Abstract
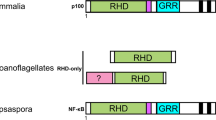
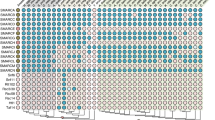
From the sequences of Rel/NF-κB and IκB proteins, we constructed an alignment of their Rel Homology Domain (RHD) and ankyrin repeat domain. Using this alignment, we performed tree reconstruction with both distance matrix and parsimony analysis and estimated the branching robustness using bootstrap resampling methods. We defined four subfamilies of Rel/NF-κB transcription factors: (i) cRel, RelA, RelB, Dorsal and Dif; (ii) NF-κB1 and NF-κB2; (iii) Relish and (iv) NF-AT factors, the most divergent members. Subfamilies I and II are clustered together whereas Relish diverged earlier than other Rel/NF-κB proteins. Three subfamilies of IκB inhibitors were also defined: (i) NF-κB1 and NF-κB2; (ii) close to subfamily I, the short IκB proteins IκBα, IκBβ and Bcl-3; (iii) Relish that diverged earlier than other IκB inhibitors. Our definition of groups and subfamilies fits to structural and functional features of the Rel/NF-κB and IκB proteins. We also showed that ankyrin repeats of NF-κB1, NF-κB2 and Relish are short IκB-specific ankyrin motifs. These proteins defining a link between Rel/NF-κB and IκB families, we propose that all these factors evolved from a common ancestral RHD-ankyrin structure within a unique superfamily, explaining the specificities of interaction between the different Rel/NF-κB dimers and the various IκB inhibitors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Huguet, C., Crepieux, P. & Laudet, V. Rel/NF-κB transcription factors and IκB inhibitors: Evolution from a unique common ancestor. Oncogene 15, 2965–2974 (1997). https://doi.org/10.1038/sj.onc.1201471
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1201471
Keywords
This article is cited by
-
G3BP2 is involved in isoproterenol-induced cardiac hypertrophy through activating the NF-κB signaling pathway
Acta Pharmacologica Sinica (2018)
-
Transcription factors involved in basal immunity in mammals and plants interact with the same MAMP-responsive cis-sequence from Arabidopsis thaliana
Plant Molecular Biology (2018)
-
Transcription factor NF-κB is modulated by symbiotic status in a sea anemone model of cnidarian bleaching
Scientific Reports (2017)
-
The transcription factor NF-κB in the demosponge Amphimedon queenslandica: insights on the evolutionary origin of the Rel homology domain
Development Genes and Evolution (2008)
-
Rel homology domain-containing transcription factors in the cnidarian Nematostella vectensis
Development Genes and Evolution (2007)