Abstract
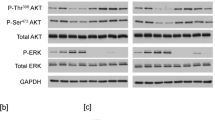
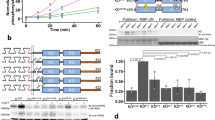
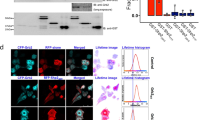
Cellular transformation occurs only in cells that express both ErbB1 and ErbB4 receptors, but not in cells expressing only one or the other of these receptors. However, when both receptors are coexpressed and ligand-stimulated, they interact with virtually the same adaptor/effector proteins as when expressed singly. To reveal the underlying regulatory mechanism of the kinase/phosphatase network in ErbB homo- and heterodimer receptor signaling, extracellular signal-regulated kinase (ERK) and Akt activities were evaluated in the presence of several enzyme inhibitors in ligand-induced cells expressing ErbB1 (E1), ErbB4 (E4), and ErbB1/ErbB4 (E1/4) receptor. The PP2A inhibitor okadaic acid showed receptor-specific inhibitory profiles for ERK and Akt activities. Moreover, B-Raf isolated only from E1/4 cells could induce in vitro phosphorylation for MEK; this B-Raf kinase activity was abolished by pretreatment of the cells with okadaic acid. Our study further showed that the E1/4 cell-specific B-Raf activity was stimulated by PLCγ and subsequent Rap1 activation. The present study suggests that B-Raf kinase, which was specifically activated in the cells coexpressing ErbB1 and ErbB4 receptors, elevates total ERK activity within the cell and, therefore, can induce cellular transformation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Abbreviations
- CHO:
-
Chinese hamster ovary
- EGFR:
-
EGF receptor
- EGF:
-
epidermal growth factor
- HRG:
-
heregulin
- MAPK:
-
mitogen-activated protein kinase
- MEK:
-
extracellular signal-regulated kinase kinase
- ERK:
-
extracellular signal-regulated kinase
- SH2:
-
Src homology 2 domain
- Grb2:
-
growth factor receptor-binding protein 2
- Shc:
-
Src homology and collagen domain protein
- PI3K:
-
phosphatidylinositol 3′-kinase
- PLCγ:
-
phosphoinositide-specific phospholipase C-γ
- PKA:
-
protein kinase A
- PH domain:
-
pleckstrin homology domain
- PP2A:
-
protein phosphatase 2A
References
Abraham D, Podar K, Pacher M, Kubicek M, Welzel N, Hemmings BA, Dilworth SM, Mischak H, Kolch W and Baccarini M . (2000). J. Biol. Chem., 275, 22300–22304.
Andjelkovic M, Jakubowicz T, Cron P, Ming XF, Han JW and Hemmings BA . (1996). Proc. Natl. Acad. Sci. USA, 93, 5699–5704.
Andjelkovic M, Maira SM, Cron P, Parker PJ and Hemmings BA . (1999). Mol. Cell Biol., 19, 5061–5072.
Aoki M, Batista O, Bellacosa A, Tsichlis P and Vogt PK . (1998). Proc. Natl. Acad. Sci. USA, 95, 14950–14955.
Brose MS, Volpe P, Feldman M, Kumar M, Rishi I, Gerrero R, Einhorn E, Herlyn M, Minna J, Nicholson A, Roth JA, Albelda SM, Davies H, Cox C, Brignell G, Stephens P, Futreal PA, Wooster R, Stratton MR and Weber BL . (2002). Cancer Res., 62, 6997–7000.
Busca R, Abbe P, Mantoux F, Aberdam E, Peyssonnaux C, Eychene A, Ortonne JP and Ballotti R . (2000). EMBO J., 19, 2900–2910.
Chong H, Vikis HG. and Guan K-L . (2003). Cell. Signal., 15, 463–469.
Cohen BD, Kiener PA, Green JM, Foy L, Fell HP and Zhang K . (1996). J. Biol. Chem., 271, 30897–30903.
Dhillon AS, Pollock C, Steen H, Shaw PE, Mischak H and Kolch W . (2002). Mol. Cell Biol., 22, 3237–3246.
Guo FF, Kumahara E and Saffen D . (2001). J. Biol. Chem., 276, 25568–25581.
Hatakeyama M, Kimura S, Naka T, Kawasaki T, Yumoto N, Ichikawa M, Kim JH, Saito K, Saeki M, Shirouzu M, Yokoyama S and Konagaya A . (2003). Biochem. J., 373, 451–463.
Heinrich R, Neel BG and Rapoport TA . (2002). Mol. Cell, 9, 957–970.
Ivaska J, Nissinen L, Immonen N, Eriksson JE, Kahari VM and Heino J . (2002). Mol. Cell Biol., 22, 1352–1359.
Jaumot M and Hancock JF . (2001). Oncogene, 20, 3949–3958.
Jones JT, Akita RW and Sliwkowski MX . (1999). FEBS Lett., 447, 227–231.
Kao S, Jaiswal RK, Kolch W and Landreth GE . (2001). J. Biol. Chem., 276, 18169–18177.
Keyse SM . (2000). Curr. Opin. Cell Biol., 12, 186–192.
Kim JH, Saito K and Yokoyama S . (2002). Eur. J. Biochem., 269, 2323–2329.
