Abstract
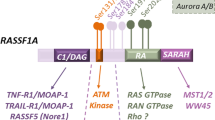
RASSF family proteins are tumor suppressors that are frequently downregulated during the development of human cancer. The best-characterized member of the family is RASSF1A, which is downregulated by promoter methylation in 40–90% of primary human tumors. We now identify and characterize a novel member of the RASSF family, RASSF6. Like the other family members, RASSF6 possesses a Ras Association domain and binds activated Ras. Exogenous expression of RASSF6 promoted apoptosis, synergized with activated K-Ras to induce cell death and inhibited the survival of specific tumor cell lines. Suppression of RASSF6 enhanced the tumorigenic phenotype of a human lung tumor cell line. Furthermore, RASSF6 is often downregulated in primary human tumors. RASSF6 shares some similar overall properties as other RASSF proteins. However, there are significant differences in biological activity between RASSF6 and other family members including a discrete tissue expression profile, cell killing specificity and impact on signaling pathways. Moreover, RASSF6 may play a role in dictating the degree of inflammatory response to the respiratory syncytial virus. Thus, RASSF6 is a novel RASSF family member that demonstrates the properties of a Ras effector and tumor suppressor but exhibits biological properties that are unique and distinct from those of other family members.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout









Similar content being viewed by others
Abbreviations
- GST:
-
glutathione S transferase
- RA:
-
Ras association
- RSV:
-
respiratory syncytial virus
References
Agathanggelou A, Cooper WN, Latif F . (2005). Role of the Ras-association domain family 1 tumor suppressor gene in human cancers. Cancer Res 65: 3497–3508.
Akino K, Toyota M, Suzuki H, Mita H, Sasaki Y, Ohe-Toyota M et al. (2005). The Ras effector RASSF2 is a novel tumor-suppressor gene in human colorectal cancer. Gastroenterology 129: 156–169.
Baksh S, Tommasi S, Fenton S, Yu VC, Martins LM, Pfeifer GP et al. (2005). The tumor suppressor RASSF1A and MAP-1 link death receptor signaling to Bax conformational change and cell death. Mol Cell 18: 637–650.
Bitko V, Garmon NE, Cao T, Estrada B, Oakes JE, Lausch RN et al. (2004). Activation of cytokines and NF-kappa B in corneal epithelial cells infected by respiratory syncytial virus: potential relevance in ocular inflammation and respiratory infection. BMC Microbiol 4: 28.
Burbee DG, Forgacs E, Zochbauer-Muller S, Shivakumar L, Fong K, Gao B et al. (2001). Epigenetic inactivation of RASSF1A in lung and breast cancers and malignant phenotype suppression. J Natl Cancer Inst 93: 691–699.
Chen J, Lui WO, Vos MD, Clark GJ, Takahashi M, Schoumans J et al. (2003). The t(1;3) breakpoint-spanning genes LSAMP and NORE1 are involved in clear cell renal cell carcinomas. Cancer Cell 4: 405–413.
Clark GJ, Drugan JK, Terrell RS, Bradham C, Der CJ, Bell RM et al. (1996). Peptides containing a consensus Ras binding sequence from Raf-1 and theGTPase activating protein NF1 inhibit Ras function. Proc Natl Acad Sci USA 93: 1577–1581.
Cox AD, Der CJ . (2003). The dark side of Ras: regulation of apoptosis. Oncogene 22: 8999–9006.
Dammann R, Li C, Yoon JH, Chin PL, Bates S, Pfeifer GP . (2000). Epigenetic inactivation of a RAS association domain family protein from the lung tumour suppressor locus 3p21.3. Nat Genet 25: 315–319.
Diep CB, Teixeira MR, Thorstensen L, Wiig JN, Eknaes M, Nesland JM et al. (2004). Genome characteristics of primary carcinomas, local recurrences, carcinomatoses, and liver metastases from colorectal cancer patients. Mol Cancer 3: 6.
Eckfeld K, Hesson L, Vos MD, Bieche I, Latif F, Clark GJ . (2004). RASSF4/AD037 is a potential ras effector/tumor suppressor of the RASSF family. Cancer Res 64: 8688–8693.
Ellis CA, Vos MD, Howell H, Vallecorsa T, Fults DW, Clark GJ . (2002). Rig is a novel Ras-related protein and potential neural tumor suppressor. Proc Natl Acad Sci USA 99: 9876–9881.
Feig LA, Buchsbaum RJ . (2002). Cell signaling: life or death decisions of ras proteins. Curr Biol 12: R259–R261.
Fiordalisi JJ, Johnson RL 2nd, Ulku AS, Der CJ, Cox AD . (2001). Mammalian expression vectors for Ras family proteins: generation and use of expression constructs to analyze Ras family functions. Meth Enzymology 332: 3–36.
Frame S, Balmain A . (2000). Integration of positive and negative growth signals during ras pathway activation in vivo. Curr Opin Genet Dev 10: 106–113.
Hueber AO, Evan GI . (1998). Traps to catch unwary oncogenes. Trends Genet 14: 364–367.
Hull J, Rowlands K, Lockhart E, Sharland M, Moore C, Hanchard N et al. (2004). Haplotype mapping of the bronchiolitis susceptibility locus near IL8. Hum Genet 114: 272–279.
Khokhlatchev A, Rabizadeh S, Xavier R, Nedwidek M, Chen T, Zhang XF et al. (2002). Identification of a novel Ras-regulated proapoptotic pathway. Curr Biol 12: 253–265.
