Abstract
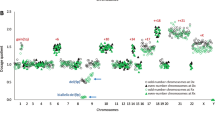
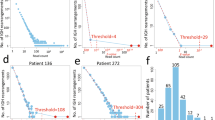
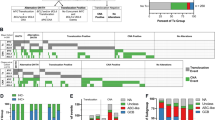
DNA amplifications are important mechanisms for proto-oncogene activation. Comparative genomic hybridization (CGH) to metaphase chromosome preparations has revealed amplifications in 10–20% of B-cell lymphomas (B-NHL). We analysed a series of 16 aggressive non-Hodgkin lymphomas by the new approach termed Matrix-CGH (M-CGH) using genomic DNA microarrays as hybridization target. For M-CGH, a dedicated B-cell lymphoma chip was constructed containing 496 genomic targets covering oncogenes, tumor suppressor genes as well as chromosome regions frequently altered in B-NHL. In 10 of 16 samples a total of 15 DNA amplifications were identified. The amplicons included BCL2, REL, CCND1, CCND2, JAK2, FGF4 and MDM2. Four of the 15 amplifications remained undetected by chromosomal CGH. The respective amplicons mapped to bands 2p13, 9p13–p21 and 12q24 and, were confirmed by fluorescence in situ hybridization. Furthermore, for four genomically amplified genes real-time quantitative reverse transcription polymerase chain reaction revealed elevated mRNA expression levels. These data show the superior diagnostic sensitivity of the newly developed diagnostic tool. As only a small portion of the genome (approximately 1.5%) has been analysed by the present DNA array, it is likely that gene amplifications are much more common in aggressive lymphomas than previously assumed.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Alizadeh AA, Eisen MB, Davis RE, Ma C, Lossos IS, Rosenwald A, Boldrick JC, Sabet H, Tran T, Yu X, Powell JI, Yang L, Marti GE, Moore T, Hudson Jr J, Lu L, Lewis DB, Tibshirani R, Sherlock G, Chan WC, Greiner TC, Weisenburger DD, Armitage JO, Warnke R and Staudt LM . (2000). Nature, 403, 503–511.
Barth TF, Bentz M, Leithauser F, Stilgenbauer S, Siebert R, Schlotter M, Schlenk RF, Dohner H and Moller P . (2001). Genes Chromosomes Cancer, 31, 316–325.
Bentz M, Plesch A, Stilgenbauer S, Döhner H and Lichter P . (1998). Genes Chromosomes Cancer, 21, 172–175.
Brodeur GM, Seeger RC, Schwab M, Varmus HE and Bishop JM . (1984). Science, 224, 1121–1124.
Du Manoir S, Speicher MR, Joos S, Schrock E, Popp S, Dohner H, Kovacs G, Robert-Nicoud M, Lichter P and Cremer T . (1993). Hum. Genet., 90, 590–610.
Gribben JG, Freedman A, Woo SD, Blake K, Shu RS, Freeman G, Longtine JA, Pinkus GS and Nadler LM . (1991). Blood, 78, 3275–3280.
Houldsworth J, Mathew S, Rao PH, Dyomina K, Louie DC, Parsa N, Offit K and Chaganti RS . (1996). Blood, 87, 25–29.
Jaffe ES, Harris NL, Stein H and Vardiman JW . (2001). World Health Organization Classification of Tumors. Pathology and Genetics of Tumors of the Hematopoetic and Lymphoid Tissues. IARC Press: Lyon, France.
Joos S, Otano-Joos MI, Ziegler S, Bruderlein S, du Manoir S, Bentz M, Moller P and Lichter P . (1996). Blood, 87, 1571–1578.
Kallioniemi A, Kallioniemi OP, Sudar D, Rutovitz D, Gray JW, Waldman F and Pinkel D . (1992). Science, 258, 818–821.
Kirchhoff M, Gerdes T, Maahr J, Rose H, Bentz M, Dohner H and Lundsteen C . (1999). Genes Chromosomes Cancer, 25, 410–413.
Klein CA, Schmidt-Kittler O, Schardt JA, Pantel K, Speicher MR and Riethmuller G . (1999). Proc. Natl. Acad. Sci., 96, 4494–4499.
Knuutila S, Bjorkqvist AM, Autio K, Tarkkanen M, Wolf M, Monni O, Szymanska J, Larramendy ML, Tapper J, Pere H, El-Rifai W, Hemmer S, Wasenius VM, Vidgren V and Zhu Y . (1998). Am. J. Pathol., 152, 1107–1123.
Korz C, Pscherer A, Benner A, Mertens D, Schaffner C, Leupolt E, Döhner H, Stilgenbauer S and Lichter P . (2002). Blood, 99, 4554–4561.
