Abstract
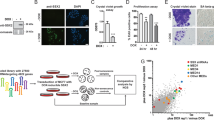
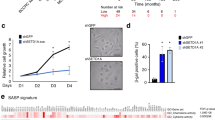
Senescence is a mechanism that limits cellular lifespan and constitutes a barrier against cellular immortalization. To identify new senescence regulatory genes that might play a role in tumorigenesis, we have designed and performed a large-scale antisense-based genetic screen in primary mouse embryo fibroblasts (MEFs). Out of this screen, we have identified five different genes through which loss of function partially bypasses senescence. These genes belong to very different biochemical families: csn2 (component of the Cop9 signalosome), aldose reductase (a metabolic enzyme) and brf1 (subunit of the RNA polymerase II complex), S-adenosyl homocysteine hydrolase and Bub1. Inactivation, at least partial, of these genes confers resistance to both p53- and p16INK4a-induced proliferation arrest. Furthermore, such inactivation inhibits p53 but not E2F1 transcriptional activity and impairs DNA-damage-induced transcription of p21. Since the aim of the screen was to identify new regulators of tumorigenesis, we have tested their inactivation in human tumors. We have found, either by northern blot or quantitative reverse transcriptase–PCR analysis, that the expression of three genes, Csn2, Aldose reductase and Brf1, is lost at different ratios in tumors of different origins. These genes are located at common positions of loss of heterogeneity (15q21.2, 7q35 and 14q32.33); therefore,we have measured genomic losses of these specific genes in different tumors. We have found that Csn2 and Brf1 also show genomic losses of one allele in different tumors. Our data suggest that the three genes identified in the genome-wide loss-of-function genetic screen are putative tumor suppressors located at 15q21.2; 7q35 and 14q32.33.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Bech-Otschir D, Kraft R, Huang X, Henklein P, Kapelari B, Pollmann C et al. (2001). COP9 signalosome-specific phosphorylation targets p53 to degradation by the ubiquitin system. Embo J 20: 1630–1639.
Bech-Otschir D, Seeger M, Dubiel W . (2002). The COP9 signalosome: at the interface between signal transduction and ubiquitin-dependent proteolysis. J Cell Sci 115: 467–473.
Berns K, Hijmans E M, Mullenders J, Brummelkamp TR, Velds A, Heimerikx M et al. (2004). A large-scale RNAi screen in human cells identifies new components of the p53 pathway. Nature 428: 431–437.
Braig M, Lee S, Loddenkemper C, Rudolph C, Peters AH, Schlegelberger B et al. (2005). Oncogene-induced senescence as an initial barrier in lymphoma development. Nature 436: 660–665.
Campisi J . (2001). Cellular senescence as a tumor-suppressor mechanism. Trends Cell Biol 11: S27–S31.
Campisi J . (2005). Suppressing cancer: the importance of being senescent. Science 309: 886–887.
Carnero A, Beach DH . (2004). Absence of p21WAF1 cooperates with c-myc in bypassing Ras-induced senescence and enhances oncogenic cooperation. Oncogene 23: 6006–6011.
Carnero A, Hudson JD, Hannon GJ, Beach DH . (2000). Loss-of-function genetics in mammalian cells: the p53 tumor suppressor model. Nucleic Acids Res 28: 2234–2241.
Collado M, Gil J, Efeyan A, Guerra C, Schuhmacher AJ, Barradas M et al. (2005). Tumour biology: senescence in premalignant tumours. Nature 436: 642.
Dimri GP, Lee X, Basile G, Acosta M, Scott G, Roskelley C et al. (1995). A biomarker that identifies senescent human cells in culture and in aging skin in vivo. Proc Natl Acad Sci USA 92: 9363–9367.
Eichhorn K, Jackson SP . (2001). A role for TAF3B2 in the repression of human RNA polymerase III transcription in nonproliferating cells. J Biol Chem 276: 21158–21165.
Greider CW, Blackburn EH . (2004). Tracking telomerase. Cell 116: S83–S86, 1 p following S86.
Hanahan D, Weinberg RA . (2000). The hallmarks of cancer. Cell 100: 57–70.
Larminie CG, Cairns CA, Mital R, Martin K, Kouzarides T, Jackson S P et al. (1997). Mechanistic analysis of RNA polymerase III regulation by the retinoblastoma protein. Embo J 16: 2061–2071.
Lefrancois-Martinez AM, Bertherat J, Val P, Tournaire C, Gallo-Payet N et al. (2004). Decreased expression of cyclic adenosine monophosphate-regulated aldose reductase (AKR1B1) is associated with malignancy in human sporadic adrenocortical tumors. J Clin Endocrinol Metab 89: 3010–3019.
