Abstract
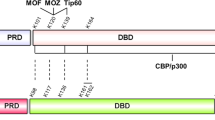
Reversible acetylation on protein lysine residues has been shown to regulate the function of both nuclear proteins such as histones and p53 and cytoplasmic proteins such as α-tubulin. To identify novel acetylated proteins, we purified several proteins by the affinity to an anti-acetylated-lysine antibody from cells treated with trichostatin A (TSA). Among the proteins identified, here we report acetylation of the SV40 large T antigen (T-Ag). The acetylation site was determined to be lysine-697, which is located adjacent to the C-terminal Cdc4 phospho-degron (CPD). Overexpression of the CBP acetyltransferase acetylated T-Ag, whereas HDAC1, HDAC3 and SIRT1 bound and deacetylated T-Ag. The acetylation and deacetylation occurred independently of p53, a binding partner of T-Ag, but the acetylation was enhanced in the presence of p53. T-Ag in the cells treated with TSA and NA or the acetylation mimic mutant (K697Q) became unstable in COS-7 cells, suggesting that acetylation regulates stability of T-Ag. Indeed, NIH3T3 cells stably expressing K697Q showed decreased anchorage-independent growth compared with those expressing wild type or the K697R mutant. These results demonstrate that acetylation destabilizes T-Ag and regulates the transforming activity of T-Ag in NIH3T3 cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Avantaggiati ML, Carbone M, Graessmann A, Nakatani Y, Howard B, Levine AS . (1996). EMBO J 15: 2236–2248.
Bitterman KJ, Anderson RM, Cohen HY, Latorre-Esteves M, Sinclair DA . (2002). J Biol Chem 277: 45099–45107.
Brown M, McCormack M, Zinn KG, Farrell MP, Bikel I, Livingston DM . (1986). J Virol 60: 290–293.
Caron C, Boyault C, Khochbin S . (2005). Bioessays 27: 408–415.
Caron C, Col E, Khochbin S . (2003). Bioessays 25: 58–65.
Chan HM, Krstic-Demonacos M, Smith L, Demonacos C, La Thangue NB . (2001). Nat Cell Biol 3: 667–674.
Chen L, Fischle W, Verdin E, Greene WC . (2001). Science 293: 1653–1657.
Eckner R, Ludlow JW, Lill NL, Oldread E, Arany Z, Modjtahedi N et al. (1996). Mol Cell Biol 16: 3454–3464.
Fukazawa H, Mizuno S, Uehara Y . (1995). Anal Biochem 228: 83–90.
Fukazawa H, Noguchi K, Masumi A, Murakami Y, Uehara Y . (2004). Mol Cancer Ther 3: 1281–1288.
Furumai R, Komatsu Y, Nishino N, Khochbin S, Yoshida M, Horinouchi S . (2001). Proc Natl Acad Sci USA 98: 87–92.
Giandomenico V, Simonsson M, Gronroos E, Ericsson J . (2003). Mol Cell Biol 23: 2587–2599.
Gu W, Roeder RG . (1997). Cell 90: 595–606.
Hubbert C, Guardiola A, Shao R, Kawaguchi Y, Ito A, Nixon A et al. (2002). Nature 417: 455–458.
Imai S, Armstrong CM, Kaeberlein M, Guarente L . (2000). Nature 403: 795–800.
Ito A, Kawaguchi Y, Lai CH, Kovacs JJ, Higashimoto Y, Appella E et al. (2002). EMBO J 21: 6236–6245.
Jeong JW, Bae MK, Ahn MY, Kim SH, Sohn TK, Bae MH et al. (2002). Cell 111: 709–720.
Komatsu Y, Yukutake Y, Yoshida M . (2003). J Immunol Methods 272: 161–175.
Kouzarides T . (2000). EMBO J 19: 1176–1179.
Leduc C, Claverie P, Eymin B, Col E, Khochbin S, Brambilla E et al. (2006). Oncogene (in press).
Lill NL, Tevethia MJ, Eckner R, Livingston DM, Modjtahedi N . (1997). J Virol 71: 129–137.
