Abstract
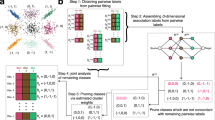
Differential display (DD) is widely employed for identifying uniquely expressed genes within two different cell populations. While potentially powerful, DD is problematic because apparent positive clones require time-consuming verification which may be made even more difficult if only small amounts of starting material are available. We have devised a screening approach to address these issues in primary human hematopoietic cells. Candidate clones are identified in a slot-blot format and verified by 'Virtual Northern' blot analyses using globally amplified cDNA as the verification probe. This method is fast, and since it requires only 0.2 μg of total RNA, it is particularly useful when only limited amounts of study tissue are available.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Divisions of Hematology/Oncology, Department of Medicine and Hematology/Molecular Diagnosis, University of Pennsylvania School of Medicine, Philadelphia, PA, USA
Rights and permissions
About this article
Cite this article
Hung, HL., Song, F. & Gewirtz, A. A method for identifying differentially expressed genes in rare populations of primary human hematopoietic cells. Leukemia 13, 295–297 (1999). https://doi.org/10.1038/sj.leu.2401274
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.leu.2401274
Keywords
This article is cited by
-
Seeking for Senile-Related Gene Expression in Cerebral Tissue of Senescence-Accelerated Mouse
Cellular and Molecular Neurobiology (2004)