Abstract
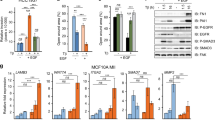
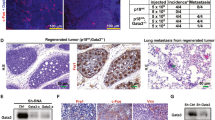
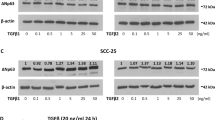
We report here that human MFGE8 encoding milk fat globule-EGF factor 8 protein (MFG-E8), also termed 46 kDa breast epithelial antigen and lactadherin, is transcriptionally activated by p63, or TP63, a p53 (TP53) family protein frequently overexpressed in head-and-neck squamous cell carcinomas, mammary carcinomas and so on. Despite that human MFG-E8 was originally identified as a breast cancer marker, and has recently been reported to provide peptides for cancer immunotherapy, its transcriptional control remains an open question. Observations in immunohistochemical analyses, a tetracycline-induced p63 expression system and keratinocyte cultures suggested a physiological link between p63 and MFGE8. By reporter assays with immediately upstream regions of MFGE8, we determined that the trans-activator (TA) isoforms of p63 activate MFGE8 transcription though a p53/p63 motif at −370, which was confirmed by a chromatin immunoprecipitation experiment. Upon siRNA-mediated p63 silencing in a squamous cell carinoma line, MFG-E8 production decreased to diminish Saos-2 cell adhesion. Interestingly, the ΔN-p63 isoform lacking the TA domain enhanced the MFGE8-activating function of TA-p63, if ΔN-p63 was dominant over TA-p63 as typically observed in undifferentiated keratinocytes and squamous cell carcinomas, implying a self-regulatory mechanism of p63 by the TA:ΔN association. MFG-E8 may provide a novel pathway of epithelial–nonepithelial cell interactions inducible by p63, probably in pathological processes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Brunner HG, Hamel BCJ, van Bokhoven H . (2002). The p63 gene in EEC and other syndromes. J Med Genet 39: 377–381.
Carmon L, Bobilev-Priel I, Brenner B, Bobilev D, Paz A, Bar-Haim E et al. (2002). Characterization of novel breast carcinoma-associated BA46-derived peptides in HLA-A2.1/D(b)-beta2m transgenic mice. J Clin Invest 110: 453–462.
Carroll DK, Carroll JS, Leong CO, Cheng F, Brown M, Mills AA et al. (2006). p63 regulates an adhesion programme and cell survival in epithelial cells. Nat Cell Biol 8: 551–561.
Celli J, Duijf P, Hamel BC, Bamshad M, Kramer B, Smits AP et al. (1999). Heterozygous germline mutations in the p53 homolog p63 are the cause of EEC syndrome. Cell 99: 143–153.
DiRenzo J, Signoretti S, Nakamura N, Rivera-Gonzalez R, Sellers W, Loda M et al. (2002). Growth factor requirements and basal phenotype of an immortalized mammary epithelial cell line. Cancer Res 62: 89–98.
el-Deiry WS, Kern SE, Pietenpol JA, Kinzler KW, Vogelstein B . (1992). Definition of a consensus binding site for p53. Nat Genet 1: 45–49.
Ensslin MA, Shur BD . (2003). Identification of mouse sperm SED1, a bimotif EGF repeat and discoidin-domain protein involved in sperm-egg binding. Cell 114: 405–417.
Gailit J, Clarke C, Newman D, Tonnesen MG, Mosesson MW, Clark RAF . (1997). Human fibroblasts bind directly to fibrinogen at RGD sites through integrin [alpha]v[beta]3. Exp Cell Res 232: 118–126.
Hanayama R, Nagata S . (2005). Impaired involution of mammary glands in the absence of milk fat globule EGF factor 8. Proc Natl Acad Sci USA 102: 16886–16891.
Hanayama R, Tanaka M, Miwa K, Shinohara A, Iwamatsu A, Nagata S . (2002). Identification of a factor that links apoptotic cells to phagocytes. Nature 417: 182–187.
Hanayama R, Tanaka M, Miyasaka K, Aozasa K, Koike M, Uchiyama Y et al. (2004). Autoimmune disease and impaired uptake of apoptotic cells in MFG-E8-deficient mice. Science 304: 1147–1150.
Helton ES, Zhu J, Chen X . (2006). The unique NH2-terminally deleted (DeltaN) residues, the PXXP motif, and the PPXY motif are required for the transcriptional activity of the DeltaN variant of p63. J Biol Chem 281: 2533–2542.
Hibi K, Trink B, Patturajan M, Westra WH, Caballero OL, Hill DE et al. (2000). AIS is an oncogene amplified in squamous cell carcinoma. Proc Natl Acad Sci 97: 5462–5467.
