Abstract
In line with the complexity of disease networks, diverse combination therapies have been demonstrated potential in the treatment of different patients with complex diseases in a personal combination profile. However, the identification of rational, compatible and effective drug combinations remains an ongoing challenge. Based on a holistic theory integrated with reductionism, Fangjiomics systematically develops multiple modes of array-designed combination therapies. We define diverse “magic shotgun” vertical, horizontal, focusing, siege and dynamic arrays according to different spatiotemporal distributions of hits on targets, pathways and networks. Through these multiple adaptive modes for treating complex diseases, Fangjiomics may help to identify rational drug combinations with synergistic or additive efficacy but reduced adverse side effects that reverse complex diseases by reconstructing or rewiring multiple targets, pathways and networks. Such a novel paradigm for combination therapies may allow us to achieve more precise treatments by developing phenotype-driven quantitative multi-scale modeling for rational drug combinations.
Similar content being viewed by others
Introduction
Growing evidence suggests that many human diseases cannot be attributed to the dysfunction of a single factor or genetic variation but instead arise due to complex interactions among a multitude of genetic mutations, polymorphisms, and environmental factors1. Of the 25 000 genes in the human genome, approximately 1800 are known to be involved in the causes of various diseases, including cancer, hypertension, ulcer, etc2,3. In spite of great advances in the treatment of certain diseases over past decades, novel target therapies such as p53-, NF-κB-, and epidermal growth factor receptor (EGFR)-targeted treatments for complex diseases have encountered with failure than success3,4. Thus, changes are needed to overcome the challenges in the healthcare and pharmaceutical industries; for example, the high costs of healthcare, low effectiveness of drugs, and high incidence of adverse drug reactions (ADRs)5. Some combination therapies have more proven effective than single drug therapies for complex diseases, including malignant cancer6,7, vascular diseases8, chronic obstructive pulmonary disease9, etc. Nevertheless, “more is not always better”10. For instance, long-term dual-antiplatelet therapy was reported to be linked with a higher risk for intracerebral hemorrhage (ICH) than clopidogrel monotherapy in the ischemic stroke population but provided no increased benefit in overall recurrent stroke risk reduction11; the combination of two but not more than three current targeted drugs was demonstrated to improve therapy of chronic myeloid leukemia12. Therefore, the optimization of combination therapies plays a key role in the improvement of the effectiveness and safety of treatments for complex diseases.
The traditional strategy to design combination therapy in clinical practice is to empirically combine agents with validated clinical efficacy13. The combination effects of these multiple agents are usually evaluated using mathematical models such as the Bliss independence model14, the Loewe additivism model15, and the Combination Index theorem16. However, this strategy can only achieve success in a case-by-case approach. Combination drugs manifest their therapeutic activities by modulating multi-targets, but these multi-target interactions are either largely unknown or insufficiently understood in most cases17,18,19. Furthermore, uncovering effective drug combinations by direct screening may be an impossible mission due to the exceedingly high number of potential combinations. In contrast, with the wealth of data available from molecular studies on complex diseases, especially the large-scale generation and integration of “omics” profile data, including genomic, proteomic, signaling, metabolomic, phenomics data, etc, we can now rationalize novel combinations using computational methods. Recently, the computational methods of evaluating the effects of combination therapies have mainly focused on two approaches20. The first is to identify and optimize multiple target interventions to solve small-scale problems by modeling signaling pathways or specific processes21,22, and the second is to evaluate the efficacy of multi-target drugs by using network biology approaches3,23. Nevertheless, the association between drug activity and network properties is not precisely understood. Thus, novel systematic approaches are urgently required for the feasible and efficient identification of rational combinations.
Fangjiomics is an emerging science for the design, production, and evaluation of combination therapy rationally selected from diverse agents in the holistic treatment of complex diseases24. In contrast to traditional “omics” techniques focusing on a certain level of cell, tissue, or organ, this holistic therapeutic strategy uses rational drug combinations with higher efficacy but fewer adverse effects in a controlled array design by integrating diverse scale omics data on gene, protein, and metabolic interactions. This minireview focuses on the key issues concerning this novel holistic array-designed paradigm for rational drug combinations based on the philosophy of Fangjiomics, which can be applied in clinical practice beyond network medicine.
Array design based on the philosophy of Fangjiomics
In the millennia-old history of Chinese medicine, a certain type of combination therapy with an array-designed mode, which was termed “Fangji”, was the essential therapeutic method for diverse diseases. Thus far, more than tens of thousands of Fangjis have been recorded in different books24. Fangjiomics is the large-scale study of combination therapies, including their combination modes, the drug-drug interactions involved in the combinations, the mechanism of the pharmacological actions of the combinations, etc. As discussed in our previous paper, the philosophy of Fangjiomics is distinct from conventional approaches in the aspects of theory, hypothesis, objective, and subject pharmacology, as well as in the process24. This therapeutic strategy targets multi-scale biological network levels25, including the molecular, cellular, tissue, and organ levels, which are related to clinical outcomes as profiled effects based on a holistic theory integrated with reductionism. This strategy supports patient-centered care based on the integration of experimental and clinical pharmacological mechanisms, which would treat patients with abnormal conditions more precisely and more effectively.
