Abstract
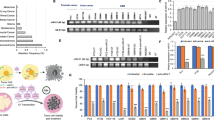
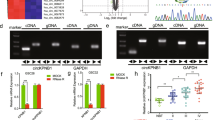
Extracellular proteases have been shown to cooperatively influence matrix degradation and tumor cell invasion through proteolytic cascades, with individual proteases having distinct roles in tumor growth, invasion, migration and angiogenesis. Matrix metalloproteases (MMP)-9 and cathepsin B have been shown to participate in the processes of tumor growth, vascularization and invasion of gliomas. In the present study, we used a cytomegalovirus promoter-driven DNA template approach to induce hairpin RNA (hpRNA)-triggered RNA interference (RNAi) to block MMP-9 and cathepsin B gene expression with a single construct. Transfection of a plasmid vector-expressing double-stranded RNA (dsRNA) for MMP-9 and cathepsin B significantly inhibited MMP-9 and cathepsin B expression and reduced the invasive behavior of SNB19, glioblastoma cell line in Matrigel and spheroid invasion models. Downregulation of MMP-9 and cathepsin B using RNAi in SNB19 cells reduced cell–cell interaction of human microvascular endothelial cells, resulting in the disruption of capillary network formation in both in vitro and in vivo models. Direct intratumoral injections of plasmid DNA expressing hpRNA for MMP-9 and cathepsin B significantly inhibited established glioma tumor growth and invasion in intracranial tumors in vivo. Further intraperitoneal (ip) injections of plasmid DNA expressing hpRNA for MMP-9 and cathepsin B completely regressed pre-established tumors for a long time (4 months) without any indication of these tumor cells. For the first time, these observations demonstrate that the simultaneous RNAi-mediated targeting of MMP-9 and cathepsin B has potential application for the treatment of human gliomas.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Abbreviations
- MMP-9:
-
matrix metalloproteases -9
- RNAi:
-
RNA interference vector expressing siRNA for cathepsin B and MMP-9 (pCM)
- CMV:
-
cytomegalovirus
- SV40:
-
simian virus type 40
- PCR:
-
polymerase chain reaction
- PBS:
-
phosphate-buffered saline
- FITC:
-
fluoresceine-5-isothiocyanate
- DiI:
-
1′-dioctadecyl-3,3,3′,3′-tetramethylindocarbocyanineperchlorate
- DiO:
-
3,3′-dioctadecyloxacarbocyanine perchlorate
- GFP:
-
green fluorescent protein
- ECM:
-
extracellular matrix
References
Arkona C and Wiederanders B . (1996). Biol. Chem., 377, 695–702.
Bergers G, Brekken R, McMahon G, Vu TH, Itoh T, Tamaki K, Tanzawa K, Thorpe P, Itohara S, Werb Z and Hanahan D . (2000). Nat. Cell Biol., 2, 737–744.
Bertrand JR, Pottier M, Vekris A, Opolon P, Maksimenko A and Malvy C . (2002). Biochem. Biophys. Res. Commun., 296, 1000–1004.
Bervar A, Zajc I, Sever N, Katunuma N, Sloane BF and Lah TT . (2003). Biol. Chem., 384, 447–455.
Boike G, Lah T, Sloane BF, Rozhin J, Honn K, Guirguis R, Stracke ML, Liotta LA and Schiffmann E . (1992). Melanoma Res., 1, 333–340.
Chandrasekar N, Jasti S, Alfred-Yung WK, Ali-Osman F, Dinh DH, Olivero WC, Gujrati M, Kyritsis AP, Nicolson GL, Rao JS and Mohanam S . (2000). Clin. Exp. Metast., 18, 337–342.
Coussens LM, Tinkle CL, Hanahan D and Werb Z . (2000). Cell, 103, 481–490.
Egeblad M and Werb Z . (2002). Nat. Rev. Cancer, 2, 161–174.
Elbashir SM, Harborth J, Lendeckel W, Yalcin A, Weber K and Tuschl T . (2001). Nature, 411, 494–498.
Emmert-Buck MR, Roth MJ, Zhuang Z, Campo E, Rozhin J, Sloane BF, Liotta LA and Stetler-Stevenson WG . (1994). Am. J. Pathol., 145, 1285–1290.
Go Y, Chintala SK, Mohanam S, Gokaslan Z, Venkaiah B, Bjerkvig R, Oka K, Nicolson GL, Sawaya R and Rao JS . (1997). Clin. Exp. Metast., 15, 440–446.
Guinec N, Dalet-Fumeron V and Pagano M . (1993). Biol. Chem. Hoppe Seyler, 374, 1135–1146.
Hammond SM, Boettcher S, Caudy AA, Kobayashi R and Hannon GJ . (2001). Science, 293, 1146–1150.
Himelstein BP, Canete-Soler R, Bernhard EJ, Dilks DW and Muschel RJ . (1994). Invas. Metast., 14, 246–258.
Hua J and Muschel RJ . (1996). Cancer Res., 56, 5279–5284.
Itoh T, Tanioka M, Matsuda H, Nishimoto H, Yoshioka T, Suzuki R and Uehira M . (1999). Clin. Exp. Metast., 17, 177–181.
Jen KY and Gewirtz AM . (2000). Stem Cells, 18, 307–319.
Kachra Z, Beaulieu E, Delbecchi L, Mousseau N, Berthelet F, Moumdjian R, Del Maestro R and Beliveau R . (1999). Clin. Exp. Metast., 17, 555–566.
Kobayashi H, Moniwa N, Sugimura M, Shinohara H, Ohi H and Terao T . (1993). Biochim. Biophys. Acta, 1178, 55–62.