Hoffmann S, He S, Jin M, Ehren M, Wiedemann P, Ryan S et al. (2005). A selective cyclic integrin antagonist blocks the integrin receptors alphavbeta3 and alphavbeta5 and inhibits retinal pigment epithelium cell attachment, migration and invasion. BMC Ophthalmol 5: 16.
Hvarregaard J, Andersen MH, Berglund L, Rasmussen JT, Petersen TE . (1996). Characterization of glycoprotein PAS-6/7 from membranes of bovine milk fat globules. Eur J Biochem 240: 628–636.
Ihrie RA, Marques MR, Nguyen BT, Horner JS, Papazoglu C, Bronson RT et al. (2005). Perp is a p63-regulated gene essential for epithelial integrity. Cell 120: 843–856.
Koga F, Kawakami S, Kumagai J, Takizawa T, Ando N, Arai G et al. (2003). Impaired delta Np63 expression associates with reduced beta-catenin and aggressive phenotypes of urothelial neoplasms. Br J Cancer 88: 740–747.
Koistinen P, Pulli T, Uitto V-J, Nissinen L, Hyypia T, Heino J . (1999). Depletion of [alpha]V integrins from osteosarcoma cells by intracellular antibody expression induces bone differentiation marker genes and suppresses gelatinase (MMP-2) synthesis. Matrix Biol 18: 239–251.
Kojima T, Ikawa Y, Katoh I . (2001). Analysis of molecular interactions of the p53-family p51(p63) gene products in a yeast two-hybrid system: homotypic and heterotypic interactions and association with p53-regulatory factors. Biochem Biophys Res Commun 281: 1170–1175.
Kruger W, Lohner R, Jung R, Kroger N, Zander AR . (2000). Expression of human milk fat globulin proteins in cells of haemopoietic origin. Br J Cancer 83: 874–879.
Kurata S, Okuyama T, Osada M, Watanabe T, Tomimori Y, Sato S et al. (2004). p51/p63 Controls subunit alpha3 of the major epidermis integrin anchoring the stem cells to the niche. J Biol Chem 279: 50069–50077.
Larocca D, Peterson JA, Urrea R, Kuniyoshi J, Bistrain AM, Ceriani RL . (1991). A Mr 46,000 human milk fat globule protein that is highly expressed in human breast tumors contains factor VIII-like domains. Cancer Res 51: 4994–4998.
Li ER, Owens DM, Djian P, Watt FM . (2000). Expression of involucrin in normal, hyperproliferative and neoplastic mouse keratinocytes. Exp Dermatol 9: 431–438.
Liu Y, Chiriva-Internati M, You C, Luo R, You H, Prasad CK et al. (2005). Use and specificity of breast cancer antigen/milk protein BA46 for generating anti-self-cytotoxic T lymphocytes by recombinant adeno-associated virus-based gene loading of dendritic cells. Cancer Gene Ther 12: 304–312.
Massion PP, Taflan PM, Jamshedur Rahman SM, Yildiz P, Shyr Y, Edgerton ME et al. (2003). Significance of p63 amplification and Ooerexpression in lung cancer development and prognosis. Cancer Res 63: 7113–7121.
Mather IH, Banghart LR, Lane WS . (1993). The major fat-globule membrane proteins, bovine components 15/16 and guinea-pig GP 55, are homologous to MGF-E8, a murine glycoprotein containing epidermal growth factor-like and factor V/VIII-like sequences. Biochem Mol Biol Int 29: 545–554.
Mills AA, Zheng B, Wang XJ, Vogel H, Roop DR, Bradley A . (1999). p63 is a p53 homologue required for limb and epidermal morphogenesis. Nature 398: 708–713.
Okada Y, Osada M, Kurata S, Sato S, Aisaki K, Kageyama Y et al. (2002). p53 gene family p51(p63)-encoded, secondary transactivator p51B(TAp63alpha) occurs without forming an immunoprecipitable complex with MDM2, but responds to genotoxic stress by accumulation. Exp Cell Res 276: 194–200.
Osada M, Inaba R, Shinohara H, Hagiwara M, Nakamura M, Ikawa Y . (2001). Regulatory domain of protein stability of human P51/TAP63, a P53 homologue. Biochem Biophys Res Commun 283: 1135–1141.
Osada M, Nagakawa Y, Park HL, Yamashita K, Wu G, Kim MS et al. (2005a). p63-specific activation of the BPAG-1e promoter. J Invest Dermatol 125: 52–60.
Osada M, Ohba M, Kawahara C, Ishioka C, Kanamaru R, Katoh I et al. (1998). Cloning and functional analysis of human p51, which structurally and functionally resembles p53. Nat Med 4: 839–843.
