytogenetic study Blood 1991 78: 775–779
Dohner H, Stilgenbauer S, Benner A, Leupolt E, Krober A, Bullinger L, Dohner K, Bentz M, Lichter P . Genomic aberrations and survival in chronic lymphocytic leukemia N Engl J Med 2000 343: 1910–1916
Zhu Y, Monni O, El-Rifai W, Siitonen SM, Vilpo L, Vilpo J, Knuutila S . Discontinuous deletions at 11q23 in B cell chronic lymphocytic leukemia Leukemia 1999 13: 708–712
Dohner H, Stilgenbauer S, James MR, Benner A, Weilguni T, Bentz M, Fischer K, Hunstein W, Lichter P . 11q deletions identify a new subset of B-cell chronic lymphocytic leukemia characterized by extensive nodal involvement and inferior prognosis Blood 1997 89: 2516–2522
DeRisi J, Penland L, Brown PO, Bittner ML, Meltzer PS, Ray M, Chen Y, Su YA, Trent JM . Use of a cDNA microarray to analyse gene expression patterns in human cancer Nat Genet 1996 14: 457–460
Golub TR, Slonim DK, Tamayo P, Huard C, Gaasenbeek M, Mesirov JP, Coller H, Loh ML, Downing JR, Caligiuri MA, Bloomfield CD, Lander ES . Molecular classification of cancer: class discovery and class prediction by gene expression monitoring Science 1999 286: 531–537
Rai KR, Sawitsky A, Cronkite EP, Chanana AD, Levy RN, Pasternack BS . Clinical staging of chronic lymphocytic leukemia Blood 1975 46: 219–234
Bennett JM, Catovsky D, Daniel MT, Flandrin G, Galton DA, Gralnick HR, Sultan C . Proposals for the classification of chronic (mature) B and T lymphoid leukaemias. French–American–British (FAB) Cooperative Group J Clin Pathol 1989 42: 567–584
Zhu Y, Monni O, Franssila K, Elonen E, Vilpo J, Joensuu H, Knuutila S . Deletions at 11q23 in different lymphoma subtypes Haematologica 2000 85: 908–912
Takeshita T, Arita T, Higuchi M, Asao H, Endo K, Kuroda H, Tanaka N, Murata K, Ishii N, Sugamura K . STAM, signal transducing adaptor molecule, is associated with Janus kinases and involved in signaling for cell growth and c-myc induction Immunity 1997 6: 449–457
Fu C, Chan AC . Identification of two tyrosine phosphoproteins, pp70 and pp68, which interact with phospholipase Cgamma, Grb2, and Vav after B cell antigen receptor activation J Biol Chem 1997 272: 27362–27368
Pivniouk V, Tsitsikov E, Swinton P, Rathbun G, Alt FW, Geha RS . Impaired viability and profound block in thymocyte development in mice lacking the adaptor protein SLP-76 Cell 1998 94: 229–238
Klein E, Teramoto N, Gogolak P, Nagy N, Bjorkholm M . LMP-1, the Epstein–Barr virus-encoded oncogene with a B cell activating mechanism similar to CD40 Immunol Lett 1999 68: 147–154
Petrella T, Yaziji N, Collin F, Rifle G, Morlevat F, Arnould L, Fargeot P, Depret O . Implication of the Epstein–Barr virus in the progression of chronic lymphocytic leukaemia/small lymphocytic lymphoma to Hodgkin-like lymphomas Anticancer Res 1997 17: 3907–3913
Cawthon RM, Andersen LB, Buchberg AM, Xu GF, O'Connell P, Viskochil D, Weiss RB, Wallace MR, Marchuk DA, Culver M, Stevens J, Jenkins NA, Copeland NG, Collins FS, White R . cDNA sequence and genomic structure of EV12B, a gene lying within an intron of the neurofibromatosis type 1 gene Genomics 1991 9: 446–460
Kaufmann D, Gruener S, Braun F, Stark M, Griesser J, Hoffmeyer S, Bartelt B . EVI2B, a gene lying in an intron of the neurofibromatosis type 1 (NF1) gene, is as the NF1 gene involved in differentiation of melanocytes and keratinocytes and is overexpressed in cells derived from NF1 neurofibromas DNA Cell Biol 1999 18: 345–356
Buchberg AM, Bedigian HG, Jenkins NA, Copeland NG . Evi-2, a common integration site involved in murine myeloid leukemogenesis Mol Cell Biol 1990 10: 4658–4666
Keating MJ . Chronic lymphocytic leukemia Semin Oncol 1999 26: 107–114
