Abstract
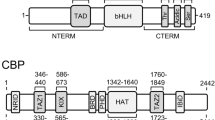
The Microphthalmia basic-Helix – Loop – Helix-Leucine Zipper (bHLH-LZ) transcription factor plays a crucial role in the genesis of melanocytes; mice deficient for a functional microphthalmia gene product lack all pigment cells. We show here that the Mi activation domain resides N-terminal to the DNA-binding domain and that as little as 18 amino acids are sufficient to mediate transcription activation. The minimal activation region of Mi is highly conserved in the related transcription factor TFE3 and is predicted to adopt an amphipathic alpha-helical conformation. This region of Mi is also highly conserved with a region of E1A known to be essential for binding the CBP/p300 transcription cofactor. Consistent with these observations, the Mi activation domain can interact in vitro with CBP specifically through a region of CBP required for complex formation with E1A, P/CAF and c-Fos, and anti p300 antibodies can co-immunoprecipitate Mi from both melanocyte and melanoma cell lines. In addition, co-transfection of a vector expressing CBP2 fused to the VP16 activation domain potentiated the ability of Mi to activate transcription, confirming the significance of the CBP-Mi interaction observed in vitro. These data suggest that transcription activation by Mi is achieved at least in part by recruitment of CBP. The parallels between transcription regulation by Microphthalmia in melanocytes and MyoD in muscle cells are discussed.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Sato, S., Roberts, K., Gambino, G. et al. CBP/p300 as a co-factor for the Microphthalmia transcription factor. Oncogene 14, 3083–3092 (1997). https://doi.org/10.1038/sj.onc.1201298
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1201298
Keywords
This article is cited by
-
Restrained Mitf-associated autophagy by Mulberroside A ameliorates osteoclastogenesis and counteracts OVX-Induced osteoporosis in mice
Cell Death Discovery (2024)
-
Acetylation reprograms MITF target selectivity and residence time
Nature Communications (2023)
-
The C-terminal transactivation domain of MITF interacts promiscuously with co-activator CBP/p300
Scientific Reports (2023)
-
Molecular mechanisms of syncytin-1 in tumors and placental development related diseases
Discover Oncology (2023)
-
Past, present, and future perspectives of transcription factor EB (TFEB): mechanisms of regulation and association with disease
Cell Death & Differentiation (2022)