Based on the holistic theory integrated with reductionism, Fangjiomics systematically develops multiple modes of array-designed combination therapies. Array design is the application of multiple agents that regulate multiple targets, pathways, or networks in a certain sequence according to their pharmacodynamic effects on certain physiological or pathophysiological states. Each array should consist of multiple compatible ingredients with polypharmacologically profiled effects. Different contributing agents play diverse pharmacological roles in the combination therapy. For example, Realgar-Indigo naturalis formula has been identified to be very effective in treating acute promyelocytic leukemia (APL) in clinical practice26. This formula contains multiple ingredients, and the main ingredients include tetraarsenic tetrasulfide (A), indirubin (I), and tanshinone IIA (T). These three ingredients, by playing different pharmacological roles in array design (mainly as a vertical array, which is discussed below), together exert a synergistic effect for the treatment of APL. The promyelocytic leukemia (PML)-retinoic acid receptor-α (RAR-α) oncoprotein is the causative oncoprotein of APL. Compound A, which is considered to be the principal component in this formula, directly affects this oncoprotein and induces APL cell differentiation, thus playing the primary pharmacological effect of the combination. Compound I and T synergistically enhance A-triggered relocalization and the ubiquitination of the oncoprotein, thus vertically affecting and leading to the degradation of PML-RAR-α. Compound I and T can also significantly up-regulate the expression level of the transmembrane protein AQP9, the key transporter for uptaking arsenic and determining cellular arsenic sensitivity, to facilitate the delivery of the compound A to APL cells, thus enhancing the formula's treatment effect. Therefore, compound I and T function as adjuvant ingredients in this combination.
Diverse array-designed modes of combination therapies in Fangjiomics
Combination therapy is a promising strategy for combating complex disorders due to its improved efficacy and reduced side effects. However, the exhaustive screening of rational drug combinations is impractical given all of the possible combinations between drugs. Based on an array-designed mode, Fangjiomics-based combination therapy could effectively use rational drug combinations to treat complex diseases at diverse scales of biological organization. We will discuss 6 common array-designed modes of combination therapies in Fangjiomics below.
Hitting on multi-targets in a “magic shotgun” array
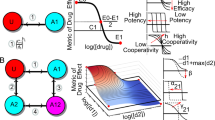
In Fangjiomics-based combination therapy, a "magic shotgun" array aims to sift through the known universe of chemicals to find the few special molecules that can broadly disrupt an entire disease process27. Drug combinations in a “magic shotgun” array will selectively hit multiple targets to effectively treat malaria28, schizophrenia29, cancer30, epilepsy31, and central nervous system disorders32, rather than previous seeking to find the “magic bullet”, chemicals which specifically attack one gene or protein involved in one particular part of a disease process27. Combining kinase focused chemistry, kinome-wide profiling and Drosophila genetics, a “magic shotgun” array identified the on-targets RET, RAF, SRC, S6K, and anti-target mTOR for the treatment of cancer, and this combination provided high efficacy and few adverse reactions, with a maximal therapeutic index30 (Figure 1A). A comparative analysis of stroke-related gene expression profiles among jasminoidin monotherapy, ursodeoxycholic acid monotherapy, and their combination demonstrated that the combination treatment exerted a synergistic effect in the “magic shotgun” array by up-regulating the expression of the genes Hspa1a, Fgf12, Rara, Map3k4, and down-regulating the PXN gene33. Moreover, some computational algorithms have been used to identify effective drug targets and potential optimal combinations of interventions that can best reverse a disease state to a normal state, including a robust computational algorithm for finding multiple target optimal intervention (MTOI) solutions34 and a quantitative composition-activity relationship (QCAR) model for multi-component drug design35.
Diverse array-designed modes of hitting on multiple targets or pathways in Fangjiomics. Fangjiomics-based combination therapy arranges the contributing agents in “magic shotguns” array (A) to hit on the multi-targets level. Vertical array (B) is usually applied to hit on different sites or stages of the same pathway. Besides, several combination therapies act on different targets in related pathways (C) or parallel pathways (D) in horizontal array to improve efficacy and reduce side effects.
Hitting the same pathway in a vertical array
Drug combinations in a vertical array are defined as combinations that simultaneously or sequentially act on different sites or stages of the same pathway to induce synergistic and additive interactions24. The point mutations (Val600Glu) in the serine/threonine-protein kinase BRAF [BRAF(V600E)] activate RAF-MEK-ERK signaling for tumor cell growth in malignant melanomas36. The vertical pathway combination of BRAF (GSK2118436) and MEK (GSK1120212) inhibitors, which has a sound rational basis in seeking to overcome several of the identified mechanisms of resistance to BRAF inhibitors, as well as blocking the undesirable paradoxical activation of CRAF in healthy cells (through RAF dimer formation) following treatment with BRAF inhibitors37,38, has resulted in additive activity against melanoma with surprisingly manageable skin toxicity and encouraging efficacy39 (Figure 1B).
Hitting different targets of related pathways or parallel pathways in a horizontal array
Horizontal array combination therapies refer to combinations that hit different targets in related pathways or parallel pathways to improve efficacy and reduce side effects24.
