Abstract
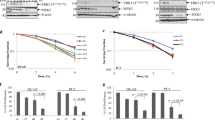
During metastases, cancer cells are temporarily exposed to the condition in which interactions with extracellular environment can be restricted (anchorage-independence). We demonstrate that the sensitivity of prostate cancer cell lines, DU145 and PC-3, to genotoxic treatment (cisplatin and γ-irradiation) increased several folds when cells were forced to grow in anchorage-independence. This enhanced drug sensitivity was associated with a severe impairment of homologous recombination-directed DNA repair (HRR). The mechanism involves Rad51, which is the major enzymatic component of HRR. The protein level of Rad51 and its recruitment to DNA double-strand breaks (DSBs) were both attenuated. Rad51 deficiency in anchorage-independence was not associated with Rad51 promoter activity and was not compensated by a constitutive overexpression of Rad51 cDNA. Instead, Rad51 protein level and its ability to colocalize with DSBs were restored in the presence of proteosome inhibitors, or when cells from the suspension cultures were allowed reattachment. Presented results indicate that anchorage-independence sensitizes prostate cancer cells to genotoxic agents; however, it also attenuates faithful component of DNA repair by targeting stability of Rad51. This temporal attenuation of HRR may contribute to the accumulation mutations after DNA damage and possibly the selection of new adaptations in cells, which survived genotoxic treatment.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Baumann P and West SC . (1998). Trends Biochem. Sci., 23, 247–251.
Bindra RS, Schaffer PJ, Meng A, Woo J, Maseide K, Roth ME, Lizardi P, Hedley DW, Bristow RG and Glazer PM . (2004). Mol. Cell. Biol., 24, 8504–8518.
Bondar VM and McConkey DJ . (2002). Prostate, 51, 42–49.
Chen G, Yuan SS, Liu W, Xu Y, Trujillo K, Song B, Cong F, Goff SP, Wu Y, Arlinghaus R, Baltimore D, Gasser PJ, Park MS, Sung P and Lee EY . (1999). J. Biol. Chem., 274, 12748–12752.
Chen Y, Wang J, Fraig MM, Henderson K, Bissada NK, Watson DK and Schweinfest CW . (2003). Int. J. Oncol., 22, 1033–1043.
Chen Y, Wang J, Fraig MM, Metcalf J, Turner WR, Bissada NK, Watson DK and Schweinfest CW . (2001). Cancer Res., 61, 4112–4121.
Chendil D, Ranga RS, Meigooni D, Sathishkumar S and Ahmed MM . (2004). Oncogene, 23, 1599–1607.
Collis SJ, Tighe A, Scott SD, Roberts SA, Hendry JH and Margison GP . (2001). Nucleic Acids Res., 29, 1534–1538.
Davies AA, Masson JY, McIlwraith MJ, Stasiak AZ, Stasiak A, Venkitaraman AR and West SC . (2001). Mol. Cell, 7, 273–282.
Del Valle L, Enam S, Lassak A, Wang JY, Croul S, Khalili K and Reiss K . (2002). Clin. Cancer Res., 8, 1822–1830.
Essers J, Houtsmuller AB, van Veelen L, Paulusma C, Nigg AL, Pastink A, Vermeulen W, Hoeijmakers JH and Kanaar R . (2002). EMBO J., 21, 2030–2037.
Flygare J, Falt S, Ottervald J, Castro J, Dackland AL, Hellgren D and Wennborg A . (2001). Exp. Cell Res., 268, 61–69.
Hoeijmakers JH . (2001). Nature, 411, 366–374.
Huang Y, Nakada S, Ishiko T, Utsugisawa T, Datta R, Kharbanda S, Yoshida K, Talanian RV, Weichselbaum R, Kufe D and Yuan ZM . (1999). Mol. Cell. Biol., 19, 2986–2997.
Karan D, Lin MF, Johansson SL and Batra SK . (2003). Int. J. Cancer, 103, 285–293.
Khanna KK and Jackson SP . (2001). Nat. Genet., 27, 247–254.
Kovalenko OV, Plug AW, Haaf T, Gonda DK, Ashley T, Ward DC, Radding CM and Golub EI . (1996). Proc. Natl. Acad. Sci. USA, 93, 2958–2963.
Labhart P . (1999). Eur. J. Biochem., 265, 849–861.
Lim DS and Hasty P . (1996). Mol. Cell. Biol., 16, 7133–7143.
Ludwig T, Eggenschwiler J, Fisher P, D’Ercole AJ, Davenport ML and Efstratiadis A . (1996). Dev. Biol., 177, 517–535.
Lundin C, Erixon K, Arnaudeau C, Schultz N, Jenssen D, Meuth M and Helleday T . (2002). Mol. Cell. Biol., 22, 5869–5878.
Mills KD, Ferguson DO and Alt FW . (2003). Immunol. Rev., 194, 77–95.
Peruzzi F, Prisco M, Dews M, Salomoni P, Grassilli E, Romano G, Calabretta B and Baserga R . (1999). Mol. Cell. Biol., 19, 7203–7215.
