Abstract
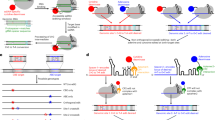
Much of our current understanding of microbiology is based on the application of genetic engineering procedures. Since their inception (more than 30 years ago), methods based largely on allelic exchange and two-step selection processes have become a cornerstone of contemporary bacterial genetics. While these tools are established for adapted laboratory strains, they have limited applicability in clinical or environmental isolates displaying a large and unknown genetic repertoire that are recalcitrant to genetic modifications. Hence, new tools allowing genetic engineering of intractable bacteria must be developed to gain a comprehensive understanding of them in the context of their biological niche. Herein, we present a method for precise, efficient and rapid engineering of the opportunistic pathogen Pseudomonas aeruginosa. This procedure relies on recombination of short single-stranded DNA facilitated by targeted double-strand DNA breaks mediated by a synthetic Cas9 coupled with the efficient Ssr recombinase. Possible applications include introducing single-nucleotide polymorphisms, short or long deletions, and short DNA insertions using synthetic single-stranded DNA templates, drastically reducing the need of PCR and cloning steps. Our toolkit is encoded on two plasmids, harboring an array of different antibiotic resistance cassettes; hence, this approach can be successfully applied to isolates displaying natural antibiotic resistances. Overall, this toolkit substantially reduces the time required to introduce a range of genetic manipulations to a minimum of five experimental days, and enables a variety of research and biotechnological applications in both laboratory strains and difficult-to-manipulate P. aeruginosa isolates.
Key points
-
This protocol provides an expanded toolkit for engineering genetically intractable Pseudomonas aeruginosa isolates, utilizing two plasmids to enable Ssr-mediated recombineering and CRISPR–Cas9 counterselection of mutated bacterial colonies in a single-step procedure.
-
The toolkit markedly reduces the time required to introduce a range of genetic manipulations compared with previous allelic exchange methods, facilitating a variety of research and biotechnological applications in both laboratory strains and difficult-to-manipulate P. aeruginosa isolates.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The authors declare that no new data were generated during the development of this protocol. The original publication describing the method expanded herein is available in the primary research paper (https://doi.org/10.1038/s41522-022-00268-1). All plasmids described in this protocol are available at Addgene (Table 1). Source data are provided with this paper.
References
Silby, M. W., Winstanley, C., Godfrey, S. A., Levy, S. B. & Jackson, R. W. Pseudomonas genomes: diverse and adaptable. FEMS Microbiol. Rev. 35, 652–680 (2011).
Tacconelli, E. et al. Discovery, research, and development of new antibiotics: the WHO priority list of antibiotic-resistant bacteria and tuberculosis. Lancet Infect. Dis. 18, 318–327 (2018).
Antimicrobial Resistance Collaborators. Global burden of bacterial antimicrobial resistance in 2019: a systematic analysis. Lancet 399, 629–655 (2022).
Lee, D. G. et al. Genomic analysis reveals that Pseudomonas aeruginosa virulence is combinatorial. Genome Biol. 7, R90 (2006).
Klockgether, J., Cramer, N., Wiehlmann, L., Davenport, C. F. & Tummler, B. Pseudomonas aeruginosa genomic structure and diversity. Front. Microbiol. 2, 150 (2011).
Freschi, L. et al. The Pseudomonas aeruginosa pan-genome provides new insights on its population structure, horizontal gene transfer, and pathogenicity. Genome Biol. Evol. 11, 109–120 (2019).
Ciofu, O., Riis, B., Pressler, T., Poulsen, H. E. & Hoiby, N. Occurrence of hypermutable Pseudomonas aeruginosa in cystic fibrosis patients is associated with the oxidative stress caused by chronic lung inflammation. Antimicrob. Agents Chemother. 49, 2276–2282 (2005).
Rossi, E. et al. Pseudomonas aeruginosa adaptation and evolution in patients with cystic fibrosis. Nat. Rev. Microbiol. 19, 331–342 (2021).
Culyba, M. J. & Van Tyne, D. Bacterial evolution during human infection: adapt and live or adapt and die. PLoS Pathog. 17, e1009872 (2021).
Schweizer, H. P. Allelic exchange in Pseudomonas aeruginosa using novel ColE1-type vectors and a family of cassettes containing a portable oriT and the counter-selectable Bacillus subtilis sacB marker. Mol. Microbiol. 6, 1195–1204 (1992).