Hitting different targets of related pathways: one example of the rational, horizontal, combinatorial hitting different targets of related pathways that regulate the same process is the combination of aplidin and cytarabine with a synergic effect. Both drugs complement each other's activity by inducing apoptosis via two major apoptotic cascades. Aplidin activates and clusters death receptors through the Fas ligand40, which subsequently activates the receptor-mediated extrinsic cascade41, while cytarabine increases cellular stress and reduces survival protein MCl1, which subsequently activates CAsPs and apoptosis42 and triggers the mitochondrial intrinsic cascade41 (Figure 1C).
Hitting parallel pathways
A synergy screen with 14 targeted drugs in a cell line derived from a DDLS patient with the 12q13–15 amplicon was performed to identify effective, synergistic drug combinations for DDLS through a hybrid experimental and computational approach for deriving context-specific signaling models. The synergistic effect of the combination of CDK4 and IGF1R inhibitors was identified by inhibiting two seemingly parallel and non-overlapping pathways that control cell viability (the AKT/mTOR pathway by IGF1R, and the retinoblastoma (RB) pathway by CDK4)43 (Figure 1D). By optimizing the fusion of more pathways through compounds with a greater contribution to the combination therapy, the combination of jasminoidin and ursodeoxycholic acid was found to exert a synergistic effect in the horizontal array44. Moreover, some mathematical models have been introduced into the analysis of the mechanism of drug combinations in the horizontal array, including an outcome-dependent global similarity analysis (GSI)45, additive index46, and multi-objective evolutionary algorithm47.
Hitting network hubs in a focusing array
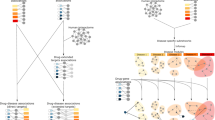
Combination therapy in a focusing array, also termed a “central hit” strategy, aims to damage network integrity by hitting network hubs in a selective manner. It is useful to target diseases characterized as flexible networks (eg, cancer, infectious disease)48. Towards this end, it would be helpful to obtain the detailed information of the topological structural differences between the host and parasite or the healthy and malignant networks. The identification of the hubs or central nodes/edges of various networks is an essential procedure in this strategy (Figure 2A). For instance, essential enzymes of metabolic networks are usually considered as drug targets in infectious diseases49,50 and in cancer51. When we design drug combinations against a network in an infectious organism or against cancer cells, many parameters such as the network topology, metabolic fluxes, conditions of the afflicted organisms, external environment interactions, stressor effects, etc should be taken into consideration. In the directed, hierarchical networks, the central hit strategy should attack the nodes at the top of the hierarchy. The high position nodes in hierarchical networks can be identified by the random upstream regions52. Recent studies on the connections of essential reactions and on super-essential reactions indicate that essential reactions form a core of metabolic networks and that super-essential reactions are needed in all organisms53,54. The cancer-specific targeting of signaling networks is mostly used in current anticancer strategies. The key aims of anti-cancer strategies include the identification of targets and the efficient combination of drugs to overcome the robustness of cancer-specific cellular networks with the least toxicity and potential resistance development55,56,57. Cytostatic drug targets have also been identified through the analysis of cancer-specific human metabolic networks58. A large-scale RNAi screen identified feedback activation of EGFR as a cause of colon cancer cell resistance to BRAF inhibition, suggesting the use of a synergistic combination of BRAF and EGFR inhibitors in BRAF-mutant, EGFR-expressing colon tumors59.
Network array-based modes in Fangjiomics. The arranged profiles of the contributing combination agents are focusing array (A) or siege array (B) in targeted network. Network array-based modes in Fangjiomics. The arranged profiles of the contributing combination agents are focusing array (A) or siege array (B) in targeted network. Dynamic array (C) represents hitting targets that alter with time at the network level.
Rewiring networks from a diseased state to healthy state in a siege array
In contrast to a focusing array, the combination in a “siege array” aims to shift a dysfunctional network to its normal state, which is also termed a “network influence” approach48. This mode is often applied in rewiring more rigid systems (eg, type 2 diabetes mellitus). Rigid systems are “well-defined” and transmit (but not dissipate) perturbations well. Thus, the optimal modification of rigid systems may be achieved by an indirect, “under-defined” attack of the neighbors of their central nodes or rigid clusters (Figure 2B). To this end, an understanding of the network dynamics both in healthy and diseased states is required, as well as knowledge of the existing drug targets of the particular disease. The network influence strategy is much less developed than the central hit strategy. With the network influence strategy, breaking down the system's robustness to push the system from one that favors the diseased state to one that favors a healthy state is a difficult task. The preferred targets of the network influence strategy are considered as the connected nodes located in vulnerable points of disease-related networks such as in inter-modular, bridging positions60,61,62,63,64,65. In signaling networks, the strategy of influencing preferred nodes in the network inhibits certain outputs of the signaling network, while leaving others intact to redirect the signal flow in the network66,67,68. The treatment strategy in a “siege array” often targets network segments (eg, disease-modules69). The targeting the central nodes/edges of systems with low plasticity may easily “over-saturate” the system, leading to a change in the system that becomes too substantial to be selective and causes side effects and toxicity. Therefore, the “siege array” often requires an indirect approach [eg, the neighbors of the real target are targeted (allo-network drugs) or multiple targets are targeted 'mildly' (multi-target drugs)], and their indirect and/or superposed effects result in the reconfiguration of a diseased network state back into a normal one. A strong synergy was identified between danusertib and bosutinib that exclusively affected CML cells harboring BCR-ABLT315I. Both compounds targeted MAPK pathways, downstream of BCR-ABL, resulting in the impaired activity of c-Myc. Thus, this drug combination synergistically targeted the dependency of BCR-ABLT315I CML cells on c-Myc through nonobvious off-targets70.