Liu L, Scolnick DM, Trievel RC, Zhang HB, Marmorstein R, Halazonetis TD et al. (1999). Mol Cell Biol 19: 1202–1209.
Luo J, Nikolaev AY, Imai S, Chen D, Su F, Shiloh A et al. (2001). Cell 107: 137–148.
Luo J, Su F, Chen D, Shiloh A, Gu W . (2000). Nature 408: 377–381.
Matsuyama A, Shimazu T, Sumida Y, Saito A, Yoshimatsu Y, Seigneurin-Berny D et al. (2002). EMBO J 21: 6820–6831.
Mitsudomi T, Steinberg SM, Nau MM, Carbone D, D'Amico D, Bodner S et al. (1992). Oncogene 7: 171–180.
Muth V, Nadaud S, Grummt I, Voit R . (2001). EMBO J 20: 1353–1362.
Nash P, Tang X, Orlicky S, Chen Q, Gertler FB, Mendenhall MD et al. (2001). Nature 414: 514–521.
North BJ, Marshall BL, Borra MT, Denu JM, Verdin E . (2003). Mol Cell 11: 437–444.
Polesskaya A, Duquet A, Naguibneva I, Weise C, Vervisch A, Bengal E et al. (2000). J Biol Chem 275: 34359–34364.
Poulin DL, Kung AL, DeCaprio JA . (2004). J Virol 78: 8245–8253.
Sakaguchi K, Herrera JE, Saito S, Miki T, Bustin M, Vassilev A et al. (1998). Genes Develop 12: 2831–2841.
Sartorelli V, Puri PL, Hamamori Y, Ogryzko V, Chung G, Nakatani Y et al. (1999). Mol Cell 4: 725–734.
Spence SL, Pipas JM . (1994). J Virol 68: 4227–4240.
Stacy T, Chamberlain M, Cole CN . (1989). J Virol 63: 5208–5215.
Sullivan CS, Pipas JM . (2002). Microbiol Mol Biol Rev 66: 179–202.
Vaziri H, Dessain SK, Ng Eaton E, Imai SI, Frye RA, Pandita TK et al. (2001). Cell 107: 149–159.
Welcker M, Clurman BE . (2005). J Biol Chem 280: 7654–7658.
Xie AY, Bermudez VP, Folk WR . (2002). Mol Cell Biol 22: 7907–7918.
Yada M, Hatakeyama S, Kamura T, Nishiyama M, Tsunematsu R, Imaki H et al. (2004). EMBO J 23: 2116–2125.
Yoshida M, Kijima M, Akita M, Beppu T . (1990). J Biol Chem 265: 17174–17179.
Acknowledgements
We thank Tso-Pang Yao for providing the CBP, SIRT1-5 and p53, Xiang-Jiao Yang for the PCAF, Saadi Khochbin for the HA-TIP60. We are grateful for the support of RIKEN BSI's Research Resources Center for the LC-MS/MS analysis. This work was supported in part by the Program for the Promotion of Fundamental studies in Health Science of the National Institute of Biomedical Innovation (NIBIO), and a Grant-in-Aid for Scientific Research on Priority Area ‘Cancer’ from The Ministry of Education, Culture, Sports, Science and Technology, Japan.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Shimazu, T., Komatsu, Y., Nakayama, K. et al. Regulation of SV40 large T-antigen stability by reversible acetylation. Oncogene 25, 7391–7400 (2006). https://doi.org/10.1038/sj.onc.1209731
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1209731
Keywords
This article is cited by
-
Histone deacetylase III interactions with BK polyomavirus large tumor antigen may affect protein stability
Virology Journal (2023)
-
Nuclear receptor co-repressors are required for the histone-deacetylase activity of HDAC3 in vivo
Nature Structural & Molecular Biology (2013)
-
GCN5-dependent acetylation of HIV-1 integrase enhances viral integration
Retrovirology (2010)
-
Degradation of cyclin A is regulated by acetylation
Oncogene (2009)
-
Liver-specific deletion of histone deacetylase 3 disrupts metabolic transcriptional networks
The EMBO Journal (2008)