Hoang, T. T., Karkhoff-Schweizer, R. R., Kutchma, A. J. & Schweizer, H. P. A broad-host-range Flp-FRT recombination system for site-specific excision of chromosomally-located DNA sequences: application for isolation of unmarked Pseudomonas aeruginosa mutants. Gene 212, 77–86 (1998).
Choi, K. H. & Schweizer, H. P. An improved method for rapid generation of unmarked Pseudomonas aeruginosa deletion mutants. BMC Microbiol. 5, 30 (2005).
Shanks, R. M., Caiazza, N. C., Hinsa, S. M., Toutain, C. M. & O’Toole, G. A. Saccharomyces cerevisiae-based molecular tool kit for manipulation of genes from gram-negative bacteria. Appl. Environ. Microbiol. 72, 5027–5036 (2006).
Martinez-Garcia, E. & de Lorenzo, V. Engineering multiple genomic deletions in Gram-negative bacteria: analysis of the multi-resistant antibiotic profile of Pseudomonas putida KT2440. Environ. Microbiol. 13, 2702–2716 (2011).
Hmelo, L. R. et al. Precision-engineering the Pseudomonas aeruginosa genome with two-step allelic exchange. Nat. Protoc. 10, 1820–1841 (2015).
Ran, F. A. et al. Genome engineering using the CRISPR–Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Wirth, N. T., Kozaeva, E. & Nikel, P. I. Accelerated genome engineering of Pseudomonas putida by I-SceI-mediated recombination and CRISPR-Cas9 counterselection. Microb. Biotechnol. 13, 233–249 (2020).
Aparicio, T., Jensen, S. I., Nielsen, A. T., de Lorenzo, V. & Martinez-Garcia, E. The Ssr protein (T1E_1405) from Pseudomonas putida DOT-T1E enables oligonucleotide-based recombineering in platform strain P. putida EM42. Biotechnol. J. 11, 1309–1319 (2016).
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl Acad. Sci. Usa. 97, 6640–6645 (2000).
Lesic, B. & Rahme, L. G. Use of the lambda Red recombinase system to rapidly generate mutants in Pseudomonas aeruginosa. BMC Mol. Biol. 9, 20 (2008).
Aparicio, T., de Lorenzo, V. & Martinez-Garcia, E. CRISPR/Cas9-based counterselection boosts recombineering efficiency in Pseudomonas putida. Biotechnol. J. 13, e1700161 (2018).
Aparicio, T., de Lorenzo, V. & Martinez-Garcia, E. CRISPR/Cas9-enhanced ssDNA recombineering for Pseudomonas putida. Microb. Biotechnol. 12, 1076–1089 (2019).
Jeske, A., Arce-Rodriguez, A., Thoming, J. G., Tomasch, J. & Haussler, S. Evolution of biofilm-adapted gene expression profiles in lasR-deficient clinical Pseudomonas aeruginosa isolates. NPJ Biofilms Microbiomes 8, 6 (2022).
Calero, P. & Nikel, P. I. Chasing bacterial chassis for metabolic engineering: a perspective review from classical to non-traditional microorganisms. Microb. Biotechnol. 12, 98–124 (2019).
Xin, X. F., Kvitko, B. & He, S. Y. Pseudomonas syringae: what it takes to be a pathogen. Nat. Rev. Microbiol. 16, 316–328 (2018).
Chen, W. et al. CRISPR/Cas9-based genome editing in Pseudomonas aeruginosa and cytidine deaminase-mediated base editing in Pseudomonas species. iScience 6, 222–231 (2018).
Ricaurte, D. E. et al. A standardized workflow for surveying recombinases expands bacterial genome-editing capabilities. Microb. Biotechnol. 11, 176–188 (2018).
Volke, D. C., Martino, R. A., Kozaeva, E., Smania, A. M. & Nikel, P. I. Modular (de)construction of complex bacterial phenotypes by CRISPR/nCas9-assisted, multiplex cytidine base-editing. Nat. Commun. 13, 3026 (2022).
Konstantakos, V., Nentidis, A., Krithara, A. & Paliouras, G. CRISPR–Cas9 gRNA efficiency prediction: an overview of predictive tools and the role of deep learning. Nucleic Acids Res. 50, 3616–3637 (2022).