Adapting temporal and spatial variations of a disease network in a dynamic array
Growing evidence has indicated that a dynamic nature is a critical property of biological networks. In line with this property of disease networks, drug combinations in a dynamic array should be applied by dynamically altering the administration of ingredients in combinations to reverse a complex disease network (Figure 2C). Two common approaches are Boolean dynamics71,72, in which each node can exist in two states (inactive or active), or using concentrations of the nodes with dynamic models based on ordinary differential equations71,73. The latter strategy is most commonly used in pharmacokinetic-pharmacodynamic models. Following network modeling, Yaffe and colleagues managed to decode the signaling network dynamics that drive resistance to DNA-damaging chemotherapy. This information was used to sensitize otherwise resistant triple-negative breast cancer cells to conventional DNA-damaging chemotherapy by administering doxorubicin (Adriamycin, Doxil) and erlotinib (Tarceva) in an order- and time-dependent fashion74. Moreover, the application of evolutionary models in drug-resistant non-small cell lung cancer (NSCLC), along with cell-based studies, has revealed that sequential therapy using cytotoxic agents with either erlotinib (Tarceva) or gefitinib (Iressa) was more effective than monotherapy or concurrent combinatorial dosing75.
Conclusions and perspectives
Through array-designed multi-scale level interventions, Fangjiomics-based combination therapy appears to be a promising treatment strategy for complex diseases. However, much work remains to be undertaken, and we must overcome certain obstacles.
Developing a phenotype-driven strategy “from bench to bedside”
A major challenge in optimizing combination therapy is to translate experimental and computational modeling into clinical practice. Although Fangjiomics-based combination therapy is based on the integration of clinical knowledge with multi-scale omics data, the translational approaches are still far from mature. One recommendation is to develop a phenotype-driven strategy based on the clinical outcomes treated by combination therapy. This strategy should elucidate the relationship between the clinical phenome (symptoms and signs) and integrate information from multi-scale omics data such as metabolomics, proteomics, transcriptomics, and genomics, which would yield more information on a biological process than the analysis of a single type of data76. However, the gathering of clinical phenotype data likely presents a greater challenge than high-throughput sequencing projects, due to the range of phenotype measurements and the complexity of the data77. The use of ontologies was proposed as an approach to semantic standardization78 for semantically categorizing phenodeviance79 such as Semantic Web technologies80, Systems Biology Markup Language81, and Web Ontology Language (OWL)82,83. Moreover, in pharmacological research, PhenomeDrug84 is another approach for predicting novel associations between drugs and diseases based on the PhenomeNET85 method for comparing phenotypes across species.
Quantitative multi-scale pharmacodynamic modeling techniques to predict the efficacy of combination therapies
Another challenge is the development of a mechanistic understanding of how multi-scale omics networks control variability in combination therapy responses at the organismal level. An enhanced pharmacodynamic model coupling detailed models of cellular regulatory networks with measurable pharmacokinetic and pharmacodynamic parameters has been used to to quantitatively predict the response of anti-EGFR therapy in decreasing tumor size86. The characterization of the topology of cellular regulatory networks and an understanding of the dynamic capability of the network topology can help to explain both the therapeutic and adverse effects of combination therapy87. For the calcium-sensing receptor and PTH, the identification of the molecular components that participate in the negative feedback loop and the means through which they can be modulated can help us to design better antagonists of the calcium-sensing receptor or develop polypharmacology for the treatment of osteoporosis88. In Fangjiomics-based therapy, we require better quantitative models of pharmacological mechanism at all scales, which should use available omics and pharmacodynamics data to correctly predict the effects of known omics changes on the response rates to combination therapy. Thus, Quantitative and Systems Pharmacology (QSP) will be introduced to identify and validate target (and druggable) networks, uncover drug-response biomarkers, design improved drug combinations, select appropriate doses and dosage regimens, and identify those patients most likely to respond to novel therapeutic combinations89.
Modularizing networks to deconstruct the relationship between complex diseases and combination therapy
Organized modularity is ubiquitous in various network systems90 such as metabolic91, transcriptional regulation92, and protein-protein interaction (PPI)93 networks. The application of computational and mathematical modeling approaches to achieving combinatorial selectivity through the use of drug combinations requires drugs with adequately modular structures. Hence, the identification of functional modules from multi-scale omics networks is becoming extremely important and necessary. The term "module" here refers to the minimal functional unit in a biological or pharmacological profile to reveal the features of organisms or drugs, as well as their mutual interactions90,94. Using a module map, different tumors could be differentiated through the activation of modules specific to particular types of tumors95. Because highly connected substrates may represent the critical connections between modules that control distinct metabolic functions96, a modular approach may provide insights into possible novel mechanisms of action for a wide range of drugs and may also identify potential new targets for combination therapy97.
Ultimately, based on the integration of multi-scale omics and quantitative modularized modeling of the relationship between complex diseases and combination therapy, Fangjiomics-based combination therapies are likely to pave the way to achieving precision medicine, which would ensure that patients receive the right treatment at the right dose at the right time, with minimum ill consequences and maximum efficacy.