Malumbres M, Barbacid M . (2003). RAS oncogenes: the first 30 years. Nat Rev Cancer 3: 459–465.
Malumbres M, Pellicer A . (1998). RAS pathways to cell cycle control and cell transformation. Front Biosci 3: d887–d912.
Marshall MS . (1993). The effector interactions of p21ras. Trends Biochem Sci 18: 250–254.
Mayo MW, Wang CY, Cogswell PC, Rogers-Graham KS, Lowe SW, Der CJ et al. (1997). Requirement of NF-kappaB activation to suppress p53-independent apoptosis induced by oncogenic Ras. Science 278: 1812–1815.
Nicke B, Bastien J, Khanna SJ, Warne PH, Cowling V, Cook SJ et al. (2005). Involvement of MINK, a Ste20 family kinase, in Ras oncogene-induced growth arrest in human ovarian surface epithelial cells. Mol Cell 20: 673–685.
Okada T, Masuda T, Shinkai M, Kariya K, Kataoka T . (1996). Post-translational modification of H-Ras is required for activation of, but not for association with, B-Raf. J Biol Chem 271: 4671–4678.
Pfeifer GP, Yoon JH, Liu L, Tommasi S, Wilczynski SP, Dammann R . (2002). Methylation of the RASSF1A gene in human cancers. Biol Chem 383: 907–914.
Ponting DP, Benjamin DR . (1996). A novel family of Ras-binding domains. Trends Biochem Sci 21: 422–425.
Serrano M, Lin AW, McCurrach ME, Beach D, Lowe SW . (1997). Oncogenic ras provokes premature cell senescence associated with accumulation of p53 and p16INK4a. Cell 88: 593–602.
Shields JM, Pruitt K, McFall A, Shaub A, Der CJ . (2000). Understanding Ras: ‘it ain’t over ‘til it's over’. Trends Cell Biol 10: 147–154.
Smyth RL, Openshaw PJ . (2006). Bronchiolitis. Lancet 368: 312–322.
Solski PA, Quilliam LA, Coats SG, Der CJ, Buss JE . (1995). Targeting proteins to membranes using signal sequences for lipid modification. Methods Enzymol 250: 435–454.
Tommasi S, Dammann R, Jin SG, Zhang XX, Avruch J, Pfeifer GP . (2002). RASSF3 and NORE1: identification and cloning of two human homologues of the putative tumor suppressor gene RASSF1. Oncogene 21: 2713–2720.
Tommasi S, Dammann R, Zhang Z, Wang Y, Liu L, Tsark WM et al. (2005). Tumor susceptibility of Rassf1a knockout mice. Cancer Res 65: 92–98.
Vos MD, Dallol A, Eckfeld K, Allen NP, Donninger H, Hesson L et al. (2006). The RASSF1A tumor suppressor activates Bax via MOAP-1. J Biol Chem 281: 4557–4563.
Vos MD, Ellis CA, Bell A, Birrer MJ, Clark GJ . (2000). Ras uses the novel tumor suppressor RASSF1 as an effector to mediate apoptosis. J Biol Chem 275: 35669–35672.
Vos MD, Ellis CA, Elam C, Ulku AS, Taylor BJ, Clark GJ . (2003a). RASSF2 is a novel K-Ras-specific effector and potential tumor suppressor. J Biol Chem 278: 28045–28051.
Vos MD, Martinez A, Elam C, Dallol A, Taylor BJ, Latif F et al. (2004). A role for the RASSF1A tumor suppressor in the regulation of tubulin polymerization and genomic stability. Cancer Res 64: 4244–4250.
Vos MD, Martinez A, Ellis CA, Vallecorsa T, Clark GJ . (2003b). The pro-apoptotic Ras effector Nore1 may serve as a Ras-regulated tumor suppressor in the lung. J Biol Chem 278: 21938–21943.
White MA, Nicolette C, Minden A, Polverino A, Van AL, Karin M et al. (1995). Multiple Ras functions can contribute to mammalian cell transformation. Cell 80: 533–541.
Williams JG, Drugan JK, Yi GS, Clark GJ, Der CJ, Campbell SL . (2000). Elucidation of binding determinants and functional consequences of Ras/Raf-cysteine-rich domain interactions. J Biol Chem 275: 22172–22179.
Acknowledgements
This work was supported in part by Intramural funds of the National Cancer Institute and RR018733 (GJC), FL is supported in part by Cancer Research UK.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Allen, N., Donninger, H., Vos, M. et al. RASSF6 is a novel member of the RASSF family of tumor suppressors. Oncogene 26, 6203–6211 (2007). https://doi.org/10.1038/sj.onc.1210440
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1210440
Keywords
This article is cited by
-
The Hippo signaling pathway contributes to the 2,5-Hexadion-induced apoptosis of ovarian granulosa cells
Journal of Ovarian Research (2023)
-
The role of miR-155 in cigarette smoke-induced pulmonary inflammation and COPD
Mucosal Immunology (2020)
-
TCF4 (E2-2) harbors tumor suppressive functions in SHH medulloblastoma
Acta Neuropathologica (2019)
-
Tumor suppressor C-RASSF proteins
Cellular and Molecular Life Sciences (2018)
-
Comparative Modeling, Molecular Docking, and Revealing of Potential Binding Pockets of RASSF2; a Candidate Cancer Gene
Interdisciplinary Sciences: Computational Life Sciences (2017)