Abstract
Global riverine nitrous oxide (N2O) emissions have increased more than 4-fold in the last century. It has been estimated that the hyporheic zones in small streams alone may contribute approximately 85% of these N2O emissions. However, the mechanisms and pathways controlling hyporheic N2O production in stream ecosystems remain unknown. Here, we report that ammonia-derived pathways, rather than the nitrate-derived pathways, are the dominant hyporheic N2O sources (69.6 ± 2.1%) in agricultural streams around the world. The N2O fluxes are mainly in positive correlation with ammonia. The potential N2O metabolic pathways of metagenome-assembled genomes (MAGs) provides evidence that nitrifying bacteria contain greater abundances of N2O production-related genes than denitrifying bacteria. Taken together, this study highlights the importance of mitigating agriculturally derived ammonium in low-order agricultural streams in controlling N2O emissions. Global models of riverine ecosystems need to better represent ammonia-derived pathways for accurately estimating and predicting riverine N2O emissions.
Similar content being viewed by others
Introduction
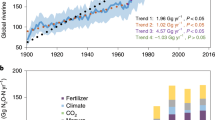
Nitrous oxide (N2O) is a potent greenhouse gas and dominant ozone-depleting substance, and its atmospheric mole fraction has increased by 23% since 17501. Riverine N2O emissions have increased fourfold since 1900 and are important components of the global N2O budget2. A 2020 global modeling study3 showed that small streams (lower-order streams rather than high-order streams) contribute up to 85% of the global riverine nitrous oxide (N2O) emissions3, most of which are produced in hyporheic zones (beneath stream beds where stream waters exchange with adjacent sediments). However, in current process-based N2O models, the N2O/N2 ratio is used to represent the N2O production rate during denitrification3. Model parameters associated with microbially mediated hyporheic N2O production are poorly represented3,4. Although the IPCC Guidelines for national greenhouse gas inventories include N2O from nitrification4, previous studies have been focused on quantifying denitrification and N2O emissions resulting from nitrate (NO3−)5,6. Furthermore, it is difficult to distinguish between nitrifier denitrification, nitrification-coupled denitrification, and heterotrophic denitrification N2O production pathways, as these processes occur via homologous genes under similar conditions, such as low O2 conditions7. Hyporheic N2O production has been shown to be very heterogeneous8,9,10, but this heterogeneity is extremely difficult to account for in models and, therefore, is often excluded5. Currently, the microbial mechanisms underlying N2O production in hyporheic exchange zones are largely unknown. These processes may significantly influence global N2O budgets and be potentially severely underestimated.
Accordingly, the aim of this study was to investigate the microbial sources and mechanisms that are responsible for hyporheic N2O production in riverbed and riparian zone sediments. We first investigated the spatiotemporal characteristics of N2O production along transects of the Baiyangdian riverine network, the largest riverine network in the North China Plains, using isotopic 15N-18O and 15N tracing, quantitative reverse-transcription PCR (RT‒qPCR), and metagenome analysis at the site and regional scales. The North China Plains cover an area of 300,000 km2 and account for 23% of the Chinese cropland area. They account for more than 25% of global fertilizer N use, making them a global N2O emission hotspot2,11,12. However, N2O emissions from agricultural streams in China13,14 or in the North China Plains specifically15,16 have only been reported in a few studies. We further compared these results to those measured globally in temperate and tropical streams to obtain comprehensive conclusions.
Results and discussion
Site-scale investigation of heterogeneous N2O emission fluxes along a transect of a riverine hyporheic zone
Over a period of 4 years, the N2O emission fluxes in the riparian zone and riverbed sediments along a transect of a riverine hyporheic zone were continuously measured using a closed-chamber method (Fig. 1a and Supplementary Data S1). The emission fluxes were significantly greater in summer than in winter (t-test, p < 0.01), but there were no significant differences between the two zones. Ammonium (NH4+) showed the most positive correlation with N2O flux, irrespective of the sampling time and zone (Supplementary Data S2 and Table S1), as reported in many previous studies on rivers, agricultural catchments, and estuaries (Supplementary Table S2). In contrast, nitrate (NO3-) was negatively correlated with N2O flux, in contrast with previous findings17,18.