References
Chen B, Butte AJ . Network medicine in disease analysis and therapeutics. Clin Pharmacol Ther 2013; 94: 627–9.
Hamosh A, Scott AF, Amberger JS, Bocchini CA, McKusick VA . Online mendelian inheritance in man (OMIM), a knowledgebase of human genes and genetic disorders. Nucleic Acids Res 2005; 33: D514-7.
Azmi AS, Wang Z, Philip PA, Mohammad RM, Sarkar FH . Proof of concept: network and systems biology approaches aid in the discovery of potent anticancer drug combinations. Mol Cancer Ther 2010; 9: 3137–44.
Caskey CT . The drug development crisis: efficiency and safety. Annu Rev Med 2007; 58: 1–16.
Yan Q . Toward the integration of personalized and systems medicine: challenges, opportunities and approaches. Per Med 2011; 8: 1–4.
Jiang G, Li RH, Sun C, Jia HY, Lei TC, Liu YQ . Efficacy and safety between temozolomide alone and temozolomide-based double therapy for malignant melanoma: a meta-analysis. Tumor Biol 2014; 35: 315–22.
Basch E, Autio K, Ryan CJ, Mulders P, Shore, N, Kheoh T, et al. Abiraterone acetate plus prednisone versus prednisone alone in chemotherapy-naive men with metastatic castration-resistant prostate cancer: patient-reported outcome results of a randomised phase 3 trial. Lancet Oncol 2013; 14: 1193–9.
Gouya G, Arrich J, Wolzt M, Huber K, Verheugt FWA, Gurbel PA, et al. Antiplatelet treatment for prevention of cerebrovascular events in patients with vascular diseases a systematic review and meta-analysis. Stroke 2014; 45: 492–503.
Nannini LJ, Poole P, Milan SJ, Kesterton A . Combined corticosteroid and long-acting beta(2)-agonist in one inhaler versus inhaled corticosteroids alone for chronic obstructive pulmonary disease. Cochrane Database Syst Rev 2013; 8: CD006826.
Mayer RJ . Targeted therapy for advanced colorectal cancer-more is not always better. New Engl J Med 2009; 360: 623–5.
Lee M, Saver JL, Hong KS, Rao NM, Wu YL, Ovbiagele B . Risk-benefit profile of long-term dual-versus single-antiplatelet therapy among patients with ischemic stroke. Ann Intern Med 2013; 159: 463–70.
Komarova NL, Katouli A A, Wodarz D . Combination of two but not three current targeted drugs can improve therapy of chronic myeloid leukemia. PLoS One 2009; 4: e4423.
Santos N, Wenger JB, Havre P, Liu Y, Dagan R, Imanirad I, et al. Combination therapy for renal cell cancer: what are possible options? Oncology 2011; 81: 220–9.
Bliss CI . The calculation of microbial assays. Bacteriol Rev 1956; 20: 243–58.
Loewe S . The problem of synergism and antagonism of combined drugs. Arzneimittelforschung 1953; 3: 285–90.
Chou TC . Talalay P: Analysis of combined drug effects: a new look at a very old problem. Trends Pharmacol Sci 1983; 4: 450–4.
Zheng M, Liu X, Xu Y, Li H, Luo C, Jiang H . Computational methods for drug design and discovery: focus on China. Trends Pharmacol Sci 2013; 34: 549–59.
Mestres J, Gregori-Puigjane E, Valverde S, Sole RV . The topology of drug–target interaction networks: implicit dependence on drug properties and target families. Mol Biosyst 2009; 5: 1051–7.
Koutsoukas A, Simms B, Kirchmair J, Bond PJ, Whitmore AV, Zimmer S, et al. From in silico target prediction to multi-target drug design: current databases, methods and applications. J Proteomics 2011; 74: 2554–74.
Li S, Zhang B, Zhang N . Network target for screening synergistic drug combinations with application to traditional Chinese medicine. BMC Syst Biol 2011; 5 Suppl 1: S10.
Fitzgerald JB, Schoeberl B, Nielsen UB, Sorger PK . Systems biology and combination therapy in the quest for clinical efficacy. Nat Chem Biol 2006; 2: 458–66.
Araujo RP, Liotta LA, Petricoin EF . Proteins, drug targets and the mechanisms they control: the simple truth about complex networks. Nat Rev Drug Discov 2007; 6: 871–80.
Csermely P, Agoston V, Pongor, S . The efficiency of multi-target drugs: the network approach might help drug design. Trends Pharmacol Sci 2005; 26: 178–82.
Wang Z, Liu J, Cheng Y, Wang Y . Fangjiomics: in search of effective and safe combination therapies. J Clin Pharmacol 2011; 51: 1132–51.
Zhao S, Iyengar R . Systems pharmacology: network analysis to identify multiscale mechanisms of drug action. Annu Revi Pharmacol Toxicol 2012; 52: 505–21.
Wang L, Zhou GB, Liu P, Song JH, Liang Y, Yan XJ, et al. Dissection of mechanisms of Chinese medicinal formula Realgar-Indigo naturalis as an effective treatment for promyelocytic leukemia. Proc Natl Acad Sci U S A 2008; 105: 4826–31.