Osada M, Park HL, Nagakawa Y, Yamashita K, Fomenkov A, Kim MS et al. (2005b). Differential recognition of response elements determines target gene specificity for p53 and p63. Mol Cell Biol 25: 6077–6089.
Oshima K, Aoki N, Kato T, Kitajima K, Matsuda T . (2002). Secretion of a peripheral membrane protein, MFG-E8, as a complex with membrane vesicles. Eur J Biochem 269: 1209–1218.
Pellegrini G, Dellambra E, Golisano O, Martinelli E, Fantozzi I, Bondanza S et al. (2001). p63 identifies keratinocyte stem cells. Proc Natl Acad Sci USA 98: 3156–3161.
Peterson JA, Zava DT, Duwe AK, Blank EW, Battifora H, Ceriani RL . (1990). Biochemical and histological characterization of antigens preferentially expressed on the surface and cytoplasm of breast carcinoma cells identified by monoclonal antibodies against the human milk fat globule. Hybridoma 9: 221–235.
Quade BJ, Yang A, Wang Y, Sun D, Park J, Sheets EE et al. (2001). Expression of the p53 homologue p63 in early cervical neoplasia. Gynecol Oncol 80: 24–29.
Rossi M, Aqeilan RI, Neale M, Candi E, Salomoni P, Knight RA et al. (2006). The E3 ubiquitin ligase Itch controls the protein stability of p63. Proc Natl Acad Sci USA 103: 12753–12758.
Shi J, Gibert GE . (2003). Lactadherin inhibits enzyme complexes of blood coagulation by completing for phospholipid-binding sites. Blood 101: 2628–2636.
Silvestre JS, Thery C, Hamard G, Boddaert J, Aguilar B, Delcayre A et al. (2005). Lactadherin promotes VEGF-dependent neovascularization. Nat Med 11: 499–506.
Smith RA, Giorgio TD . (2004). Quantitation and kinetics of CD51 surface receptor expression: implications for targeted delivery. Ann Biomed Eng 32: 635–644.
Taylor MR, Couto JR, Scallan CD, Ceriani RL, Peterson JA . (1997). Lactadherin (formerly BA46), a membrane-associated glycoprotein expressed in human milk and breast carcinomas, promotes Arg-Gly-Asp (RGD)-dependent cell adhesion. DNA Cell Biol 16: 861–869.
Watanabe T, Totsuka R, Miyatani S, Kurata S, Sato S, Katoh I et al. (2005). Production of the long and short forms of MFG-E8 by epidermal keratinocytes. Cell Tissue Res 321: 185–193.
Wu G, Osada M, Guo Z, Fomenkov A, Begum S, Zhao M et al. (2005). DeltaNp63alpha up-regulates the Hsp70 gene in human cancer. Cancer Res 65: 758–766.
Yamaguchi K, Wu L, Caballero OL, Hibi K, Trink B, Resto V et al. (2000). Frequent gain of the p40/p51/p63 gene locus in primary head and neck squamous cell carcinoma. Int J Cancer 86: 684–689.
Yang A, Kaghad M, Wang Y, Gillett E, Fleming MD, Dotsch V et al. (1998). p63, a p53 homolog at 3q27-29, encodes multiple products with transactivating, death-inducing, and dominant-negative activities. Mol Cell 2: 305–316.
Yang A, Schweitzer R, Sun D, Kaghad M, Walker N, Bronson RT et al. (1999). p63 is essential for regenerative proliferation in limb, craniofacial and epithelial development. Nature 398: 714–718.
Acknowledgements
This work was supported by Grant-in Aids from MEXT Japan, for Scientific Research on Priority Areas (YI) and for High-Tech Research Center Project (RH) and by Strategic Research Fund from University of Yamanashi (IK). We thank N Shirato, Photography Department, Tokyo Medical and Dental University, for data file making.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Oncogene web site (http://www.nature.com/onc).
Supplementary information
Rights and permissions
About this article
Cite this article
Okuyama, T., Kurata, S., Tomimori, Y. et al. p63(TP63) elicits strong trans-activation of the MFG-E8/lactadherin/BA46 gene through interactions between the TA and ΔN isoforms. Oncogene 27, 308–317 (2008). https://doi.org/10.1038/sj.onc.1210646
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1210646
Keywords
This article is cited by
-
Milk fat globule epithelial growth factor VIII (MFG-E8) sustains survival of cancer cells by prompting tumor angiogenesis and suppressing host immunities ⁎
Oncology and Translational Medicine (2017)
-
Role of milk fat globule-epidermal growth factor 8 in colonic inflammation and carcinogenesis
Journal of Gastroenterology (2015)