Blin, K., Pedersen, L. E., Weber, T. & Lee, S. Y. CRISPy-web: an online resource to design sgRNAs for CRISPR applications. Synth. Syst. Biotechnol. 1, 118–121 (2016).
Concordet, J. P. & Haeussler, M. CRISPOR: intuitive guide selection for CRISPR/Cas9 genome editing experiments and screens. Nucleic Acids Res. 46, W242–W245 (2018).
Aparicio, T. et al. Mismatch repair hierarchy of Pseudomonas putida revealed by mutagenic ssDNA recombineering of the pyrF gene. Environ. Microbiol. 22, 45–58 (2020).
Silva-Rocha, R. et al. The Standard European Vector Architecture (SEVA): a coherent platform for the analysis and deployment of complex prokaryotic phenotypes. Nucleic Acids Res. 41, D666–D675 (2013).
Martinez-Garcia, E. et al. SEVA 4.0: an update of the Standard European Vector Architecture database for advanced analysis and programming of bacterial phenotypes. Nucleic Acids Res. 51, D1558–D1567 (2023).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Fu, Y., Sander, J. D., Reyon, D., Cascio, V. M. & Joung, J. K. Improving CRISPR–Cas nuclease specificity using truncated guide RNAs. Nat. Biotechnol. 32, 279–284 (2014).
Ran, F. A. et al. Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell 154, 1380–1389 (2013).
Wang, T., Wei, J. J., Sabatini, D. M. & Lander, E. S. Genetic screens in human cells using the CRISPR–Cas9 system. Science 343, 80–84 (2014).
Lin, Y. et al. CRISPR/Cas9 systems have off-target activity with insertions or deletions between target DNA and guide RNA sequences. Nucleic Acids Res. 42, 7473–7485 (2014).
Xu, H. et al. Sequence determinants of improved CRISPR sgRNA design. Genome Res. 25, 1147–1157 (2015).
Doench, J. G. et al. Rational design of highly active sgRNAs for CRISPR–Cas9-mediated gene inactivation. Nat. Biotechnol. 32, 1262–1267 (2014).
Liang, G., Zhang, H., Lou, D. & Yu, D. Selection of highly efficient sgRNAs for CRISPR/Cas9-based plant genome editing. Sci. Rep. 6, 21451 (2016).
Ellis, H. M., Yu, D., DiTizio, T. & Court, D. L. High efficiency mutagenesis, repair, and engineering of chromosomal DNA using single-stranded oligonucleotides. Proc. Natl Acad. Sci. USA 98, 6742–6746 (2001).
Shen, H., Han, F., Lin, Y. & Yu, W. A high efficient electroporation of Pseudomonas sp. QDA pretreated with alginate lyase. Enzym. Microb. Technol. 39, 677–682 (2006).
Hanahan, D. & Meselson, M. in Methods in Enzymology (eds. Wu, R. et al.) Vol. 100, 333–342 (Academic Press, 1983).
Rahme, L. G. et al. Common virulence factors for bacterial pathogenicity in plants and animals. Science 268, 1899–1902 (1995).
Holloway, B. W. Genetic recombination in Pseudomonas aeruginosa. Microbiology 13, 572–581 (1955).
Hornischer, K. et al. BACTOME—a reference database to explore the sequence- and gene expression-variation landscape of Pseudomonas aeruginosa clinical isolates. Nucleic Acids Res. 47, D716–D720 (2019).
Boyer, H. W. & Roulland-Dussoix, D. A complementation analysis of the restriction and modification of DNA in Escherichia coli. J. Mol. Biol. 41, 459–472 (1969).
Keen, N. T., Tamaki, S., Kobayashi, D. & Trollinger, D. Improved broad-host-range plasmids for DNA cloning in gram-negative bacteria. Gene 70, 191–197 (1988).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Winsor, G. L. et al. Enhanced annotations and features for comparing thousands of Pseudomonas genomes in the Pseudomonas genome database. Nucleic Acids Res. 44, D646–D653 (2016).
Benson, D. A. et al. GenBank. Nucleic Acids Res 41, D36–D42 (2013).
Poulsen, B. E. et al. Defining the core essential genome of Pseudomonas aeruginosa. Proc. Natl Acad. Sci. USA 116, 10072–10080 (2019).