University of California-San Francisco. Cancer's next magic bullet may be magic shotgun. ScienceDaily. 2012 Jun [cited 2014 Aug 22]. Available from: http://www.sciencedaily.com/releases/2012/06/120615141716.htm
Sullivan DJ . Plasmodium drug targets outside the genetic control of the parasite. Curr Pharm Des 2013; 19: 282–9.
Roth BL, Sheffler DJ, Kroeze WK . Magic shotguns versus magic bullets: selectively non-selective drugs for mood disorders and schizophrenia. Nat Rev Drug Discov 2004; 3: 353–9.
Dar AC, Das TK, Shokat KM, Cagan RL . Chemical genetic discovery of targets and anti-targets for cancer polypharmacology. Nature 2012; 486: 80–4.
Margineanu DG . Systems biology impact on antiepileptic drug discovery. Epilepsy Res 2012; 98: 104–15.
Talevi A, Bellera CL, Di Ianni M, Gantner M, Bruno-Blanch LE, Castro EA . CNS drug development-lost in translation? Mini Rev Med Chem 2012; 12: 959–70.
Liu J, Zhou CX, Zhang ZJ, Wang LY, Jing ZW, Wang Z . Synergistic mechanism of gene expression and pathways between jasminoidin and ursodeoxycholic acid in treating focal cerebral ischemia-reperfusion injury. CNS Neurosci Ther 2012; 18: 674–82.
Yang K, Bai H, Ouyang Q, Lai L, Tang C . Finding multiple target optimal intervention in disease-related molecular network. Mol Syst Biol 2008; 4: 228.
Wang Y, Yu L, Zhang L, Qu H, Cheng Y . A novel methodology for multicomponent drug design and its application in optimizing the combination of active components from Chinese medicinal formula Shenmai. Chem Biol Drug Des 2010; 75: 318–24.
Johannessen CM, Johnson LA, Piccioni F, Townes A, Frederick DT, Donahue MK, et al. A melanocyte lineage program confers resistance to MAP kinase pathway inhibition. Nature 2013; 504: 138–42.
Heidorn SJ, Milagre C, Whittaker S, Nourry A, Niculescu-Duvas I, Dhomen N, et al. Kinase-dead BRAF and oncogenic RAS cooperate to drive tumor progression through CRAF. Cell 2010; 140: 209–21.
Poulikakos PI, Zhang C, Bollag G, Shokat KM, Rosen N . RAF inhibitors transactivate RAF dimers and ERK signalling in cells with wild-type BRAF. Nature 2010; 464: 427–30.
Infante JR, Falchook GS, Lawrence DP, Weber JS, Kefford RF, Bendell JC, et al. Phase I/II study to assess safety, pharmacokinetics, and efficacy of the oral MEK 1/2 inhibitor GSK1120212 (GSK212) dosed in combination with the oral BRAF inhibitor GSK2118436 (GSK436). J Clin Oncol (Meeting Abstracts) 2011; 29: CRA8503.
Gajate C, Mollinedo F . Cytoskeleton-mediated death receptor and ligand concentration in lipid rafts forms apoptosis-promoting clusters in cancer chemotherapy. J Biol Chem 2005; 280: 11641–7.
Hajra KM, Liu JR . Apoptosome dysfunction in human cancer. Apoptosis 2004; 9: 691–704.
de Vries JF, Falkenburg JH, Willemze R, Barge RM . The mechanisms of Ara-C-induced apoptosis of resting B-chronic lymphocytic leukemia cells. Haematologica 2006; 91: 912–9.
Miller ML, Molinelli EJ, Nair JS, Sheikh T, Samy R, Jing X, et al. Drug synergy screen and network modeling in dedifferentiated liposarcoma identifies CDK4 and IGF1R as synergistic drug targets. Sci Signal 2013; 6: ra85.
Wang Z, Jing ZW, Zhou CX, Zhang L, Cheng J, Zhang ZJ, et al. Fusion of core pathways reveals a horizontal synergistic mechanism underlying combination therapy. Eur J Pharmacol 2011; 667: 278–86.
Liu J, Zhang ZJ, Zhou CX, Wang Y, Cheng YY, Duan DY, et al. Outcome-dependent global similarity analysis of imbalanced core signaling pathways in ischemic mouse hippocampus. CNS Neurol Disord Drug Targets 2012; 11: 1070–82.
Zhang YY, Li HX, Chen YY, Fang H, Yu YN, Liu J, et al. Convergent and divergent pathways decoding hierarchical additive mechanisms in treating cerebral ischemia-reperfusion injury. CNS Neurosci Ther 2014; 20: 253–63.
Small BG, McColl BW, Allmendinger R, Pahle J, López-Castejón G, Rothwell NJ, et al. Efficient discovery of anti-inflammatory small-molecule combinations using evolutionary computing. Nat Chem Biol 2011; 7: 902–8
Csermely P, Korcsmáros T, Kiss HJ, London G, Nussinov R . Structure and dynamics of molecular networks: a novel paradigm of drug discovery: a comprehensive review. Pharmacol Ther 2013; 138: 333–408.
Fatumo S, Plaimas K, Mallm JP, Schramm G, Adebiyi E, Oswald M, et al. Estimating novel potential drug targets of Plasmodium falciparum by analysing the metabolic network of knock-out strains in silico. Infect Genet Evol 2009; 9: 351–8.
