Abstract
OBJECTIVE: To investigate whether the C-60G polymorphism and other markers in the hormone-sensitive lipase (LIPE) gene are associated with baseline body composition and free-fatty acid (FFA) concentrations measured at rest and during low-intensity exercise in white and black subjects participating in the HERITAGE Family Study.
SUBJECTS: Adult sedentary white (245 men and 258 women) and black (91 men and 185 women) subjects.
MEASUREMENTS: body mass index (BMI); fat mass (FAT); percentage body fat (%FAT); fat-free mass (FATFR); sum of eight skinfolds (SF8); subcutaneous (ASF), visceral (AVF) and total (ATF) abdominal fat areas assessed by CT scan; plasma FFA concentrations measured at rest (FFAR), at a power output of 50 W (FFA50) and at a relative power output of 60% of VO2max (FFA60%); and fasting insulin (INS).
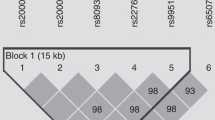
STATISTICAL ANALYSIS: Association between the C-60G polymorphism of the LIPE gene and each phenotype was tested separately in men and women using ANCOVA with the effects of age and race as covariates and with further adjustment for FAT for ASF, AVF, ATF, FFAR, FFA50 and FFA60%. Secondly, owing to significant gene-by-race interaction, associations were investigated separately in each of the two race groups. Linkage was tested with the C-60G polymorphism, a dinucleotide repeat polymorphism in the intron 7 of the LIPE gene and two microsatellites markers (D19S178 and D19S903) flanking the LIPE gene.
RESULTS: There were no race differences in the allele frequencies of the C-60G polymorphism of the LIPE gene. No association or gene-by-race interaction was observed in men. However, in women, strong gene-by-race interactions were observed for BMI (P=0.0005), FAT (P=0.0007), %FAT (P=0.0003), SF8 (P=0.0001), ASF (P=0.03) and ATF (P=0.01). When the analysis was performed separately in each race, white women carriers of the -60G allele exhibited lower %FAT (P=0.005) and SF8 (P=0.01) than non-carriers, while in black women, the -60G allele was associated with higher BMI (P=0.004), FAT (P=0.009), %FAT (P=0.01) and SF8 (P=0.0009). These associations were no longer significant after adjusting for INS. Evidence of linkage was observed in whites with ATF, FFAR, FFA50 and FFA60%.
CONCLUSION: These results suggest that the C-60G polymorphism in the LIPE gene plays a role in determining body composition and that its effect is sex-, race- and insulin-dependent.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Langin D, Holm C, Lafontan M . Adipocyte hormone-sensitive lipase: a major regulator of lipid metabolism Proc Nutr Soc 1996 55: 93–109.
Carey GB . Mechanisms regulating adipocyte lipolysis Adv Exp Med Biol 1998 441: 157–170.
Large V, Arner P . Regulation of lipolysis in humans. Pathophysiological modulation in obesity, diabetes, and hyperlipidaemia Diabetes Metab 1998 24: 409–418.
Arner P . Impact of exercise on adipose tissue metabolism in humans Int J Obes Relat Metab Disord 1995 19(Suppl 4): S18–21.
Romijn JA, Coyle EF, Sidossis LS, Gastaldelli A, Horowitz JF, Endert E, Wolfe RR . Regulation of endogenous fat and carbohydrate metabolism in relation to exercise intensity and duration Am J Physiol 1993 265: E380–391.
Hellström L, Langin D, Reynisdottir S, Dauzats M, Arner P . Adipocyte lipolysis in normal weight subjects with obesity among first-degree relatives Diabetologia 1996 39: 921–928.
Large V, Arner P, Reynisdottir S, Grober J, Van Harmelen V, Holm C, Langin D . Hormone-sensitive lipase expression and activity in relation to lipolysis in human fat cells J Lipid Res 1998 39: 1688–1695.
Large V, Reynisdottir S, Langin D, Fredby K, Klannemark M, Holm C, Arner P . Decreased expression and function of adipocyte hormone-sensitive lipase in subcutaneous fat cells of obese subjects J Lipid Res 1999 40: 2059–2066.