Liberati, N. T. et al. An ordered, nonredundant library of Pseudomonas aeruginosa strain PA14 transposon insertion mutants. Proc. Natl Acad. Sci. USA 103, 2833–2838 (2006).
Jacobs, M. A. et al. Comprehensive transposon mutant library of Pseudomonas aeruginosa. Proc. Natl Acad. Sci. USA 100, 14339–14344 (2003).
Hofacker, I. L. Vienna RNA secondary structure server. Nucleic Acids Res. 31, 3429–3431 (2003).
Kessler, B., de Lorenzo, V. & Timmis, K. N. A general system to integrate lacZ fusions into the chromosomes of gram-negative eubacteria: regulation of the Pm promoter of the TOL plasmid studied with all controlling elements in monocopy. Mol. Gen. Genet. 233, 293–301 (1992).
Simon, R., Priefer, U. & Pühler, A. A broad host range mobilization system for in vivo genetic engineering: transposon mutagenesis in Gram negative bacteria. Nat. Biotechnol. 1, 784–791 (1983).
Sambrook, J., Fritsch, E. F. & Maniatis, T. Molecular Cloning: A Laboratory Manual 2nd edn (Cold Spring Harbor Laboratory, 1989).
Roland, K., Curtiss, R. 3rd & Sizemore, D. Construction and evaluation of a delta cya delta crp Salmonella typhimurium strain expressing avian pathogenic Escherichia coli O78 LPS as a vaccine to prevent airsacculitis in chickens. Avian Dis. 43, 429–441 (1999).
Choi, K. H., Kumar, A. & Schweizer, H. P. A 10-min method for preparation of highly electrocompetent Pseudomonas aeruginosa cells: application for DNA fragment transfer between chromosomes and plasmid transformation. J. Microbiol. Methods 64, 391–397 (2006).
Acknowledgements
We thank V. de Lorenzo and E. Martinez-García for their inspiring work in recombineering methods and for sharing valuable materials. This work was supported by grants from the European Union (ERC Consolidator grant COMBAT 724290) and the excellence cluster RESIST (Resolving Infection Susceptibility; EXC 2155—project number 390874280). Furthermore, S.H. received funding from the German Research Foundation (Deutsche Forschungsgemeinschaft grant SPP 1879), the Lower Saxony Ministry for Science and Culture (BacData grant ZN3428) and the Novo Nordisk Foundation (grant NNF18OC0033946). P.I.N. received financial support from the Novo Nordisk Foundation through grants NNF20CC0035580, LiFe (NNF18OC0034818) and TARGET (NNF21OC0067996), the Danish Council for Independent Research (SWEET, DFF–research project 8021-00039B) and the European Union’s Horizon 2020 Research and Innovation Programme under grant agreement no. 814418 (SinFonia). Biorender (Biorender.com) was used to create the figures.
Author information
Authors and Affiliations
Contributions
A.A.-R., S.H. and P.I.N. designed the protocol. P.I.N. developed the original CRISPR–Cas9-bearing plasmid. D.P., N.O.G., A.A.-R., A.N. and A.M. constructed the new vector set for the toolkit. D.P., A.A.-R., N.O.G., A.N., A.M. and F.A.-R. performed the experiments and validated the protocol in laboratory strains. M.G. and D.P. performed the experiments in clinical isolates. A.A.-R., S.H. and P.I.N. supervised the project. A.A.-R., D.P, N.O.G. and S.H. wrote the protocol.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Protocols thanks Andrea Smania and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Jeske, A. et al. NPJ Biofilms Microbiomes 8, 6 (2022): https://doi.org/10.1038/s41522-022-00268-1
Borgert, S. R. et al. Nat. Commun. 13, 7402 (2022): https://doi.org/10.1038/s41467-022-35030-w
Supplementary information
Supplementary Table 1
Oligonucleotides used in this study.
Source data
Source Data Fig. 4
Unprocessed agarose gels for Fig. c–e.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Pankratz, D., Gomez, N.O., Nielsen, A. et al. An expanded CRISPR–Cas9-assisted recombineering toolkit for engineering genetically intractable Pseudomonas aeruginosa isolates. Nat Protoc 18, 3253–3288 (2023). https://doi.org/10.1038/s41596-023-00882-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-023-00882-z
This article is cited by
-
CRISPR–Cas9 applications in T cells and adoptive T cell therapies
Cellular & Molecular Biology Letters (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.