Fatumo S, Plaimas K, Adebiyi E, Konig R . Comparing metabolic network models based on genomic and automatically inferred enzyme information from Plasmodium and its human host to define drug targets in silico. Infect Genet Evol 2011; 11: 708–15.
Yu LR, Issaq HJ, Veenstra TD . Phosphoproteomics for the discovery of kinases as cancer biomarkers and drug targets. Proteomics Clin Appl 2007; 1: 1042–57.
Liu YY, Slotine JJ, Barabasi AL . Control centrality and hierarchical structure in complex networks. PLoS One 2012; 7: e44459.
Barve A, Rodrigues JF, Wagner A . Superessential reactions in metabolic networks. Proc Natl Acad Sci U S A 2012; 109: E1121–30.
Ma J, Zhang X, Ung CY, Chen YZ, Li B . Metabolic network analysis revealed distinct routes of deletion effects between essential and non-essential genes. Mol Biosyst 2012; 8: 1179–86.
Kitano HH . Cancer as a robust system: implications to anticancer therapy. Nat Rev Cancer 2004; 4: 227–35.
Kitano HH . A robustness-based approach to systems-oriented drug design. Nat Rev Drug Discov 2007; 6: 202–10.
Cheng TM, Gulati S, Agius R, Bates PA . Understanding cancer mechanisms through network dynamics. Brief Funct Genomics 2012; 11: 543–60.
Folger O, Jerby L, Frezza C, Gottlieb E, Ruppin E, Shlomi T . Predicting selective drug targets in cancer through metabolic networks. Mol Syst Biol 2011; 7: 501.
Prahallad A, Sun C, Huang S, Di Nicolantonio F, Salazar R, Zecchin D, et al. Unresponsiveness of colon cancer to BRAF(V600E) inhibition through feedback activation of EGFR. Nature 2012; 483: 100–3.
Antal MA, Böde C, Csermely P . Perturbation waves in proteins and protein networks: Applications of percolation and game theories in signaling and drug design. Curr Prot Pept Sci 2009; 10: 161–72.
Hase T, Tanaka H, Suzuki Y, Nakagawa S, Kitano H . Structure of protein interaction networks and their implications on drug design. PLoS Comput Biol 2009; 5: e1000550.
Zanzoni A, Soler-Lopez M, Aloy P . A network medicine approach to human disease. FEBS Lett 2009; 583: 1759–65.
Fliri AF, Loging WT, Volkmann RA . Cause-effect relationships in medicine: a protein network perspective. Trends Pharmacol Sci 2010; 31: 547–55.
Farkas IJ, Korcsmáros T, Kovács I A, Mihalik Á, Palotai R, Simkó G I, et al. Network-based tools in the identification of novel drug-targets. Sci Signal 2011; 4: pt3.
Yu Q, Huang JF . The analysis of the druggable families based on topological features in the protein-protein interaction network. Lett Drug Des Discov 2012; 9: 426–30.
Dasika MS, Burgard A, Maranas CD . A computational framework for the topological analysis and targeted disruption of signal transduction networks. Biophys J 2006; 91: 382–98.
Ruths DA, Nakhleh L, Iyengar MS, Reddy SA, Ram PT . Hypothesis generation in signaling networks. J Comput Biol 2006; 13: 1546–57.
Pawson T, Linding R . Network medicine. FEBS Lett 2008; 582: 1266–70.
Cho DY, Kim YA, Przytycka TM . Chapter 5: Network biology approach to complex diseases. PLoS Comput Biol 2012; 8: e1002820.
Winter GE, Rix U, Carlson SM, Gleixner KV, Grebien F, Gridling M, et al. Systems-pharmacology dissection of a drug synergy in imatinib-resistant CML. Nat Chem Biol 2012; 8: 905–12.
Ferrell JE, Tsai TYC, Yang Q . Modeling the cell cycle: why do certain circuits oscillate? Cell 2011; 144: 874–85.
Singhania R, Sramkoski RM, Jacobberger JW, Tyson JJ . A hybrid model of mammalian cell cycle regulation. PLoS Comput Biol 2011; 7: e1001077.
Neves SR, Tsokas P, Sarkar A, Grace EA, Rangamani P, et al. Cell shape and negative links in regulatory motifs together control spatial information flow in signaling networks. Cell 2008; 133: 666–80.
Lee MJ, Ye AS, Gardino AK, Heijink AM, Sorger PK, MacBeath G, et al. Sequential application of anticancer drugs enhances cell death by rewiring apoptotic signaling networks. Cell 2012; 149: 780–94.
Mumenthaler SM, Foo J, Leder K, Choi NC, Agus DB, Pao W, et al. Evolutionary modeling of combination treatment strategies to overcome resistance to tyrosine kinase inhibitors in non-small cell lung cancer. Mol Pharm 2011; 8: 2069–79.
Snyder M, Du J, Gerstein M . Personal genome sequencing: current approaches and challenges. Genes Dev 2010; 24: 423–31.
Gkoutos GV, Schofield PN, Hoehndorf R . Computational tools for comparative phenomics: the role and promise of ontologies. Mamm Genome 2012; 23: 669–79.
Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 2000; 25: 25–9.