Lleonart ME, Vidal F, Gallardo D, Diaz-Fuertes M, Rojo F, Cuatrecasas M et al. (2006). New p53 related genes in human tumors: significant downregulation in colon and lung carcinomas. Oncol Rep 16: 603–608.
Lykke-Andersen K, Schaefer L, Menon S, Deng XW, Miller J B, Wei N . (2003). Disruption of the COP9 signalosome Csn2 subunit in mice causes deficient cell proliferation, accumulation of p53 and cyclin E, and early embryonic death. Mol Cell Biol 23: 6790–6797.
Michaloglou C, Vredeveld LC, Soengas MS, Denoyelle C, Kuilman T, van der Horst CM et al. (2005). BRAFE600-associated senescence-like cell cycle arrest of human naevi. Nature 436: 720–724.
Morales CP, Holt SE, Ouellette M, Kaur KJ, Yan Y, Wilson KS et al. (1999). Absence of cancer-associated changes in human fibroblasts immortalized with telomerase. Nat Genet 21: 115–118.
Sage J, Mulligan GJ, Attardi LD, Miller A, Chen S, Williams B et al. (2000). Targeted disruption of the three Rb-related genes leads to loss of G(1) control and immortalization. Genes Dev 14: 3037–3050.
Schmitt CA, Fridman JS, Yang M, Lee S, Baranov E, Hoffman RM et al. (2002). A senescence program controlled by p53 and p16INK4a contributes to the outcome of cancer therapy. Cell 109: 335–346.
Schouten JP, McElgunn CJ, Waaijer R, Zwijnenburg D, Diepvens F, Pals G . (2002). Relative quantification of 40 nucleic acid sequences by multiplex ligation-dependent probe amplification. Nucleic Acids Res 30: e57.
Serrano M, Blasco MA . (2001). Putting the stress on senescence. Curr Opin Cell Biol 13: 748–753.
Serrano M, Lin W, McCurrach ME, Beach D, Lowe SW . (1997). Oncogenic ras provokes premature cell senescence associated with accumulation of p53 and p16INK4a. Cell 88: 593–602.
Shay JW, Roninson IB . (2004). Hallmarks of senescence in carcinogenesis and cancer therapy. Oncogene 23: 2919–2933.
Sherr CJ, McCormick F . (2002). The RB and p53 pathways in cancer. Cancer Cell 2: 103–112.
Suzuki K, Mori I, Nakayama Y, Miyakoda M, Kodama S, Watanabe M . (2001). Radiation-induced senescence-like growth arrest requires TP53 function but not telomere shortening. Radiat Res 155: 248–253.
Wei N, Tsuge T, Serino G, Dohmae N, Takio K, Matsui M et al. (1998). The COP9 complex is conserved between plants and mammals and is related to the 26S proteasome regulatory complex. Curr Biol 8: 919–922.
Yamasaki L, Bronson R, Williams BO, Dyson N J, Harlow E, Jacks T . (1998). Loss of E2F-1 reduces tumorigenesis and extends the lifespan of Rb1(+/-)mice. Nat Genet 18: 360–364.
Yang X, Menon S, Lykke-Andersen K, Tsuge T, Di X, Wang X et al. (2002). The COP9 signalosome inhibits p27(kip1) degradation and impedes G1-S phase progression via deneddylation of SCF Cul1. Curr Biol 12: 667–672.
Yoneda-Kato N, Tomoda K, Umehara M, Arata Y, Kato JY . (2005). Myeloid leukemia factor 1 regulates p53 by suppressing COP1 via COP9 signalosome subunit 3. Embo J 24: 1739–1749.
Acknowledgements
We thank the CNIO Tumor Bank for providing the tumor samples. This work has been partly supported by grants from Spanish Ministry of Health (FIS-02/0126), Fundación Mutua Madrileña and the Spanish Ministry of Education and Science (SAF2005-00944) to AC and MRC and Wellcome Trust (to DHB).
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Oncogene website (http://www.nature.com/onc).
Rights and permissions
About this article
Cite this article
Leal, J., Fominaya, J., Cascón, A. et al. Cellular senescence bypass screen identifies new putative tumor suppressor genes. Oncogene 27, 1961–1970 (2008). https://doi.org/10.1038/sj.onc.1210846
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1210846
Keywords
This article is cited by
-
Role of NEDD8 and neddylation dynamics in DNA damage response
Genome Instability & Disease (2021)
-
Levels of active tyrosine kinase receptor determine the tumor response to Zalypsis
BMC Cancer (2014)
-
SIRT7 links H3K18 deacetylation to maintenance of oncogenic transformation
Nature (2012)
-
New insights into p53 activation
Cell Research (2010)
-
Bypassing cellular senescence by genetic screening tools
Clinical and Translational Oncology (2010)