Kolch W . (2000). Biochem J., 351, 289–305.
Kubicek M, Pacher M, Abraham D, Podar K, Eulitz M and Baccarini M . (2002). J. Biol. Chem., 277, 7913–7919.
MacNicol MC and MacNicol AM . (1999). J. Biol. Chem., 274, 13193–13197.
Oehrl W, Rubio I and Wetzker R . (2003). J. Biol. Chem., 278, 17819–17826.
Olayioye MA, Graus-Porta D, Beerli RR, Rohrer J, Gay B and Hynes NE . (1998). Mol. Cell Biol., 18, 5042–5051.
Olayioye MA, Neve RM, Lane HA and Hynes NE . (2000). EMBO J., 19, 3159–3167.
Papin C, Denouel-Galy A, Laugier D, Calothy G and Eychène A . (1998). J. Biol. Chem., 273, 24939–24947.
Payne DM, Rossomando AJ, Martino P, Erickson AK, Her JH, Shabanowitz J, Hunt DF, Weber MJ and Sturgill TW . (1991). EMBO J., 10, 885–892.
Peraldi P, Frodin M, Barnier JV, Calleja V, Scimeca JC, Filloux C, Calothy G and Van Obberghen E . (1995). FEBS Lett., 357, 290–296.
Resjö S, Göransson O, Härndahl L, Zolnierowicz S, Manganiello V and Degerman E . (2002). Cell. Signal., 14, 231–238.
Riese II DJ, van Raaij TM, Plowman GD, Andrews GC and Stern DF . (1995). Mol. Cell Biol., 15, 5770–5776.
Riese DJ, Kim ED, Elenius K, Buckley S, Klagsbrun M, Plowman GD and Stern DF . (1996). J. Biol. Chem., 271, 20047–20052.
Rodrigues GA, Falasca M, Zhang Z, Ong SH and Schlessinger J . (2000). Mol. Cell Biol., 20, 1448–1459.
Sable CL, Filippa N, Filloux C, Hemmings BA and Van Obberghen E . (1998). J. Biol. Chem., 273, 29600–29606.
Shelly M, Pinkas-Kramarski R, Guarino BC, Waterman H, Wang LM, Lyass L, Alimandi M, Kuo A, Bacus SS, Pierce JH, Andrews GC and Yarden Y . (1998). J. Biol. Chem., 273, 10496–10505.
Simon MA . (2000). Cell, 103, 13–15.
Strack S . (2002). J. Biol. Chem., 277, 41525–41532.
Thomas CC, Deak M, Alessi DR and van Aalten DMF . (2002). Curr. Biol., 12, 1256–1262.
Tzahar E, Moyer JD, Waterman H, Barbacci EG, Bao J, Levkowitz G, Shelly M, Strano S, Pinkas-Kramarski R, Pierce JH, Andrews GC and Yarden Y . (1998). EMBO J., 17, 5948–5963.
Tzahar E, Waterman H, Chen X, Levkowitz G, Karunagaran D, Lavi S, Ratzkin BJ and Yarden Y . (1996). Mol. Cell Biol., 16, 5276–5287.
Tzahar E, Pinkas-Kramarski R, Moyer JD, Klapper LN, Alroy I, Levkowitz G, Shelly M, Henis S, Eisenstein M, Ratzkin BJ, Sela M, Andrews GC and Yarden Y . (1997). EMBO J., 16, 4938–4950.
Wang LM, Kuo A, Alimandi M, Veri MC, Lee CC, Kapoor V, Ellmore N, Chen XH and Pierce JH. . (1998). Proc. Natl. Acad. Sci. USA, 95, 6809–6814.
Yarden Y and Sliwkowski MX . (2001). Nat. Rev. Mol. Cell Biol., 2, 127–137.
York RD, Yao H, Dillon T, Ellig CL, Eckert SP, McCleskey EW and Stork PJ . (1998). Nature, 392, 622–626.
Zhang K, Sun J, Liu N, Wen D, Chang D, Thomason A and Yoshinaga SK . (1996). J. Biol. Chem., 271, 3884–3890.
Zhou B, Wang Z, Zhao Y, Brautigan DL and Zhang Z . (2002). J. Biol. Chem., 277, 31818–31825.
Zwartkruis FJ, Wolthuis RM, Nabben NM, Franke B and Bos JL . (1998). EMBO J., 17, 5905–5912.
Acknowledgements
We thank Ms Mihoro Saeki for constructing CHO cells expressing ErbB1 and ErbB4 receptors. We also thank Dr Takashi Naka for critically reading the manuscript and Mr Akinobu Fukuzaki for preparing the manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hatakeyama, M., Yumoto, N., Yu, X. et al. Transformation potency of ErbB heterodimer signaling is determined by B-Raf kinase. Oncogene 23, 5023–5031 (2004). https://doi.org/10.1038/sj.onc.1207664
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1207664
Keywords
This article is cited by
-
At the crossroads: EGFR and PTHrP signaling in cancer-mediated diseases of bone
Odontology (2012)
-
Expression of the ErbB4 receptor causes reversal regulation of PP2A in the Shc signal transduction pathway in human cancer cells
Molecular and Cellular Biochemistry (2006)