a N2O emission flux over 4 years of monitoring (n = 48 independent experiments). For each box chart, the horizontal line indicates the median, the box represents the 25th and 75th percentiles, and the whisker shows the range from the 5th to the 95th percentile. b, c The microbial rate and contribution to N2O production via NH4+-derived and NO3--derived processes based on semi-in situ sediment-core incubation via the 15N tracing method (b) and slurry incubation involving the C2H2-inhibitor method (c) (n = 6 and n = 3 biologically independent samples for core and slurry incubation, respectively). Data were presented as mean values ± SEM; d Transcript abundance of N2O-production related (amoA, norB, nirS, and nirK) and N2O-reduction related (nosZ) genes (n = 3 biologically independent samples). Data were presented as mean values ± SEM. Here, amoA, ammonia monooxygenase gene encoded the NN pathway (NH4+ → NH2OH); nirSK, nitrite reductase genes (NO2- → NO) encoded the ND (nirK only), NCD and HD pathways; norB, nitric oxide reductase gene (NO → N2O) encoded the ND, NCD and HD pathways; and nosZ, nitrous oxide reductase gene (N2O → N2) encoded the NCD and HD pathways. e Schematic representation of the sampling sites along a transect of a riverine hyporheic zone.
Site-scale investigation of 15N tracing of the semi-in situ sediment core revealed NH4 +-derived and NO3 −-derived sources of hyporheic N2O production
Generally, N2O production is a biogeochemical process driven by microorganisms via two main microbial processes with four pathways: the NH4+-derived process (NH4+ as a substrate; includes nitrifier nitrification (NN), nitrifier denitrification (ND), and nitrification-coupled denitrification (NCD) pathways) and the NO3−-derived process (NO3- as a substrate; heterotrophic denitrification (HD) pathway)6,19,20,21. Our results confirmed that the contribution of biotic N2O production in our samples was more than 92–95%, and abiotic processes accounted for less than 5–8% of the total N2O production (Supplementary Data S1, S2 and Fig. S1). A 15N tracing semi-in situ sediment-core incubation was performed to investigate the semi-in situ hyporheic N2O production rate and to clarify NH4+-derived and NO3−-derived sources without distinguishing the NH4+-derived process along the transect of the Xiaoqinghe River hyporheic zone (Fig. 1b and Supplementary Data S3). Interestingly, the NH4+-derived process was the dominant hyporheic N2O source in both the riparian zone (87.3 ± 3.9%) and the riverbed sediments (92.6 ± 5.6%), and the remaining N2O production could be attributed to the NO3−-derived process.
Furthermore, we used the 0.01% C2H2-inhibitor method to confirm the above results (Fig. 1c and Supplementary Data S1, S2). First, we found that the potential rate of biotic N2O production in the riparian zone (0.61 ± 0.03 μg N2O-N kg−1 h−1) was lower than that in the riverbed zone (0.73 ± 0.09 μg N2O-N kg−1 h−1), in agreement with the 15N semi-in situ sediment-core incubation results. However, this was not in agreement with the previous, which reported a higher N2O production rate in the riparian zone in different river systems18,22,23. In the riparian and riverbed zones, NH4+-derived N2O accounted for 89.2 ± 2.9% and 76.1 ± 2.6% of the total N2O produced, respectively. The NO3−-derived process resulted in the production of the remaining N2O. These findings differ from the prevailing opinion that NO3−-derived processes are the main contributors to N2O in riverine hyporheic zones6,24.
Site-scale investigation of quantitative reverse-transcription PCR (RT‒qPCR) analysis of N2O-related gene mRNA
To further investigate the microbial mechanism and activity related to N2O production, the transcript abundances of microbial N2O-related genes were quantified via reverse-transcription PCR (RT‒qPCR) (Fig. 1d). The transcript abundances of all N2O-related (production and reduction) genes showed low heterogeneity between the riparian and riverbed zones. N2O production-related genes (amoA, norB, nirS, and nirK) had greater transcriptional abundances than the reduction gene (nosZ) in both the riparian and riverbed zones, providing evidence that the hyporheic zone had greater potential for N2O production. RT‒qPCR cannot be used to distinguish both the nirK and norB genes in the NH4+-derived and NO3−-derived N2O production pathways. However, we found that the transcript abundances of genes in the NH4+-derived N2O production pathway, including the amoA gene encoded the NN pathway and the norB gene encoded the ND and NCD pathways, were greater than those of the norB gene encoded the HD pathway, irrespective of sediment location (Fig. 1d).
