Abstract
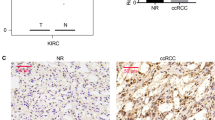
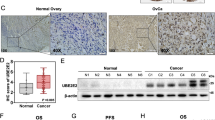
The UbcH10 gene codes for a protein that belongs to the ubiquitin-conjugating enzyme family. Previous studies of our group suggest UbcH10 expression as a valid indicator of the proliferative and aggressive status of thyroid carcinomas. Therefore, to better understand the process of ovarian carcinogenesis, and to look for possible tools to be used as prognostic markers in these neoplasias, we decided to extend the analysis of the UbcH10 expression to the ovarian neoplastic disease. We found that the UbcH10 gene was upregulated in some ovarian carcinoma cell lines analysed. Then, immunohistochemical studies demonstrate that UbcH10 expression significantly correlates with the tumor grade and the undifferentiated histotype of the ovarian carcinomas. Furthermore, a significant relationship between UbcH10 expression and overall survival was observed. Finally, the block of UbcH10 protein synthesis by RNA interference inhibited the growth of ovarian carcinoma cell lines, suggesting a role of UbcH10 overexpression in ovarian carcinogenesis. Therefore, all these data taken together suggest the possibility to use UbcH10 detection as a marker for the diagnosis and prognosis of these neoplastic diseases and open the perspective of a therapy of some ovarian carcinomas based on the suppression of the UbcH10 synthesis and/or function.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Aldovini D, Demichelis F, Doglioni C, Di Vizio D, Galligioni E, Brugnara S et al. (2006). M-CAM expression as marker of poor prognosis in epithelial ovarian cancer. Int J Cancer Jun 27 [Epub ahead of print].
Benedet JL, Bender H, Jones III H, Ngan HY, Pecorelli S . (2000). FIGO staging classifications and clinical practice guidelines in the management of gynecologic cancers. FIGO Committee on Gynecologic Oncology. Int J Gynaecol Obstet 70: 209–262.
Feki A, Irminger-Finger I . (2004). Mutational spectrum of p53 mutations in primary breast and ovarian tumors. Crit Rev Oncol Hematol 52: 103–116.
Hershko A, Ciechanover A . (1998). The ubiquitin system. Annu Rev Biochem 67: 425–479.
Jemal A, Murray T, Ward E, Samuels A, Tiwari RC, Ghafoor A et al. (2005). Cancer statistics, 2005. CA Cancer J Clin 55: 10–30.
Joazeiro CA, Weissman AM . (2000). Ring finger proteins: mediators of ubiquitin ligase activity. Cell 1 102: 549–552.
Liu X, Minin V, Huang Y, Seligson DB, Horvath S . (2004). Statistical methods for analysing tissue microarray data. J Biopharm Stat 14: 671–685.
Masciullo V, Baldassarre G, Pentimalli F, Berlingieri MT, Boccia A, Chiappetta G et al. (2003). HMGA1 protein over-expression is a frequent feature of epithelial ovarian carcinomas. Carcinogenesis 24: 1191–1198.
Ozols RF, Bookman MA, Connolly DC, Daly MB, Godwin AK, Schilder RJ et al. (2004). Focus on epithelial ovarian cancer. Cancer Cell 5: 19–24.
Pallante P, Berlingieri MT, Troncone G, Kruhoffer M, Orntoft TF, Viglietto G et al. (2005). UbcH10 overexpression may represent a marker of anaplastic thyroid carcinomas. Br J Cancer 93: 464–471.
Pierantoni GM, Rinaldo C, Esposito F, Mottolese M, Soddu S, Fusco A . (2005). High Mobility Group A1 (HMGA1) proteins interact with p53 and inhibit its apoptotic activity. Cell Death Differ (9 December): 1–10 (Epub ahead of print).
R Development Core Team (2004). A Language and Environment for Statistical Computing. R Foundation for Statistical Computing: Vienna, Austria.
Tavassoli FA, Devilee P . (2003). World Health Oraganization Classification of Tumours. Tumours of the breast and female genital organs. IARC Press: Lyon.
Troncone G, Iaccarino A, Caleo A, Bifano D, Pettinato G, Palombini L . (2003). p27 Kip1 protein expression in Hashimoto's thyroiditis. J Clin Pathol 56: 587–591.
Vacher-Lavenu MC, Le Tourneau A, Duvillard P, Godefroy N, Pinel MC . (1993). Pathological classification and grading of primary ovarian carcinoma: experience of the ARTAC ovarian study group. Bull Cancer 80: 135–141.
Wagner KW, Sapinoso LM, El-Rifai W, Frierson HF, Butz N, Mestan J et al. (2004). Overexpression, genomic amplification and therapeutic potential of inhibiting the UbcH10 ubiquitin conjugase in human carcinomas of diverse anatomic origin. Oncogene 23: 6621–6629.
Welcsh PL, King MC . (2001). BRCA1 and BRCA2 and the genetics of breast and ovarian cancer. Hum Mol Genet 10: 705–713.
Acknowledgements
This work was supported by grants from the Associazione Italiana Ricerca sul Cancro (AIRC), Progetto Strategico Oncologia Consiglio Nazionale delle Ricerche, the Ministero dell'Università e della Ricerca Scientifica e Tecnologica (MIUR), and ‘Piani di Potenziamento della Rete Scientifica e Tecnologica’ CLUSTER C-04, the Programma Italia-USA sulla Terapia dei Tumori coordinated by Professor Cesare Peschle and ‘Ministero della Salute’. This work was supported from NOGEC-Naples Oncogenomic Center. We thank the Associazione Partenopea per le Ricerche Oncologiche (APRO) for its support.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Berlingieri, M., Pallante, P., Guida, M. et al. UbcH10 expression may be a useful tool in the prognosis of ovarian carcinomas. Oncogene 26, 2136–2140 (2007). https://doi.org/10.1038/sj.onc.1210010
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1210010
Keywords
This article is cited by
-
The correlation between the expression of ubiquitin-conjugating enzyme 2C and prostate cancer prognosis
Journal of Cancer Research and Clinical Oncology (2023)
-
Ubiquitin-conjugating enzyme E2C regulates apoptosis-dependent tumor progression of non-small cell lung cancer via ERK pathway
Medical Oncology (2015)
-
High expression of ubiquitin-conjugating enzyme 2C (UBE2C) correlates with nasopharyngeal carcinoma progression
BMC Cancer (2013)
-
Inhibition of ubiquitin conjugating enzyme UBE2C reduces proliferation and sensitizes breast cancer cells to radiation, doxorubicin, tamoxifen and letrozole
Cellular Oncology (2013)
-
Expression of UbcH10 in pancreatic ductal adenocarcinoma and its correlation with prognosis
Tumor Biology (2013)