Gkoutos GV, Green EC, Mallon AM, Hancock JM, Davidson D . Building mouse phenotype ontologies. Pac Symp Biocomput 2004: 178–89.
Ruttenberg A, Clark T, Bug W, Samwald M, Bodenreider O, Chen H, et al. Advancing translational research with the Semantic Web. BMC Bioinformatics 2007; 8 Suppl 3: S2.
Hoehndorf R, Dumontier M, Gennari JH, Wimalaratne S, de Bono B, Cook DL, et al. Integrating systems biology models and biomedical ontologies. BMC Syst Biol 2011; 5: 124.
Wolstencroft K, Lord P, Tabernero L, Brass A, Stevens R . Protein classification using ontology classification. Bioinformatics 2006; 22: e530–8.
Hoehndorf R, Dumontier M, Oellrich A, Rebholz-Schuhmann D, Schofield PN, Gkoutos GV . Interoperability between biomedical ontologies through relation expansion, upper-level ontologies and automatic reasoning. PLoS One 2011; 6: e22006.
Hoehndorf R, Oellrich A, Rebholz-Schuhmann D, Schofield PN, Gkoutos GV . Linking PharmGKB to phenotype studies and animal models of disease for drug repurposing. Pac Symp Biocomput 2012: 388–99.
Hoehndorf R, Schofield PN, Gkoutos GV . PhenomeNET: a whole-phenome approach to disease gene discovery. Nucleic Acids Res 2011; 39: e119.
Iyengar R, Zhao S, Chung SW, Mager DE, Gallo JM . Merging systems biology with pharmacodynamics. Sci Transl Med 2012; 4: 126ps7.
Zhao S, Iyengar R . Systems pharmacology: network analysis to identify multiscale mechanisms of drug action. Annu Rev Pharmacol Toxicol 2012; 52: 505–21.
Abraham AK, Maurer TS, Kalgutkar AS, Gao X, Li M, Healy DR, et al. Pharmacodynamicmodel of parathyroid hormone modulation by a negative allosteric modulator of the calcium-sensing receptor. AAPS J 2011; 13: 265–73.
Sorger PK, Allerheiligen SR, Abernethy DR, Altman RB, Brouwer KL, Califano A, et al. Quantitative and systems pharmacology in the post-genomic Era: new approaches to discovering drugs and understanding therapeutic mechanisms [monograph on the Internet]. National Institutes of Health White Paper; 2011 [cited 2014 Jul 9]. Available from: http://www.nigms.nih.gov/training/documents/systemspharmawpsorger2011.pdf
Lorenz DM, Jeng A, Deem MW . The emergence of modularity in biological systems. Phys Life Rev 2011; 8: 129–60.
Ravasz E, Somera AL, Mongru DA, Oltyai ZN, Barabasi AL . Hierarchical organization of modularity in metabolic networks. Science 2002; 297: 1551–5.
Ihmels J, Friedlander G, Bergmann S, Sarig O, Ziv Y, Barkal N . Revealing modular organization in the yeast transcriptional network. Nat Genet 2002; 31: 370–7.
Han JD, Bertin N, Hao T, Goldberg DS, Berriz GF, Zhang LV, et al. Evidence for dynamically organized modularity in the yeast protein–protein interaction network. Nature 2004; 430: 88–93.
Wang Z, Wang YY . Modular pharmacology: deciphering the interacting structural organization of the targeted networks. Drug Discov Today 2013; 18: 560–6.
Segal E, Friedman N, Koller D, Regev A . A module map showing conditional activity of expression modules in cancer. Nat Genet 2004; 36: 1090–8.
Hartwell LH, Hopfield JJ, Leibler S, Murray AW . From molecular to modular cell biology. Nature 1999; 402: C47–52.
Kutalik Z, Beckmann JS, Bergmann S . A modular approach for integrative analysis of large-scale gene-expression and drug-response data. Nat Biotechnol 2008; 26: 531–9.
Acknowledgements
This study was supported by National Natural Science Foundation of China (No 81102741) and the sixth autonomously selected subject projects of the China Academy of Chinese Medical Sciences (No Z0214).
Author information
Authors and Affiliations
Corresponding author
PowerPoint slides
Rights and permissions
About this article
Cite this article
Liu, J., Wang, Z. Diverse array-designed modes of combination therapies in Fangjiomics. Acta Pharmacol Sin 36, 680–688 (2015). https://doi.org/10.1038/aps.2014.125
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/aps.2014.125
Keywords
This article is cited by
-
Milk Peptides as Novel Multi‐Targeted Therapeutic Candidates for SARS-CoV2
The Protein Journal (2021)
-
Similarity of therapeutic networks induced by a multi-component herbal remedy, Ukgansan, in neurovascular unit cells
Scientific Reports (2020)
-
New omic and network paradigms for deep understanding of therapeutic mechanisms for Fangji of traditional Chinese medicine
Acta Pharmacologica Sinica (2018)
-
Pure mechanistic analysis of additive neuroprotective effects between baicalin and jasminoidin in ischemic stroke mice
Acta Pharmacologica Sinica (2018)
-
Yangxin Tongmai Formula ameliorates impaired glucose tolerance in children with Graves' disease through upregulation of the insulin receptor levels
Acta Pharmacologica Sinica (2018)