Regional-scale investigation of microbial pathways and key parameters influencing hyporheic N2O production
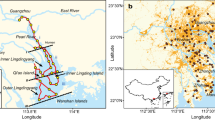
To support the above site-scale results, samples were also collected from 50 riparian zones and 50 riverbed sediments (0–20 cm depth) in 25 hyporheic zone transects at equal distances along five streams in the high- and low-water-level seasons (Fig. 2a, Supplementary Data S1, S2, and Fig. S2). The 0.01% C2H2 inhibitor results showed that N2O production in hyporheic sediments was dominated by NH4+-derived processes, the contribution of which to N2O production (72 ± 3%, n = 100) was significantly greater than that of NO3−-derived processes (28 ± 3%, n = 100) (paired t-test, p < 0.0001; Fig. 2b and Supplementary Data S2). There was less heterogeneity in hyporheic N2O emissions among the five different streams or in all streams at both the site and regional scales, but there was large temporal heterogeneity at the seasonal scale, with lower heterogeneity in winter (0.63 ± 0.22 mg N kg−1 soil d−1) than in summer (1.26 ± 0.34 mg N kg−1 soil d−1) (p < 0.05), and the values mostly depended on the NH4+ content (r = 0.851, p < 0.0001, n = 100) (Fig. 2b, c; Supplementary Fig. S2; Data S2; and Table S3). This result indicates that in hyporheic riverine sediments, microbial N2O generation is mainly driven by NH4+-derived processes rather than by NO3−-derived processes.
a Overview of the Baiyangdian riverine network and the sampling sites; b Spatiotemporal distribution of the potential rate of NH4+-derived and NO3--derived N2O production in the five streams (46–273 km length) of the Baiyangdian riverine network (n = 3 biologically independent samples). Data were presented as mean values ± SEM; c Spatiotemporal heterogeneity analyses of the potential rate and contribution of NH4+-derived and NO3−-derived pathways at the regional scale. P values were calculated with the two-tailed independent t-test (*p < 0.05, **p < 0.01 two-tailed; Except n = 44 and 56 independent experiments for upstream and downstream, respectively, n = 50 for other groups); d Potential rates and contributions of nitrifier nitrification (NN), nitrifier denitrification (ND), nitrification-coupled denitrification (NCD), and heterotrophic denitrification (HD) pathways in the sediments of the riverine hyporheic zones in the Tang River (n = 20 independent experiments). For each box chart, the horizontal line indicates the median, the box represents the 25th and 75th percentiles, and the whisker shows the range from the 5th to the 95th percentile. Bai River, B; Juma River, J; Cao River, C; Tang River, T; Zhulong River, Z; and Riparian zone, R; riverbed zone, O.
Furthermore, we used improved 15N-18O dual-isotope tracing to verify these results in one of the five streams, the Tang River. Based on our findings, NH4+-derived N2O production (73 ± 5%, n = 10) was significantly greater than NO3--derived N2O production (27 ± 5%, n = 10) (paired t-test, p < 0.0001; Fig. 2d; Supplementary Fig. S2; and Data S2, S4), which was consistent with the above 0.01 % C2H2-inhibitor results. Among the NH4+-derived pathways, the ND pathway was dominant upstream (38 ± 4%, n = 4), downstream (44 ± 3%, n = 6), in riparian zone sediments (45 ± 3%, n = 5), and in riverbed sediments (38 ± 4%, n = 5), with little heterogeneity at both the site and regional scales. The remaining NH4+-derived N2O production was attributed to the NN and NCD pathways, with a range of 8–23%. These results showed that ND was an important pathway for N2O generation, which has long been overlooked but has recently been widely reported25,26,27,28,29. However, the biogeochemical mechanism underlying the role and key factors influencing this pathway are still unknown.