Laurell H, Grober J, Holst LS, Holm C, Mohrenweiser HW, Langin D . The hormone-sensitive lipase (LIPE) gene located on chromosome 19q13.1→13.2 is not duplicated on 19p13.3 Int J Obes Relat Metab Disord 1995 19: 590–592.
Levitt RC, Liu Z, Nouri N, Meyers DA, Brandriff B, Mohrenweiser HM . Mapping of the gene for hormone sensitive lipase (LIPE) to chromosome 19q13.1→q13.2 Cytogenet Cell Genet 1995 69: 211–214.
Shimada F, Makino H, Hashimoto N, Iwaoka H, Taira M, Nozaki O, Kanatsuka A, Holm C, Langin D, Saito Y . Detection of an amino acid polymorphism in hormone-sensitive lipase in Japanese subjects Metabolism 1996 45: 862–864.
Levitt RC, Jedlicka AE, Nouri N . Dinucleotide repeat polymorphism at the hormone sensitive lipase (LIPE) locus Hum Mol Genet 1992 1: 139.
Klannemark M, Orho M, Langin D, Laurell H, Holm C, Reynisdottir S, Arner P, Groop L . The putative role of the hormone-sensitive lipase gene in the pathogenesis of type II diabetes mellitus and abdominal obesity Diabetologia 1998 41: 1516–1522.
Pajukanta P, Porkka KV, Antikainen M, Taskinen MR, Perola M, Murtomaki-Repo S, Ehnholm S, Nuotio I, Suurinkeroinen L, Lahdenkari AT, Syvanen AC, Viikari JS, Ehnholm C, Peltonen L . No evidence of linkage between familial combined hyperlipidemia and genes encoding lipolytic enzymes in Finnish families Arterioscler Thromb Vasc Biol 1997 17: 841–850.
Austin MA, Talmud PJ, Luong LA, Haddad L, Day IN, Newman B, Edwards KL, Krauss RM, Humphries SE . Candidate-gene studies of the atherogenic lipoprotein phenotype: a sib-pair linkage analysis of DZ women twins Am J Hum Genet 1998 62: 406–419.
Magre J, Laurell H, Fizames C, Antoine PJ, Dib C, Vigouroux C, Bourut C, Capeau J, Weissenbach J, Langin D . Human hormone-sensitive lipase: genetic mapping, identification of a new dinucleotide repeat, and association with obesity and NIDDM Diabetes 1998 47: 284–286.
Clément K, Dina C, Basdevant A, Chastang N, Pelloux V, Lahlou N, Berlan M, Langin D, Guy-Grand B, Froguel P . A sib-pair analysis study of 15 candidate genes in French families with morbid obesity: indication for linkage with islet 1 locus on chromosome 5q Diabetes 1999 48: 398–402.
Talmud PJ, Palmen J, Walker M . Identification of genetic variation in the human hormone-sensitive lipase gene and 5′ sequences: homology of 5′ sequences with mouse promoter and identification of potential regulatory elements Biochem Biophys Res Commun 1998 252: 661–668.
Bouchard C, Leon AS, Rao DC, Skinner JS, Wilmore JH, Gagnon J . The HERITAGE family study. Aims, design, and measurement protocol Med Sci Sports Exerc 1995 27: 721–729.
Wilmore JH, Stanforth PR, Domenick MA, Gagnon J, Daw EW, Leon AS, Rao DC, Skinner JS, Bouchard C . Reproducibility of anthropometric and body composition measurements: the Heritage Family Study Int J Obes Relat Metab Disord 1997 21: 297–303.
Ferland M, Després JP, Tremblay A, Pinault S, Nadeau A, Moorjani S, Lupien PJ, Thériault G, Bouchard C . Assessment of adipose tissue distribution by computed axial tomography in obese women: association with body density and anthropometric measurements Br J Nutr 1989 61: 139–148.
