Abstract
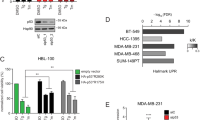
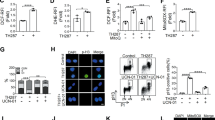
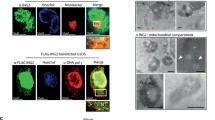
p53 can eliminate damaged cells through the induction of mitochondria-mediated apoptosis. Recent observations have provided strong evidence that a fraction of total p53 translocates to mitochondria specifically in response to a death stimulus. Unexpectedly, mutant p53, which is expressed at much higher levels than wild type in unstressed cells, is apparently always present at the mitochondria, independent of apoptotic signal. This prompted us to ask whether cell lines with intact p53-dependent apoptosis and cell cycle arrest pathways exist in which the mitochondrial localization of wild-type p53, like that of mutant, is independent of a death stimulus and instead, correlates with the total p53 levels. Here, we document that human HCT116 colorectal carcinoma cells treated with adriamycin or 5-fluorouracil (5FU) can accumulate total p53 to equally high levels, and mitochondrial p53 to proportionate levels, although only 5FU treatment provoked p53-dependent apoptosis. Along the same line, HCT116 derivatives with increased basal p53 levels, and glioblastoma cells with a doxycycline-inducible p53, also revealed proportionate mitochondrial p53 levels, and even unstressed HCT116 cells had some p53 located at the mitochondria. Finally, mitochondrial and total p53 showed distinct post-translational modifications. Thus, cell lines exist in which the mitochondrial p53 levels parallel total levels independent of apoptosis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Attardi LD, Reczek EE, Cosmas C, Demicco EG, McCurrach ME, Lowe SW and Jacks T . (2000). Genes Dev., 14, 704–718.
Bogenhagen D and Clayton DA . (1974). J. Biol. Chem., 249, 7991–7995.
Bunz F, Dutriaux A, Lengauer C, Waldman T, Zhou S, Brown JP, Sedivy JM, Kinzler KW and Vogelstein B . (1998). Science, 282, 1497–1501.
Bunz F, Fauth C, Speicher MR, Dutriaux A, Sedivy JM, Kinzler KW, Vogelstein B and Lengauer C . (2002). Cancer Res., 62, 1129–1133.
Bunz F, Hwang PM, Torrance C, Waldman T, Zhang Y, Dillehay L, Williams J, Lengauer C, Kinzler KW and Vogelstein B . (1999). J. Clin. Invest., 104, 263–269.
Caelles C, Helmberg A and Karin M . (1994). Nature, 370, 220–223.
Ding HF, Lin YL, McGill G, Juo P, Zhu H, Blenis J, Yuan J and Fisher DE . (2000). J. Biol. Chem., 275, 38905–38911.
Fortin A, Cregan SP, MacLaurin JG, Kushwaha N, Hickman ES, Thompson CS, Hakim A, Albert PR, Cecconi F, Helin K, Park DS and Slack RS . (2001). J. Cell Biol., 155, 207–216.
Gao C and Tsuchida N . (1999). Jpn. J. Cancer Res., 90, 180–187.
Gu W and Roeder RG . (1997). Cell, 90, 595–606.
Hwang PM, Bunz F, Yu J, Rago C, Chan TA, Murphy MP, Kelso GF, Smith RA, Kinzler KW and Vogelstein B . (2001). Nat. Med., 7, 1111–1117.
Iliakis G, Wang Y, Guan J and Wang H . (2003). Oncogene, 22, 5834–5847.
Jallepalli PV, Lengauer C, Vogelstein B and Bunz F . (2003). J. Biol. Chem., 278, 20475–20479.
Johnstone RW, Ruefli AA and Lowe SW . (2002). Cell, 108, 153–164.
Lin Y, Ma W and Benchimol S . (2000). Nat. Genet., 26, 122–127.
Lohr K, Moritz C, Contente A and Dobbelstein M . (2003). J. Biol. Chem., 278, 32507–32516.
Mahyar-Roemer M, Katsen A, Mestres P and Roemer K . (2001). Int. J. Cancer, 94, 615–622.
Mahyar-Roemer M, Kohler H and Roemer K . (2002). BMC Cancer, 2, 27.
Mahyar-Roemer M and Roemer K . (2001). Oncogene, 20, 3387–3398.
Marchenko ND, Zaika A and Moll UM . (2000). J. Biol. Chem., 275, 16202–16212.
Mihara M, Erster S, Zaika A, Petrenko O, Chittenden T, Pancoska P and Moll UM . (2003). Mol. Cell, 11, 577–590.
