Abstract
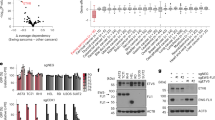
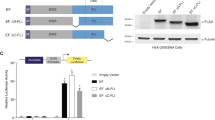
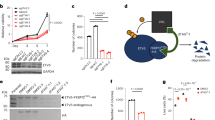
Ewing tumour is characterized by specific chromosome translocations which fuse EWS to a subset of genes encoding ETS transcription factors, most frequently FLI-1. We report the analysis of the expression of various cell cycle regulators both in Ewing tumour derived cell lines and in different cellular models with either inducible or constitutive EWS-FLI-1 cDNA expression. In Ewing cell lines, cyclin D1, CDK4, Rb, p27KIP1 and c-Myc were consistently highly expressed whereas p57KIP2, p15INK4B and p14ARF demonstrated undetectable or low expression levels. The amount of p16INK4A, p21CIP1, p18INKAC and CDK6 was variable from one cell line to the other. The inducible expression of EWS-FLI-1 led to a strong upregulation of c-Myc and a considerable downregulation of p57KIP2. Other proteins did not show evident modification. High c-Myc and very low p57KIP2 expression levels were also observed in neuroblastoma NGP cells constitutively expressing EWS-FLI-1 as compared to parental cells. Analysis of the p57KIP2 promoter indicated that EWS-FLI-1 downregulates, possibly through an indirect mechanism, the transcription of this gene. Finally, we show that ectopic expression of p57KIP2 in Ewing cells blocks proliferation through a complete G1 arrest. These results suggest that the modulation of p57KIP2 expression by EWS-FLI-1 is a fundamental step in Ewing tumorigenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Alexandrow MG, Moses HL . 1995 Cancer Res. 55: 1452–1457
Arvand A, Bastians H, Welford SM, Thompson AD, Ruderman JV, Denny CT . 1998 Oncogene 17: 2039–2045
Bailly RM, Bosselut R, Zucman J, Cormie F, Delattre O, Roussel M, Thomas G, Ghysdael J . 1994 Mol. Cell. Biol. 14: 3230–3241
Delattre O, Zucman J, Plougastel B, Desmaze C, Melot T, Peter M, Kovar H, Joubert I, de Jong P, Rouleau G, Aurias A, Thomas G . 1992 Nature 359: 162–165
Feinberg AP . 1999 Cancer Res. 59: 1743s–1746s
Gartel AL, Tyner AL . 1999 Exp. Cell. Res. 246: 280–289
Gil-Diez de Medina S, Salomon L, Colombel M, Abbou CC, Bellot J, Thiery JP, Radvanyi F, Van der Kwast TH, Chopin DK . 1998 Hum. Pathol. 29: 1005–1102
Hahm KB, Cho K, Lee C, Im YH, Chang J, Choi SG, Sorensen PH, Thiele CJ, Kim SJ . 1999 Nat. Genet. 23: 2222–2227
Hamelin R, Zucman J, Melot T, Delattre O, Thomas G . 1994 Int. J. Cancer 57: 336–340
Hashimoto Y, Kohri K, Kaneko Y, Morisaki H, Kato T, Ikeda K, Nakanishi M . 1998 J. Biol. Chem. 273: 16544–16550
Hatada I, Inazawa J, Abe T, Nakayama M, Kaneko Y, Jinno Y, Niikawa N, Ohashi H, Fukushima Y, Iida K, Yutani C, Takahashi S, Chiba Y, Ohishi S, Mukai T . 1996a Hum. Mol. Genet. 5: 783–788
Hatada I, Ohashi H, Fukushima Y, Kaneko Y, Inoue M, Komoto Y, Okada A, Ohishi S, Nabetani A, Morisaki H, Nakayama M, Niikawa N, Mukai T . 1996b Nat. Genet. 14: 171–173
Hermeking H, Rago C, Schuhmacher M, Li Q, Barrett JF, Obaya AJ, O'Connell BC, Mateyak MK, Tam W, Kohlhuber F, Dang CV, Sedivy JM, Eick D, Vogelstein B, Kinzler KW . 2000 Proc. Natl. Acad. Sci. USA 97: 2229–2234
Im YH, Kim HT, Lee C, Poulin D, Welford S, Sorensen PH, Denny CT, Kim SJ . 2000 Cancer Res. 60: 1536–1540
Jaishankar S, Zhang J, Roussel MF, Baker SJ . 1999 Oncogene 18: 5592–5597
Komuro H, Hayashi Y, Kawamura M, Hayashi K, Kaneko Y, Kamoshita S, Hanada R, Yamamoto K, Hongo T, Yamada M, Tsuchida Y . 1993 Cancer Res. 53: 5284–5288