Kobayashi H, Ohi H, Sugimura M, Shinohara H, Fujii T and Terao T . (1992). Cancer Res., 52, 3610–3614.
Kondraganti S, Mohanam S, Chintala SK, Kin Y, Jasti SL, Nirmala C, Lakka SS, Adachi Y, Kyritsis AP, Ali-Osman F, Sawaya R, Fuller GN and Rao JS . (2000). Cancer Res., 60, 6851–6855.
Konduri SD, Rao CN, Chandrasekar N, Tasiou A, Mohanam S, Kin Y, Lakka SS, Dinh D, Olivero WC, Gujrati M, Foster DC, Kisiel W and Rao JS . (2001). Oncogene, 20, 6938–6945.
Lakka SS, Jasti SL, Kyritsis AP, Yung WK, Ali-Osman F, Nicolson GL and Rao JS . (2000). Clin. Exp. Metast., 18, 245–252.
Lakka SS, Rajan M, Gondi C, Yanamandra N, Chandrasekar N, Jasti SL, Adachi Y, Siddique K, Gujrati M, Olivero W, Dinh DH, Kouraklis G, Kyritsis AP and Rao JS . (2002). Oncogene, 21, 8011–8019.
Leunig M, Yuan F, Menger MD, Boucher Y, Goetz AE, Messmer K and Jain RK . (1992). Cancer Res., 52, 6553–6560.
Lewis DL, Hagstrom JE, Loomis AG, Wolff JA and Herweijer H . (2002). Nat. Genet., 32, 107–108.
Liotta LA, Rao CN and Wewer UM . (1986). Annu. Rev. Biochem., 55, 1037–1057.
Mai J, Sameni M, Mikkelsen T and Sloane BF . (2002). Biol. Chem., 383, 1407–1413.
McCaffrey AP, Meuse L, Pham TT, Conklin DS, Hannon GJ and Kay MA . (2002). Nature, 418, 38–39.
Miyagishi M, Hayashi M and Taira K . (2003). Antisense Nucleic Acid Drug Dev., 13, 1–7.
Mohanam S, Jasti SL, Kondraganti SR, Chandrasekar N, Lakka SS, Kin Y, Fuller GN, Yung AW, Kyritsis AP, Dinh DH, Olivero WC, Gujrati M, Ali-Osman F and Rao JS . (2001). Oncogene, 20, 3665–3673.
Pozzi A, LeVine WF and Gardner HA . (2002). Oncogene, 21, 272–281.
Premzl A, Zavasnik-Bergant V, Turk V and Kos J . (2003). Exp. Cell Res., 283, 206–214.
Raithatha SA, Muzik H, Muzik H, Rewcastle NB, Johnston RN, Edwards DR and Forsyth PA . (2000). Neuro-oncology, 2, 145–150.
Rao JS, Steck PA, Mohanam S, Stetler-Stevenson WG, Liotta LA and Sawaya R . (1993). Cancer Res., 53, 2208–2211.
Redwood SM, Liu BC, Weiss RE, Hodge DE and Droller MJ . (1992). Cancer, 69, 1212–1219.
Rooprai HK, Van Meter T, Rucklidge GJ, Hudson L, Everall IP and Pilkington GJ . (1998). Int. J. Oncol., 13, 1153–1157.
Simon C, Goepfert H and Boyd D . (1998). Cancer Res., 58, 1135–1139.
Sloane BF, Rozhin J, Johnson K, Taylor H, Crissman JD and Honn KV . (1986). Proc. Natl. Acad. Sci. USA, 83, 2483–2487.
Stark GR, Kerr IM, Williams BR, Silverman RH and Schreiber RD . (1998). Annu. Rev. Biochem., 67, 227–264.
Vu TH, Shipley JM, Bergers G, Berger JE, Helms JA, Hanahan D, Shapiro SD, Senior RM and Werb Z . (1998). Cell, 93, 411–422.
Yu Q and Stamenkovic I . (1999). Genes Dev., 13, 35–48.
Zamore PD . (2001). Nat. Struct. Biol., 8, 746–750.
Acknowledgements
We thank Karen Minter for preparing the manuscript, and Sushma Jasti and Diana Meister for manuscript review. This research was supported by the National Cancer Institute CA 85216 and CA 75557 and the National Institute of Neurological Disorders and Stroke Grant NS47699 (to JSR).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lakka, S., Gondi, C., Yanamandra, N. et al. Inhibition of cathepsin B and MMP-9 gene expression in glioblastoma cell line via RNA interference reduces tumor cell invasion, tumor growth and angiogenesis. Oncogene 23, 4681–4689 (2004). https://doi.org/10.1038/sj.onc.1207616
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1207616
Keywords
This article is cited by
-
Role of lysosomes in physiological activities, diseases, and therapy
Journal of Hematology & Oncology (2021)
-
Down-regulation of cathepsin S and matrix metalloproteinase-9 via Src, a non-receptor tyrosine kinase, suppresses triple-negative breast cancer growth and metastasis
Experimental & Molecular Medicine (2018)
-
TRAIL DR5-CTSB crosstalk participates in breast cancer autophagy initiated by SAHA
Cell Death Discovery (2017)
-
Dual treatment with shikonin and temozolomide reduces glioblastoma tumor growth, migration and glial-to-mesenchymal transition
Cellular Oncology (2017)
-
Cathepsin L in tumor angiogenesis and its therapeutic intervention by the small molecule inhibitor KGP94
Clinical & Experimental Metastasis (2016)