Abstract
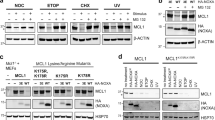
The alternative reading frame (ARF) mRNA encodes two pro-death proteins, the nucleolar p19ARF and a shorter mitochondrial isoform, named smARF (hsmARF in human). While p19ARF can inhibit cell growth by causing cell cycle arrest or type I apoptotic cell death, smARF is able to induce type II autophagic cell death. Inappropriate proliferative signals generated by proto-oncogenes, such as c-Myc and E2F1, can elevate both p19ARF and smARF proteins. Here, we reveal a novel means of regulation of smARF protein steady state levels through its interactions with the mitochondrial p32. The p32 protein physically interacts with both human and murine smARF, and colocalizes with these short isoforms to the mitochondria. Remarkably, knocking down p32 protein levels significantly reduced the steady state levels of smARF by increasing its turn over. As a consequence, the ability of ectopically expressed smARF to induce autophagy and to cause mitochondrial membrane dissipation was significantly reduced. In contrast, the protein levels of full-length p19ARF, which mainly resides in the nucleolus, were not influenced by p32 depletion, suggesting that p32 exclusively stabilizes the mitochondrial smARF protein. Thus the interaction with p32 provides a means of specifically regulating the expression of the recently identified autophagic inducer, smARF, and adds yet another layer of complexity to the multifaceted regulation of the ARF gene.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Brokstad KA, Kalland KH, Russell WC, Matthews DA . (2001). Mitochondrial protein p32 can accumulate in the nucleus. Biochem Biophys Res Commun 281: 1161–1169.
Bruni R, Roizman B . (1996). Open reading frame P – a herpes simplex virus gene repressed during productive infection encodes a protein that binds a splicing factor and reduces synthesis of viral proteins made from spliced mRNA. Proc Natl Acad Sci USA 93: 10423–10427.
Deb TB, Datta K . (1996). Molecular cloning of human fibroblast hyaluronic acid-binding protein confirms its identity with P-32, a protein co-purified with splicing factor SF2. Hyaluronic acid-binding protein as P-32 protein, co-purified with splicing factor SF2. J Biol Chem 271: 2206–2212.
Dedio J, Jahnen-Dechent W, Bachmann M, Muller-Esterl W . (1998). The multiligand-binding protein gC1qR, putative C1q receptor, is a mitochondrial protein. J Immunol 160: 3534–3542.
Desai K, Loewenstein PM, Green M . (1991). Isolation of a cellular protein that binds to the human immunodeficiency virus Tat protein and can potentiate transactivation of the viral promoter. Proc Natl Acad Sci USA 88: 8875–8879.
Fridell RA, Harding LS, Bogerd HP, Cullen BR . (1995). Identification of a novel human zinc finger protein that specifically interacts with the activation domain of lentiviral Tat proteins. Virology 209: 347–357.
Ghebrehiwet B, Lim BL, Peerschke EI, Willis AC, Reid KB . (1994). Isolation, cDNA cloning, and overexpression of a 33-kD cell surface glycoprotein that binds to the globular ‘heads’ of C1q. J Exp Med 179: 1809–1821.
Granot Z, Geiss-Friedlander R, Melamed-Book N, Eimerl S, Timberg R, Weiss AM et al. (2003). Proteolysis of normal and mutated steroidogenic acute regulatory proteins in the mitochondria: the fate of unwanted proteins. Mol Endocrinol 17: 2461–2476.
Herwald H, Dedio J, Kellner R, Loos M, Muller-Esterl W . (1996). Isolation and characterization of the kininogen-binding protein p33 from endothelial cells. Identity with the gC1q receptor. J Biol Chem 271: 13040–13047.
Jiang J, Zhang Y, Krainer AR, Xu RM . (1999). Crystal structure of human p32, a doughnut-shaped acidic mitochondrial matrix protein. Proc Natl Acad Sci USA 96: 3572–3577.
Kuo ML, den Besten W, Bertwistle D, Roussel MF, Sherr CJ . (2004). N-terminal polyubiquitination and degradation of the ARF tumor suppressor. Genes Dev 18: 1862–1874.
Lim BL, Reid KB, Ghebrehiwet B, Peerschke EI, Leigh LA, Preissner KT . (1996). The binding protein for globular heads of complement C1q, gC1qR. Functional expression and characterization as a novel vitronectin binding factor. J Biol Chem 271: 26739–26744.
