Abstract
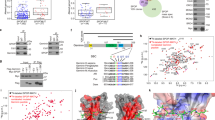
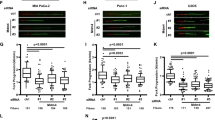
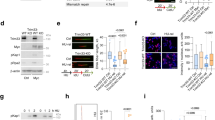
A domain-specific disruption was performed on the destruction box sequence of endogenous Geminin gene, an inhibitor of the DNA replication initiation complex, in a human cancer cell line HCT116 resulting in the formation of a protein that was stable in the G1 phase of the cell cycle. Although the total amount of Geminin in asynchronous cultures was not elevated, the G1-specific stabilization of Geminin, diminished chromatin loading of minichromosome maintenance complex, inhibited DNA replication, and resulted in the accumulation of cells in G1. The mutated Geminin suppressed in vivo tumorigenicity and in vitro cell growth. Cells carrying this mutation failed to support the replication of a plasmid bearing the oriP replicator of Epstein–Barr virus. The DNA damage checkpoint pathway was activated in the mutated cells with increased levels of p53 protein and its target, the p21 protein. All these deficits were rescued by overexpression of Cdt1, a replication initiator protein that binds to Geminin. Therefore, alteration of the cell cycle-dependent regulation of endogenous Geminin in human cells without increasing total protein level inhibits DNA replication and suppresses tumor growth.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Abbreviations
- APC:
-
anaphase-promoting complex
- ATM:
-
ataxia-telangiectasia-mutated
- BrdU:
-
bromodeoxyuridine
- EBV:
-
Epstein–Barr virus
- cdk:
-
cyclin-dependent kinases
- MCM:
-
minichromosome maintenance proteins
- ORC:
-
origin recognition complex
References
Abraham RT . (2001). Genes Dev., 15, 2177–2196.
Bell SP and Dutta A . (2002). Annu. Rev. Biochem., 71, 333–374.
Blow JJ and Hodgson B . (2002). Trends Cell Biol., 12, 72–78.
Blow JJ and Laskey RA . (1988). Nature, 332, 546–548.
Chaudhuri B, Xu H, Todorov I, Dutta A and Yates JL . (2001). Proc. Natl. Acad. Sci. USA, 98, 10085–10089.
Chehab NH, Malikzay A, Appel M and Halazonetis TD . (2000). Genes Dev., 14, 278–288.
DeCaprio JA, Ludlow JW, Figge J, Shew J-Y, Huang C-M, Lee W-H, Marsilio E, Paucha E and Livingston DM . (1988). Cell, 54, 275–282.
Dhar SK, Yoshida K, Machida Y, Khaira P, Chaudhuri B, Wohlschlegel JA, Leffak M, Yates J and Dutta A . (2001). Cell, 106, 287–296.
Diffley JF . (2001). Curr. Biol., 11, R367–R370.
Diffley JF, Cocker JH, Dowell SJ and Rowley A . (1994). Cell, 78, 303–316.
Dimitrova DS and Gilbert DM . (2000). Nat. Cell Biol., 2, 686–694.
Dyson NP, Howley M, Munger K and Harlow E . (1989). Science, 243, 934–937.
Feijoo C, Hall-Jackson C, Wu R, Jenkins D, Leitch J, Gilbert DM and Smythe C . (2001). J. Cell Biol., 154, 913–923.
Gavrieli Y, Sherman Y and Ben-Sasson SA . (1992). J. Cell Biol., 119, 493–501.
Gilbert DM, Neilson A, Miyazawa H, DePamphilis ML and Burhans WC . (1995). J. Biol. Chem., 270, 9597–9606.
Halbert CL, Demers GW and Galloway DA . (1992). J. Virol., 66, 2125–2134.
Kroll KL, Salic AN, Evans LM and Kirschner MW . (1998). Development, 125, 3247–3258.
Labib K, Kearsey SE and Diffley JF . (2001). Mol. Biol. Cell, 12, 3658–3667.
Lei M and Tye BK . (2001). J. Cell Sci., 114, 1447–1454.
Maiorano D, Moreau J and Mechali M . (2000). Nature, 404, 622–625.
McGarry TJ . (2002). Mol. Biol. Cell, 13, 3662–3671.
McGarry TJ and Kirschner MW . (1998). Cell, 93, 1043–1053.
Mihaylov IS, Kondo T, Jones L, Ryzhikov S, Tanaka J, Zheng J, Higa LA, Minamino N, Cooley L and Zhang H . (2002). Mol. Cell. Biol., 22, 1868–1880.
Nishitani H, Lygerou Z, Nishimoto T and Nurse P . (2000). Nature, 404, 625–628.
Quinn LM, Herr A, McGarry TJ and Richardson H . (2001). Genes Dev., 15, 2741–2754.
Shieh SY, Ahn J, Tamai K, Taya Y and Prives C . (2000). Genes Dev., 14, 289–300.
Shreeram S, Sparks A, Lane DP and Blow JJ . (2002). Oncogene, 21, 6624–6632.
Tada S, Li A, Maiorano D, Mechali M and Blow JJ . (2001). Nat. Cell Biol., 3, 107–113.
Todorov IT, Attaran A and Kearsey SE . (1995). J Cell Biol., 129, 1433–1445.
Waldman T, Kinzler KW and Vogelstein B . (1995). Cancer Res., 55, 5187–5190.
Werness BA, Levine AJ and Howley PM . (1990). Science, 248, 76–79.
Wohlschlegel JA, Dwyer BT, Dhar SK, Cvetic C, Walter JC and Dutta A . (2000). Science, 290, 2309–2312.
Wohlschlegel JA, Kutok JL, Weng AP and Dutta A . (2002). Am. J. Pathol., 161, 267–273.
Yanagi K, Mizuno T, You Z and Hanaoka F . (2002). J. Biol. Chem., 277, 40871–40880.
Yates JL, Camiolo SM and Bashaw JM . (2000). J. Virol., 74, 4512–4522.
Zachariae W and Nasmyth K . (1999). Genes Dev., 13, 2039–2058.
Acknowledgements
We are grateful to Yoko Kanamori for technical assistance, Haruo Onoda for histochemical staining and Shigeo Mori for help with histology. We thank the members of our laboratory for their helpful discussions during the course of this work. This study was supported in part by a grant-in-aid from the Ministry of Education, Science, Sports and Culture, Japan.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Yoshida, K., Oyaizu, N., Dutta, A. et al. The destruction box of human Geminin is critical for proliferation and tumor growth in human colon cancer cells. Oncogene 23, 58–70 (2004). https://doi.org/10.1038/sj.onc.1206987
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1206987
Keywords
This article is cited by
-
The expanding genetic and clinical landscape associated with Meier-Gorlin syndrome
European Journal of Human Genetics (2023)
-
Human cytomegalovirus miR-US5-1 inhibits viral replication by targeting Geminin mRNA
Virologica Sinica (2017)
-
Control of DNA replication and its potential clinical exploitation
Nature Reviews Cancer (2005)
-
DNA replication licensing and cell cycle kinetics of normal and neoplastic breast
British Journal of Cancer (2005)