Kovar H, Auinger A, Jug G, Aryee D, Zoubek A, Salzer-Kuntschik M, Gadner H . 1993 Oncogene 8: 2683–2690
Lasorella A, Noseda M, Beyna M, Iavarone A . 2000 Nature 5: 592–598
Lee MH, Reynisdottir I, Massague J . 1995 Genes Dev. 9: 639–649
Lessnick SL, Brau BS, Denny CT, May WA . 1995 Oncogene 10: 423–431
Lin J, Reichner C, Wu X, Levine AJ . 1996 Mol. Cell. Biol. 16: 1786–1793
Mao X, Miesfeldt S, Yang H, Leiden JM, Thompson CB . 1994 J. Biol. Chem. 269: 18216–18222
Matsuoka S, Edwards MC, Bai C, Parker S, Zhang P, Baldini A, Harper JW, Elledge SJ . 1995 Genes Dev. 9: 650–662
Matsuoka S, Thompson JS, Edwards MC, Bartletta JM, Grundy P, Kalikin LM, Harper JW, Elledge SJ, Feinberg AP . 1996 Proc. Natl. Acad. Sci. USA 93: 3026–3030
May WA, Arvand A, Thompson AD, Braun BS, Wright M, Denny CT . 1997 Nat. Genet. 17: 495–497
May WA, Lessnick SL, Braun BS, Klemsz M, Lewis BC, Lunsford LB, Hromas R, Denny CT . 1993 Mol. Cell. Biol. 13: 7393–7398
Melot T, Gruel N, Doubeikovski A, Sevenet N, Teillaud JL, Delattre O . 1997 Hybridoma 165: 457–464
Peter M, Magdelenat H, Michon J, Melot T, Oberlin O, Zucker JM, Thomas G, Delattre O . 1995 Br. J. Cancer 72: 96–100
Reid LH, Crider-Miller SJ, West A, Lee MH, Massague J, Weissman BE . 1996 Cancer Res. 56: 1214–1218
Remvikos Y, Vielh P, Padoy E, Benyahia B, Voillemot N, Magdelenat H . 1991 Br. J. Cancer 64: 501–507
Reynaud EG, Pelpel K, Guillier M, Leibovitch MP, Leibovitch SA . 1999 Mol. Cell. Biol. 19: 7621–7629
Sumarsono SH, Wilson TJ, Tymms MJ, Venter DJ, Corrick CM, Kola R, Lahoud MH, Papas TS, Seth A, Kola I . 1996 Nature 379: 534–537
Thompson AD, Braun BS, Arvand A, Stewart SD, May WA, Chen E, Korenberg J, Denny CT . 1996 Oncogene 13: 2649–2658
Tokino T, Urano T, Furuhata T, Matsushima M, Miyatsu T, Sasaki S, Nakamura Y . 1996 Hum. Genet. 97: 625–631
Xing EP, Nie Y, Song Y, Yang GY, Cai YC, Wang LD, Yang CS . 1999 Clin. Cancer Res. 510: 2704–2713
Acknowledgements
We thank Ismail Kola, Christopher Denny and Serge Leibovitch for providing the pMT-CB6+ vector, the EWS-FLI-1 retroviral vector, the GFP-p57KIP2 and the p57KIP2 vectors, respectively. We also thank Danny Rouillard for her help in cell cycle analysis, Catherine Barbarou and Patricia de Cremoux for the analysis of p53. L Dauphinot is a recipient of a fellowship from the Ligue Nationale Contre le Cancer. This work was supported in part by grants from the Association pour la Recherche contre le Cancer, the Ligue Nationale Contre le Cancer and the USPHS, CA63176.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Dauphinot, L., De Oliveira, C., Melot, T. et al. Analysis of the expression of cell cycle regulators in Ewing cell lines: EWS-FLI-1 modulates p57KIP2 and c-Myc expression. Oncogene 20, 3258–3265 (2001). https://doi.org/10.1038/sj.onc.1204437
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1204437
Keywords
This article is cited by
-
TBX3 promotes proliferation of papillary thyroid carcinoma cells through facilitating PRC2-mediated p57KIP2 repression
Oncogene (2018)
-
Design of a Lentiviral Vector for the Inducible Expression of MYC: A New Strategy for Construction Approach
Molecular Biotechnology (2017)
-
CIC-DUX sarcomas demonstrate frequent MYC amplification and ETS-family transcription factor expression
Modern Pathology (2015)
-
Molecular detection and targeting of EWSR1 fusion transcripts in soft tissue tumors
Medical Oncology (2013)
-
Molecular pathogenesis and targeted therapeutics in Ewing sarcoma/primitive neuroectodermal tumours
Clinical Sarcoma Research (2012)