Wilmore JH, Stanforth PR, Turley KR, Gagnon J, Daw EW, Leon AS, Rao DC, Skinner JS, Bouchard C . Reproducibility of cardiovascular, respiratory, and metabolic responses to submaximal exercise: the Heritage Family Study Med Sci Sports Exerc 1998 30: 259–265.
Desbuquois B, Aurbach GD . Use of polyethylene glycol to separate free and antibody-bound peptide hormones in radioimmunoassays J Clin Endocrinol Metab 1971 33: 732–738.
SAS Statistical analysis, V. 6.12 SAS Institute Inc.: Cary, North Carolina 1996.
SAGE . Statistical analysis for genetic epidemiology. Computer program package available from the Department of Biometry and Genetics, release 3.1 Edn. LSU Medical Center: New Orleans, LA 1997.
Ott J . Analysis of human genetic linkage The John Hopkins University Press: Baltimore, MD 1991.
Grober J, Laurell H, Blaise R, Fabry B, Schaak S, Holm C, Langin D . Characterization of the promoter of human adipocyte hormone-sensitive lipase Biochem J 1997 328: 453–461.
Reynisdottir S, Dauzats M, Thörne A, Langin D . Comparison of hormone-sensitive lipase activity in visceral and subcutaneous human adipose tissue J Clin Endocrinol Metab 1997 82: 4162–4166.
Lefebvre AM, Laville M, Vega N, Riou JP, van Gaal L, Auwerx J, Vidal H . Depot-specific differences in adipose tissue gene expression in lean and obese subjects Diabetes 1998 47: 98–103.
Langfort J, Ploug T, Ihlemann J, Enevoldsen LH, Stallknecht B, Saldo M, Kjaer M, Holm C, Galbo H . Hormone-sensitive lipase (HSL) expression and regulation in skeletal muscle Adv Exp Med Biol 1998 441: 219–228.
Boden G . Role of fatty acids in the pathogenesis of insulin resistance and NIDDM Diabetes 1997 46: 3–10.
Jiang X, Srinivasan SR, Radhakrishnamurthy B, Dalferes ER, Berenson GS . Racial (black–white) differences in insulin secretion and clearance in adolescents: the Bogalusa heart study Pediatrics 1996 97: 357–360.
Acknowledgements
The HERITAGE Family Study is supported by NHLBI through grants HL47323 (AS Leon), HL47317 (DC Rao), HL47327 (JS Skinner), HL47321 (JH Wilmore) and HL45670 (C Bouchard). Arthur S Leon is partially supported by the Henry L Taylor endowed Professorship in Exercise Science and Health Enhancement. Claude Bouchard is partially supported by the George A Bray Chair in Nutrition. Some of the results of this study were obtained with the program SAGE whose development was supported by the US Public Health Service Research Grant (1P41RR03655) from the National Center for Research Resources. Thanks are expressed to all the investigators, local project coordinators, research assistants, laboratory technicians and secretaries who have contributed to this study.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Garenc, C., Pérusse, L., Chagnon, Y. et al. The hormone-sensitive lipase gene and body composition: the HERITAGE Family Study. Int J Obes 26, 220–227 (2002). https://doi.org/10.1038/sj.ijo.0801872
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ijo.0801872
Keywords
This article is cited by
-
A QTL for Genotype by Sex Interaction for Anthropometric Measurements in Alaskan Eskimos (GOCADAN Study) on Chromosome 19q12–13
Obesity (2011)
-
LIPE C-60G influences the effects of physical activity on body fat and plasma lipid concentrations: the Quebec Family Study
Human Genomics (2009)
-
Obesity and polymorphisms in genes regulating human adipose tissue
International Journal of Obesity (2007)
-
The combination of ApoCIII, hepatic lipase and hormono sensitive lipase gene polymorphisms suggests an association with susceptibility to gestational hypertension
Journal of Human Genetics (2007)
-
The hormone-sensitive lipase C–60G promoter polymorphism is associated with increased waist circumference in normal-weight subjects
International Journal of Obesity (2006)