Regional-scale investigation of potential metabolic N2O production mechanism
As previously mentioned, the nir and nor genes are involved in the ND, NCD, and HD pathways; as a result, these processes cannot be distinguished based on the presence of these genes. To resolve this issue, species annotation was combined with functional gene annotation to study the genes related to nitrogen cycling in nitrifying and denitrifying bacteria and distinguish between the nir and nor genes in ND, NCD, and HD. The assembly and binning processes generated 198 high-quality (completeness >75%, contamination <15%) metagenome-assembled genomes (MAGs), 111 of which encoded genes involved in N2O production and reduction (Supplementary Data S5, S6). Through species and functional gene annotation, we obtained eight MAGs containing amoABC, hao, or nxrAB genes, which have been identified as nitrifying bacteria, and seven MAGs that contained the nirK/S gene, identified as denitrifying bacteria30,31. The annotation results of the species and functional genes were further verified via phylogenetic analysis (Fig. 3a). The relative abundances of nitrifying and denitrifying bacteria were 9524 and 8310 TPM, respectively (Fig. 3b).
a Phylogenetic diagram of the genome associated with metagenomic recombinant binning. Based on the 16S rRNA sequence, phylogenetic relationships were calculated using the maximum likelihood method. The sequences associated with the N cycle based on the NCBI database and metagenome-assembled genomes (MAGs) identified in this study are shown in black and red, respectively. Each MAG was labeled as bin.“MAG source”.“MAG number” and included River (Bai River, B; Juma River, J; Cao River, C; Tang River, T and Zhulong River, Z), Site (1, 2, 3, 4, and 5), Zone (Riparian zone, R; Riverbed zone, O), and Season (Summer, S; Winter, W). The species, abundance, and metabolic path diagrams of nitrifiers and denitrifiers are shown above and below the orange dotted line. b Relative abundance of 15 MAGs and gene recombination and characterization of N2O production- and metabolism-related microbial processes. The cyan bar graph represents the relative abundance (TPM) of 15 MAGs. Different colors in the grid represent different gene abundances (TPMs). c Metabolic relationship of substances related to N2O production. The bin.T2B.7 genome and the bin.B5Z.4 genome represent nitrifiers and denitrifiers, respectively. The black arrows in the figure indicate material transfer, transformation, or electron transport. The yellow, green, brown, and pink arrows represent the NN, ND, NCD, and HD pathways, respectively, in the nitrogen cycle. Filled blocks of different shapes or colors represent different enzymes in cells.
Subsequently, the abundances of the N2O-producing genes amoABC, hao, nirSK, and nor in nitrifying and denitrifying bacteria were determined (Fig. 3b and Supplementary Data S6). Traditionally, NH4+ oxidation is the rate-limiting step in nitrification32,33,34 and in the N cycle35. However, both the total and individual abundances of N2O-producing genes in nitrifying bacteria (26, 8, 48, and 6 TPM, respectively) were significantly greater than those in denitrifying bacteria (0, 0, 6, and 4 TPM, respectively). In contrast, the abundance of the N2O-reducing gene nosZ (N2O → N2) was 4 TPM in denitrifying bacteria, but it was not detected in nitrifying bacteria (Fig. 3b). These results further revealed the mechanism of microbial N2O production and highlighted the dominant role of nitrifying bacteria (NH4+-derived pathways) in N2O production.
The microbial mechanism of NH4+-derived and NO3--derived N2O production was further revealed from the perspective of energy metabolism by metagenome binning analysis and is shown as a single-cell draft (Fig. 3c). The transmembrane transport of nitrite and nitrate, which are necessary substrates for ND and HD, respectively, is an energy-consuming process36,37. The NO3− and NO2− produced by nitrifying bacteria can be directly used in the ND pathway as substrates38, whereas denitrifying bacteria in the HD process need to consume adenosine triphosphate (ATP) to absorb NO3− from the environment36. Additionally, the nirKS gene is significantly more abundant in nitrifying microorganisms than in denitrifying microorganisms (p = 0.003); hence, in the consumption of NO2− for detoxification39,40,41, the ND pathway has greater potential to alleviate the toxic effects of nitrite on microorganisms39,40,41 than does the HD pathway. Furthermore, nitrifiers have the potential to produce large amounts of N2O because, based on reports to date, nitrifying bacteria do not contain N2O reductase42. Denitrifying bacteria have the nosZ gene encoding N2O reductase; hence, the N2O produced during NCD and HD can be further reduced to N243. In contrast, there is no nosZ gene in nitrifying bacteria, and the N2O produced by NN and ND is not consumed within the same cells7,27,44 and can be released. Overall, there is greater potential for N2O production from the NH4+-derived pathways than from the NO3−-derived pathway because of energy savings, NO2− detoxification, and the absence of N2O reductase.