Luo Y, Yu H, Peterlin BM . (1994). Cellular protein modulates effects of human immunodeficiency virus type 1 Rev. J Virol 68: 3850–3856.
Matthews DA, Russell WC . (1998). Adenovirus core protein V interacts with p32--a protein which is associated with both the mitochondria and the nucleus. J Gen Virol 79 (Part 7): 1677–1685.
Muta T, Kang D, Kitajima S, Fujiwara T, Hamasaki N . (1997). p32 protein, a splicing factor 2-associated protein, is localized in mitochondrial matrix and is functionally important in maintaining oxidative phosphorylation. J Biol Chem 272: 24363–24370.
Reef S, Zalckvar E, Shifman O, Bialik S, Sabanay H, Oren M et al. (2006). A short mitochondrial form of p19ARF induces autophagy and caspase-independent cell death. Mol Cell 22: 463–475.
Seytter T, Lottspeich F, Neupert W, Schwarz E . (1998). Mam33p, an oligomeric, acidic protein in the mitochondrial matrix of Saccharomyces cerevisiae is related to the human complement receptor gC1q-R. Yeast 14: 303–310.
Sherr CJ . (2006). Divorcing ARF and p53: an unsettled case. Nat Rev Cancer 6: 663–673.
Simos G, Georgatos SD . (1994). The lamin B receptor-associated protein p34 shares sequence homology and antigenic determinants with the splicing factor 2-associated protein p32. FEBS Lett 346: 225–228.
Soltys BJ, Kang D, Gupta RS . (2000). Localization of P32 protein (gC1q-R) in mitochondria and at specific extramitochondrial locations in normal tissues. Histochem Cell Biol 114: 245–255.
Sunayama J, Ando Y, Itoh N, Tomiyama A, Sakurada K, Sugiyama A . (2004). Physical and functional interaction between BH3-only protein Hrk and mitochondrial pore-forming protein p32. Cell Death Differ 11: 771–781.
Tange TO, Jensen TH, Kjems J . (1996). In vitro interaction between human immunodeficiency virus type 1 Rev protein and splicing factor ASF/SF2-associated protein, p32. J Biol Chem 271: 10066–10072.
Wang Y, Finan JE, Middeldorp JM, Hayward SD . (1997). P32/TAP, a cellular protein that interacts with EBNA-1 of Epstein–Barr virus. Virology 236: 18–29.
Yu L, Loewenstein PM, Zhang Z, Green M . (1995a). In vitro interaction of the human immunodeficiency virus type 1 Tat transactivator and the general transcription factor TFIIB with the cellular protein TAP. J Virol 69: 3017–3023.
Yu L, Zhang Z, Loewenstein PM, Desai K, Tang Q, Mao D et al. (1995b). Molecular cloning and characterization of a cellular protein that interacts with the human immunodeficiency virus type 1 Tat transactivator and encodes a strong transcriptional activation domain. J Virol 69: 3007–3016.
Acknowledgements
We thank WC Russell for providing the polyclonal antibody against p32. We thank S Bialik for critical reading of the article, E Zalckvar and G Tarcic for help and advice. This work was supported by grants from the European Union (LSHB-CT-2004-511983) (to AK) and by the Center of Excellence grant from the Flight Attendant Medical Research Institute (FAMRI) to AK and MO. AK is the incumbent of Helena Rubinstein Chair of Cancer Research.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Reef, S., Shifman, O., Oren, M. et al. The autophagic inducer smARF interacts with and is stabilized by the mitochondrial p32 protein. Oncogene 26, 6677–6683 (2007). https://doi.org/10.1038/sj.onc.1210485
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1210485
Keywords
This article is cited by
-
An approach to p32/gC1qR/HABP1: a multifunctional protein with an essential role in cancer
Journal of Cancer Research and Clinical Oncology (2022)
-
Elevated expression of HABP1 is a novel prognostic indicator in triple-negative breast cancers
Tumor Biology (2015)
-
A short acidic motif in ARF guards against mitochondrial dysfunction and melanoma susceptibility
Nature Communications (2014)
-
Elevated expression of hyaluronic acid binding protein 1 (HABP1)/P32/C1QBP is a novel indicator for lymph node and peritoneal metastasis of epithelial ovarian cancer patients
Tumor Biology (2013)
-
The autophagic paradox in cancer therapy
Oncogene (2012)