Miranda RN, Briggs RC, Shults K, Kinney MC, Jensen RA, Cousar JB . Immunocytochemical analysis of MNDA in tissue sections and sorted normal bone marrow cells documents expression only in maturing normal and neoplastic myelomonocytic cells and a subset of normal and neoplastic B lymphocytes Hum Pathol 1999 30: 1040–1049
Briggs RC, Briggs JA, Ozer J, Sealy L, Dworkin LL, Kingsmore SF, Seldin MF, Kaur GP, Athwal RS, Dessypris EN . The human myeloid cell nuclear differentiation antigen gene is one of at least two related interferon-inducible genes located on chromosome 1q that are expressed specifically in hematopoietic cells Blood 1994 83: 2153–2162
Callea V, Morabito F, Oliva BM, Stelitano C, Levato D, Dattilo A, Gangemi F, Iorfida A, Iacopino P, Nobile F, Molica S, Brugiatelli M . Surface CD14 positivity in B-cell chronic lymphocytic leukaemia is related to clinical outcome Br J Haematol 1999 107: 347–352
McGillis JP, Rangnekar V, Cialella JR . A role for calcitonin gene related peptide (CGRP) in the regulation of early B lymphocyte differentiation Can J Physiol Pharmacol 1995 73: 1057–1064
Fernandez S, Knopf MA, McGillis JP . Calcitonin-gene related peptide (CGRP) inhibits interleukin-7-induced pre-B cell colony formation J Leukoc Biol 2000 67: 669–676
Francia di Celle P, Mariani S, Riera L, Stacchini A, Reato G, Foa R . Interleukin-8 induces the accumulation of B-cell chronic lymphocytic leukemia cells by prolonging survival in an autocrine fashion Blood 1996 87: 4382–4389
Garcia Vela J, Delgado I, Benito L, Monteserin M, Garcia Alonso L, Somolinos N, Andreu M, Ona F . CD79b expression in B cell chronic lymphocytic leukemia: its implication for minimal residual disease detection Leukemia 1999 13: 1501–1505
Forster R, Mattis AE, Kremmer E, Wolf E, Brem G, Lipp M . A putative chemokine receptor, BLR1, directs B cell migration to defined lymphoid organs and specific anatomic compartments of the spleen Cell 1996 87: 1037–1047
Birkenbach M, Josefsen K, Yalamanchili R, Lenoir G, Kieff E . Epstein–Barr virus-induced genes: first lymphocyte-specific G protein-coupled peptide receptors J Virol 1993 67: 2209–2220
Stratowa C, Löffler G, Lichter P, Stilgenbauer S, Haberl P, Schweifer N, Döhner H, Wilgenbus KK . cDNA microarray gene expression analysis of B-cell chronic lymphocytic leukemia proposes potential new prognostic markers involved in lymphocyte trafficking Int J Cancer 2001 91: 474–480
Acknowledgements
This work was supported by grants from the Medical Research Fund of Tampere University Hospital, the Finnish Foundation for Cancer Research, the Sigrid Jusélius Foundation, Paulo Foundation, Finnish Cultural Foundation and Helsinki University Central Hospital Research Funds. We thank the Tampere CLL Group for collaboration, and Leena Pankko and Merja Suoranta for their skilful technical assistance.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Aalto, Y., El-Rifai, W., Vilpo, L. et al. Distinct gene expression profiling in chronic lymphocytic leukemia with 11q23 deletion. Leukemia 15, 1721–1728 (2001). https://doi.org/10.1038/sj.leu.2402282
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.leu.2402282
Keywords
This article is cited by
-
Lck is a relevant target in chronic lymphocytic leukaemia cells whose expression variance is unrelated to disease outcome
Scientific Reports (2017)
-
RelB, together with RelA, sustains cell survival and confers proteasome inhibitor sensitivity of chronic lymphocytic leukemia cells from bone marrow
Journal of Molecular Medicine (2014)
-
Cell surface expression of CD25 antigen (surface IL-2 receptor alpha-chain) is not a prognostic marker in chronic lymphocytic leukemia: results of a retrospective study of 281 patients
Annals of Hematology (2012)
-
Suppression of immune system genes by methylprednisolone in exacerbations of multiple sclerosis
Journal of Neurology (2004)
-
Stimulation-induced gene expression in Ramos B-cells
Genes & Immunity (2003)