Petrini JH . (2000). Curr. Opin. Cell Biol., 12, 293–296.
Pierce AJ and Jasin M . (2001). Mol. Cell, 8, 1160–1161.
Pierce AJ, Johnson RD, Thompson LH and Jasin M . (1999). Genes Dev., 13, 2633–2638.
Raderschall E, Stout K, Freier S, Suckow V, Schweiger S and Haaf T . (2002). Cancer Res., 62, 219–225.
Reiss K, D’Ambrosio C, Tu X, Tu C and Baserga R . (1998a). Clin. Cancer Res., 4, 2647–2655.
Reiss K, Valentinis B, Tu X, Xu SQ and Baserga R . (1998b). Exp. Cell Res., 242, 361–372.
Reiss K, Wang JY, Romano G, Furnari FB, Cavenee WK, Morrione A, Tu X and Baserga R . (2000). Oncogene, 19, 2687–2694.
Reiss K, Yumet G, Shan S, Huang Z, Alnemri E, Srinivasula SM, Wang JY, Morrione A and Baserga R . (1999). J. Cell Physiol., 181, 124–135.
Richardson C and Jasin M . (2000). Mol. Cell. Biol., 20, 9068–9075.
Richardson C, Stark JM, Ommundsen M and Jasin M . (2004). Oncogene, 23, 546–553.
Rodgers KJ and Dean RT . (2003). Int. J. Biochem. Cell Biol., 35, 716–727.
Schindewolf C, Lobenwein K, Trinczek K, Gomolka M, Soewarto D, Fella C, Pargent W, Singh N, Jung T and Hrabe de Angelis M . (2000). Mamm. Genome., 11, 552–554.
Schmutte C, Tombline G, Rhiem K, Sadoff MM, Schmutzler R, von Deimling A and Fishel R . (1999). Cancer Res., 59, 4564–4569.
Sieber OM, Heinimann K and Tomlinson IP . (2003). Nat. Rev. Cancer, 3, 701–708.
Sigurdsson S, Van Komen S, Petukhova G and Sung P . (2002). J. Biol. Chem., 277, 42790–42794.
Skorski T, Nieborowska-Skorska M, Campbell K, Iozzo RV, Zon G, Darzynkiewicz Z and Calabretta B . (1995). J. Exp. Med., 182, 1645–1653.
Slupianek A, Schmutte C, Tombline G, Nieborowska-Skorska M, Hoser G, Nowicki MO, Pierce AJ, Fishel R and Skorski T . (2001). Mol. Cell, 8, 795–806.
Sonoda E, Sasaki MS, Buerstedde JM, Bezzubova O, Shinohara A, Ogawa H, Takata M, Yamaguchi-Iwai Y and Takeda S . (1998). EMBO J., 17, 598–608.
Tombline G and Fishel R . (2002). J. Biol. Chem., 277, 14417–14425.
Tombline G, Shim KS and Fishel R . (2002). J. Biol. Chem., 277, 14426–14433.
Trojanek J, Ho T, Del Valle L, Nowicki M, Wang JY, Lassak A, Peruzzi F, Khalili K, Skorski T and Reiss K . (2003). Mol. Cell. Biol., 23, 7510–7524.
Valentinis B, Reiss K and Baserga R . (1998). J. Cell Physiol., 176, 648–657.
Valerie K and Povirk LF . (2003). Oncogene, 22, 5792–5812.
Van Komen S, Petukhova G, Sigurdsson S and Sung P . (2002). J. Biol. Chem., 277, 43578–43587.
Wang JY, Del Valle L, Gordon J, Rubini M, Romano G, Croul S, Peruzzi F, Khalili K and Reiss K . (2001). Oncogene, 20, 3857–3868.
Wozniak K and Blasiak J . (2002). Acta Biochim. Pol., 49, 583–596.
Xia SJ, Shammas MA and Shmookler Reis RJ . (1997). Mol. Cell. Biol., 17, 7151–7158.
Yeh CC, Lee C and Dahiya R . (2001). Biochem. Biophys. Res. Commun., 285, 409–413.
Zhu XD, Kuster B, Mann M, Petrini JH and Lange T . (2000). Nat. Genet., 25, 347–352.
Acknowledgements
We gratefully acknowledge editorial assistance of Dr Sidney Croul and Jessica Otte, technical manager of the Center for Neurovirology and Cancer Biology, for her operational efforts in our laboratory. TS is a scholar of the Leukemia and Lymphoma Society.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Wang, J., Ho, T., Trojanek, J. et al. Impaired homologous recombination DNA repair and enhanced sensitivity to DNA damage in prostate cancer cells exposed to anchorage-independence. Oncogene 24, 3748–3758 (2005). https://doi.org/10.1038/sj.onc.1208537
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1208537
Keywords
This article is cited by
-
PKMYT1: A Potential Target for CCNE1 Amplificated Colorectal Tumors
Cell Biochemistry and Biophysics (2023)