Miyashita T and Reed JC . (1995). Cell, 80, 293–299.
Moroni MC, Hickman ES, Denchi EL, Caprara G, Colli E, Cecconi F, Muller H and Helin K . (2001). Nat. Cell Biol., 3, 552–558.
Nakano K and Vousden KH . (2001). Mol. Cell, 7, 683–694.
Oda E, Ohki R, Murasawa H, Nemoto J, Shibue T, Yamashita T, Tokino T, Taniguchi T and Tanaka N . (2000a). Science, 288, 1053–1058.
Oda K, Arakawa H, Tanaka T, Matsuda K, Tanikawa C, Mori T, Nishimori H, Tamai K, Tokino T, Nakamura Y and Taya Y . (2000b). Cell, 102, 849–862.
Owen-Schaub LB, Zhang W, Cusack JC, Angelo LS, Santee SM, Fujiwara T, Roth JA, Deisseroth AB, Zhang WW, Kruzel E and Radinsky R . (1995). Mol. Cell. Biol., 15, 3032–3040.
Prives C and Manley JL . (2001). Cell, 107, 815–818.
Robles AI, Bemmels NA, Foraker AB and Harris CC . (2001). Cancer Res., 61, 6660–6664.
Sansome C, Zaika A, Marchenko ND and Moll UM . (2001). FEBS Lett., 488, 110–115.
Schmitt CA, Fridman JS, Yang M, Lee S, Baranov E, Hoffman RM and Lowe SW . (2002). Cell, 109, 335–346.
Schuler M and Green DR . (2001). Biochem. Soc. Trans., 29, 684–688.
Scorrano L and Korsmeyer SJ . (2003). Biochem. Biophys. Res. Commun., 304, 437–444.
Van Meir EG, Polverini PJ, Chazin VR, Su Huang HJ, de Tribolet N and Cavenee WK . (1994). Nat. Genet., 8, 171–176.
Vousden KH and Lu X . (2002). Nat. Rev. Cancer, 2, 594–604.
Wagner AJ, Kokontis JM and Hay N . (1994). Genes Dev., 8, 2817–2830.
Waldman T, Kinzler KW and Vogelstein B . (1995). Cancer Res., 55, 5187–5190.
Wang L, Wu Q, Qiu P, Mirza A, McGuirk M, Kirschmeier P, Greene, JR, Wang Y, Pickett CB and Liu S . (2001). J. Biol. Chem., 276, 43604–43610.
Wu GS, Burns TF, McDonald III ER, Jiang W, Meng R, Krantz ID, Kao G, Gan DD, Zhou JY, Muschel R, Hamilton SR, Spinner NB, Markowitz S, Wu G and el-Deiry WS . (1997). Nat. Genet., 17, 141–143.
Yan Y, Shay JW, Wright WE and Mumby MC . (1997). J. Biol. Chem., 272, 15220–15226.
Yu J, Wang Z, Kinzler KW, Vogelstein B and Zhang L . (2003). Proc. Natl. Acad. Sci. USA, 100, 1931–1936.
Acknowledgements
We thank Bert Vogelstein, Johns Hopkins University, Baltimore, USA for the cell lines HCT116, HCT116 p53−/− and HCT116 p21−/−; we are also grateful to Erwin Van Meir, Emory University, Atlanta, USA for the cell line LN-Z308 p53wt. Finally, we thank Gabi Kiefer, Institute of Anatomy, for excellent technical assistance with electron microscopy. This work was supported by the German Research Foundation (DFG) Grant RO 1205/5-1 to KR.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mahyar-Roemer, M., Fritzsche, C., Wagner, S. et al. Mitochondrial p53 levels parallel total p53 levels independent of stress response in human colorectal carcinoma and glioblastoma cells. Oncogene 23, 6226–6236 (2004). https://doi.org/10.1038/sj.onc.1207637
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1207637
Keywords
This article is cited by
-
Elevation of effective p53 expression sensitizes wild-type p53 breast cancer cells to CDK7 inhibitor THZ1
Cell Communication and Signaling (2022)
-
Mutant p53 exhibits trivial effects on mitochondrial functions which can be reactivated by ellipticine in lymphoma cells
Apoptosis (2011)
-
Resistance of mitochondrial p53 to dominant inhibition
Molecular Cancer (2008)
-
p73 and caspase-cleaved p73 fragments localize to mitochondria and augment TRAIL-induced apoptosis
Oncogene (2008)
-
p53 in mitochondria enhances the accuracy of DNA synthesis
Cell Death & Differentiation (2008)