Global-scale investigation of hyporheic N2O production in streams
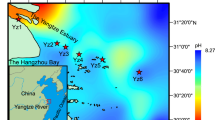
The above findings indicate that NH4+-derived pathways, rather than NO3−-derived pathways, largely influence hyporheic N2O production in the Yangtze, Yellow, Pearl, Yarlung Zangbo, Huai, Liao, Songhuajiang, and Heilongjiang river basins. To further expand our survey, we carried out a global spatiotemporal investigation across a wide range of streams around the world using the 0.01% C2H2 inhibitor method (Supplementary Data S1). Analysis conducted on a spatial scale showed that the rates of NH4+-derived N2O production were significantly greater than those of the NO3−-derived pathway at both the regional (4.56 ± 1.20 vs. 3.60 ± 1.68 μg N kg−1 d−1, n = 27) and global scales (6.00 ± 1.20 vs. 2.40 ± 0.72 μg N kg−1 d−1, n = 11, p = 0.003) (Fig. 4a, b and Supplementary Table S1). Seasonal scale analysis in the Buerhatong River (15–22 °C) and Xiaoqing River (14–23 °C) also revealed that the NH4+-derived N2O production rates (8.40 ± 2.40 μg N kg−1 d−1) were substantially greater than the NO3--derived rates (3.60 ± 1.68 μg N kg−1 d−1; n = 4, p = 0.001) and were the main contributors to N2O production (67.5 ± 6.7%) (Fig. 4c). Statistical analysis at the spatial and temporal scales revealed a similar trend, namely, that NH4+-derived pathways contributed most significantly to N2O production (64.7 ± 2.9%) (Fig. 4d). In summary, NH4+-derived pathways are the dominant hyporheic N2O sources in low-order agricultural streams around the world.
a–c The locations of the pie charts show the sampling sites; the sites for temporal-scale analysis are marked with solid orange circles. Detailed information on the sampling sites can be found in Supplementary Data S1. For each sampling site, the pie chart shows the total N2O production rate. The pie chart colors represent the relative contributions of the NH4+-derived (blue) and NO3−-derived (yellow) pathways to total N2O production. d Box charts (the horizontal line indicates the median, the box indicates the 25th and 75th percentiles, the whisker shows the range from the 5th to 95th percentiles, and the colors show the N2O production pathways) indicating the potential rate of N2O production via NH4+-derived and NO3--derived pathways and their related contributions in riverine hyporheic zones worldwide (n = 41 independent experiments).
Based on our results, the NH4+-derived process, rather than the NO3−-derived process, is the dominant hyporheic N2O source in lower-order agricultural streams. Together, these findings provide insights into better estimation of N2O emissions in global models of riverine ecosystems, which has been under debate for many years because of either under- or overestimation of riverine N2O budgets in IPCC assessments2,5,39. Furthermore, the results emphasize the importance of managing ammonium11. This is particularly important, as China has long been the largest ammonia fertilizer consumer in the world12. Approximately 60% of the fertilizer applied is not utilized by crops but leaches into stream systems, stimulating global climate warming; that is, the climate benefits of increased CO2 uptake via crop ammonia application are outweighed by stimulated greenhouse gas production from ammonia pollution12. Therefore, it is necessary to optimize fertilizer and ammonia management.