Lichter P, Bentz M and Joos S . (1995). Methods Enzymol., 254, 334–359.
Pinkel D, Segraves R, Sudar D, Clark S, Poole I, Kowbel D, Collins C, Kuo WL, Chen C, Zhai Y, Dairkee SH, Ljung BM, Gray JW and Albertson DG . (1998). Nat. Genet., 20, 207–211.
Pollack JR, Perou CM, Alizadeh AA, Eisen MB, Pergamenschikov A, Williams CF, Jeffrey SS, Botstein D and Brown PO . (1999). Nat. Genet., 23, 41–46.
Rao PH, Houldsworth J, Dyomina K, Parsa NZ, Cigudosa JC, Louie DC, Popplewell L, Offit K, Jhanwar SC and Chaganti RS . (1998) Blood, 92, 234–240.
Slamon DJ, Clark GM, Wong SG, Levin WJ, Ullrich A and McGuire WL . (1987). Science, 235, 177–182.
Rosenwald A, Wright G, Chan WC, Connors JM, Campo E, Fisher RI, Gascoyne RD, Muller-Hermelink HK, Smeland EB, Giltnane JM, Hurt EM, Zhao H, Averett L, Yang L, Wilson WH, Jaffe ES, Simon R, Klausner RD, Powell J, Duffey PL, Longo DL, Greiner TC, Weisenburger DD, Sanger WG, Dave BJ, Lynch JC, Vose J, Armitage JO, Montserrat E, Lopez-Guillermo A, Grogan TM, Miller TP, LeBlanc M, Ott G, Kvaloy S, Delabie J, Holte H, Krajci P, Stokke T and Staudt LM . (2002). N. Engl. J. Med., 346, 1937–1947.
Snijders AM, Nowak N, Segraves R, Blackwood S, Brown N, Conroy J, Hamilton G, Hindle AK, Huey B, Kimura K, Law S, Myambo K, Palmer J, Ylstra B, Yue JP, Gray JW, Jain AN, Pinkel D and Albertson DG . (2001). Nat. Genet., 29, 263–264.
Solinas-Toldo S, Lampel S, Stilgenbauer S, Nickolenko J, Benner A, Dohner H, Cremer T and Lichter P . (1997). Genes Chromosomes Cancer, 20, 399–407.
Wessendorf S, Fritz B, Wrobel G, Nessling M, Lampel S, Goettel D, Kuepper M, Joos S, Hopman T, Kokocinski F, Dohner H, Bentz M, Schwaenen C and Lichter P . (2002). Lab. Invest., 82, 47–60.
Werner CA, Dohner H, Joos S, Trumper LH, Baudis M, Barth TF, Ott G, Moller P, Lichter P and Bentz M . (1997). Am. J. Pathol., 151, 335–342.
Acknowledgements
The technical support and sharing of material by Daniel Göttel, Felix Kokocinski, Björn Fritz, Manfred Küpper, Jeanette Dörfel, Tatjana Salvi, Andrea Wittmann and Martina Enz is gratefully acknowledged.
This work was supported by grants of the Deutsche Forschungsgemeinschaft (SFB 518), the Deutsche Krebshilfe (Grant no. 70-2840-Be3, 70-2859-Ba2), the IZKF Ulm (Project C12) and the Bundesministerium für Bildung und Forschung (BMBF Project 01KW9936-9938).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wessendorf, S., Schwaenen, C., Kohlhammer, H. et al. Hidden gene amplifications in aggressive B-cell non-Hodgkin lymphomas detected by microarray-based comparative genomic hybridization. Oncogene 22, 1425–1429 (2003). https://doi.org/10.1038/sj.onc.1206297
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1206297
Keywords
This article is cited by
-
Sensitivity and resistance towards isoliquiritigenin, doxorubicin and methotrexate in T cell acute lymphoblastic leukaemia cell lines by pharmacogenomics
Naunyn-Schmiedeberg's Archives of Pharmacology (2010)
-
Sub-megabase resolution tiling (SMRT) array-based comparative genomic hybridization profiling reveals novel gains and losses of chromosomal regions in Hodgkin Lymphoma and Anaplastic Large Cell Lymphoma cell lines
Molecular Cancer (2008)
-
Further delineation of chromosomal consensus regions in primary mediastinal B-cell lymphomas: an analysis of 37 tumor samples using high-resolution genomic profiling (array-CGH)
Leukemia (2007)
-
The use of transcriptomic biomarkers for personalized medicine
Heart Failure Reviews (2007)
-
Mutations in the NF-κB signaling pathway: implications for human disease
Oncogene (2006)