Abstract
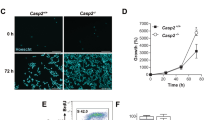
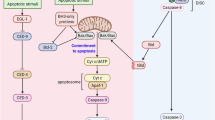
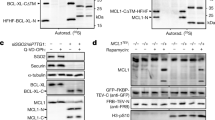
The activation of caspases is a critical event for the execution phase of programmed cell death. Caspases are highly specific in their ability to activate or inhibit many crucial proteins in the cell via cleavage. In this study, we report the identification of several caspase-3-like cleavage sites in the cell-cycle checkpoint protein Rad9. We demonstrate that human Rad9 can be specifically cleaved in cells induced to enter apoptosis by both DNA damage and staurosporine treatment. Indeed, we show that human Rad9 can be effectively cleaved both in vitro and in vivo, which can be inhibited by either a pan-caspase inhibitor or a caspase-3-specific inhibitor. Additionally, no cleavage of Rad9 can be seen in the caspase-3-deficient cell line MCF-7. Site-directed mutagenesis of three of the most conserved cleavage sites dramatically abrogates cleavage of Rad9 by caspase-3 in vitro, and in intact cells after DNA damage. Expression of the cleavage-resistant mutant Rad9 DDD/AAA appears to protect the cell from DNA damage-induced apoptosis. Immunofluorescence studies of Rad9 localization before and after induction of apoptosis show a translocation of Rad9 from the nucleus to the cytosol, concomitant to the appearance of apoptotic morphology. Furthermore, analysis of a truncated Rad9 mutant that corresponds to a putative N-terminal cleavage fragment shows that the N-terminal portion of Rad9 localizes in the cytosol, binds to Bcl-XL, and induces apoptosis. These results support a dual role for cleavage of Rad9: (1) the liberation and translocation of the BH3 domain-containing N-terminus of Rad9 to the cytosol, as a means of promoting apoptosis via antagonism of Bcl-XL, and (2) the disruption of the Rad9-Rad1-Hus1 DNA damage checkpoint complex.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Aravind L, Walker DR and Koonin EV . (1999). Nucleic Acids Res., 27, 1223–1242.
Burtelow MA, Roos-Mattjus PM, Rauen M, Babendure JR and Karnitz LM . (2001). J. Biol. Chem., 276, 25903–25909.
Caspari T and Carr AM . (2002). Curr. Biol., 12, R105–R107.
Chen MJ, Lin YT, Lieberman HB, Chen G and Lee EY . (2001). J. Biol. Chem., 276, 16580–16586.
Earnshaw WC, Martins LM and Kaufmann SH . (1999). Annu. Rev. Biochem., 68, 383–424.
Green DR and Reed JC . (1998). Science, 281, 1309–1312.
Hartwell LH and Kastan MB . (1994). Science, 266, 1821–1828.
Hayashi K, Kuniyasu H, Oue N, Shigeishi H, Kuraoka K, Nakayama H and Yasui W . (2002). Pathobiology, 70, 40–46.
Hirai I and Wang HG . (2002). J. Biol. Chem., 277, 25722–25727.
Komatsu K, Hopkins KM, Lieberman HB and Wang H-G . (2000a). FEBS Lett., 481, 122–126.
Komatsu K, Miyashita T, Hang H, Hopkins KM, Zheng W, Cuddeback S, Yamada M, Lieberman HB and Wang HG . (2000b). Nat. Cell Biol., 2, 1–6.
Nicholson DW . (1999). Cell Death Differ., 6, 1028–1042.
O'Connell MJ, Walworth NC and Carr AM . (2000). Trends Cell Biol., 10, 296–303.
Reed JC . (1998). Oncogene, 17, 3225–3236.
Salvesen GS and Dixit VM . (1997). Cell, 91, 443–446.
St Onge RP, Besley BD, Park M, Casselman R and Davey S . (2001). J. Biol. Chem., 276, 41898–41905.
St Onge RP, Udell CM, Casselman R and Davey S . (1999). Mol. Biol. Cell, 10, 1985–1995.
Stennicke HR and Salvesen GS . (1997). J. Biol. Chem., 272, 25719–25723.
Thornberry N, Rano T, Peterson E, Rasper D, Timkey T, Garcia-Calvo M, Houtzager V, Nordstrom P, Roy S, Vaillancourt J, Chapman K and Nicholson D . (1997). J. Biol. Chem., 272, 17907–17911.
Tsurimoto T . (1999). Front Biosci., 4, D849–D858.
Venclovas C and Thelen MP . (2000). Nucleic Acids Res., 28, 2481–2493.
Volkmer E and Karnitz LM . (1999). J. Biol. Chem., 274, 567–570.
Wahl GM and Carr AM . (2001). Nat. Cell Biol., 3, E277–E286.
Wang H-G and Reed JC . (1998). Histol. Histopathol., 13, 521–530.
Yoshida K, Komatsu K, Wang HG and Kufe D . (2002). Mol. Cell. Biol., 22, 3292–3300.
Zhou BB and Elledge SJ . (2000). Nature, 408, 433–439.
Zimmermann KC, Bonzon C and Green DR . (2001). Pharmacol. Ther., 92, 57–70.
Acknowledgements
We thank Dr Guy S Salvesen (The Burnham Institute) for kindly providing caspase-3 and -6 enzymes, and Dr Hirohito Yamaguchi for helpful discussion. This work was supported by grants CA82197 and CA90315 from the National Institutes of Health. Michael W Lee is a fellow of the American Heart Association.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lee, M., Hirai, I. & Wang, HG. Caspase-3-mediated cleavage of Rad9 during apoptosis. Oncogene 22, 6340–6346 (2003). https://doi.org/10.1038/sj.onc.1206729
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1206729
Keywords
This article is cited by
-
Rad9a is involved in chromatin decondensation and post-zygotic embryo development in mice
Cell Death & Differentiation (2019)
-
Pro-apoptotic cationic host defense peptides rich in lysine or arginine to reverse drug resistance by disrupting tumor cell membrane
Amino Acids (2017)
-
Decoding and unlocking the BCL-2 dependency of cancer cells
Nature Reviews Cancer (2013)
-
Far upstream element-binding protein-1, a novel caspase substrate, acts as a cross-talker between apoptosis and the c-myc oncogene
Oncogene (2009)
-
Loss of Hus1 sensitizes cells to etoposide-induced apoptosis by regulating BH3-only proteins
Oncogene (2008)