Methods
Study site and sampling
The Baiyangdian riverine network (113°40’–116°20’E, 38°10’–40°10’N) in the Haihe River Basin, with a total area of 3.12 × 104 km2, is the largest riverine network in the North China Plains. The river network density is 0.50–0.99 km/km2, representing the upper level in China, with significant temporal distribution differences. Its upper and middle regions receive 30–40 first- or second-order tributaries or small streams with a total length of ~3000 km. These small streams drain various land areas in agricultural, urban, and mountain regions and are affected by various human activities and agricultural fertilization practices. Their catchment areas exceeded 5.5% of the agricultural area. First, a site-scale study was conducted on one of these small streams, the Xiaoqinghe River (Supplementary Data S1). Nine parallel in situ sediment cores (0–20 cm depth) were collected separately from the riparian (2 m away from the water‒soil transition zone) and riverbed (the middle of the river) zones. Six sediment cores were subjected to semi-in situ 15N-tracer assays, and the other three were subjected to 0.01% C2H2 inhibitor assays and molecular analysis. Furthermore, a regional-scale study was conducted on five rivers in the Baiyangdian riverine network (Fig. 2a and Supplementary Data S1). A total of 100 sediment samples were collected in the riparian zone and from riverbed sediments in the five rivers at five sampling sites along each river during the dry (Jan to Mar 2021) and rainy (Aug to Oct 2021) seasons. Then, we chose steams located adjacent to farmlands as the study sites, characterized by their land use type being cropland land45. Twenty-eight low-order agricultural streams were sampled in major river basins worldwide, including the Mississippi River, Colorado River (North America), Amazon River, Ucayali River (South America), Elbe River, Weser River, Po River (Europe), Niger River, Zambezi River (Africa), Murray River (Oceania), Indus River, Yangtze River, Yellow River, Pearl River, Yarlung Zangbo River, Huai River, Liao River, Songhuajiang River, and Heilongjiang River (Asia) (Supplementary Data S1).
All the sediment cores were collected in triplicate using an auger (Beijing New Landmark Soil Equipment, Beijing, China) with a plexiglass tube (5.5 cm diameter, 20 cm height). The site-scale sediment cores were stored in individual plexiglass tubes, with both nozzles of each plexiglass tube sealed with caps. The regional-scale and global-scale sediment cores were placed in individual sterile plastic bags. The collected sediment cores were immediately transported to a laboratory at 4 °C for subsequent analyses (Supplementary Data S1).
15N-tracer assay for semi-in situ sediment-core incubation
The potential rates of NH4+-derived and NO3−-derived N2O production were measured via the 15N-tracer semi-in situ incubation method for the sediment core. Once the cores were at the laboratory, the gas in the headspace of the plexiglass tube was evacuated and then adjusted to standard atmospheric pressure using high-purity Ar (99.99%; Beijing Huayuan Gas, Beijing, China). The oxygen content was immediately adjusted by the injection of a known volume of high-purity O2 (99.99%; Beijing Huayuan Gas) to the site oxygen content after the same volume of Ar was withdrawn from the headspace to balance the pressure. Two treatments for 15N enrichment were applied in triplicate: (i) 15NO3− (15N at 99.19%) + 14NH4+ and (ii) 15NH4+ (15N at 99.16%) + 14NO3−. In each treatment, final enrichments of 15N-NH4+/NO3− were added to 8.0 atom% 15N excess. The final N concentrations of both NH4+ and NO3− remained close to the in situ concentrations in the sediments. The 15N-enriched cores were incubated under site temperature and oxygen conditions in a constant-temperature incubator. At defined intervals (0, 18, 36, 72, and 144 h), headspace gas was collected by using a 25-mL gas-tight syringe (Agilent, USA) and transferred into a 12-mL vacuum exetainer (Labco, UK). The 15NO3− in the sediments was first converted by sponge cadmium (1.0 g) into 15NO2− and then to 29N2. The 15NH4+ in the sediments was converted by hypobromite into 30N2.
15N-18O tracer assay for N2O production
A slightly modified version of a previously reported improved 15N-18O dual-isotope tracing method7,19,20,21 was used to carry out incubation experiments under site oxygen and temperature conditions. The improved 15N-18O dual tracing method was applied to 20 sediment samples from the Tanghe River, one of the five rivers in the Baiyangdian River. After visible roots and plant residues were removed, 5 g of homogeneous fresh sediment with 25 ml of overlying water close to the sediment (5:1, v/w) was placed into a 60-mL glass serum vial (Ochs Laborbedarf, Germany). Four treatments enriched in 18O and 15N were applied in triplicate: (i) H218O (18O at 97.2%) + NH4+ + NO3−, (ii) N18O3− (18O at 96.3%) + NH4+ + NO3−, (iii) 15NO3− (15N at 99.19%) + NH4+ + NO3−, and (iv) 15NH4+ (15N at 99.16%) + NH4+ + NO3−. In all treatments, final enrichments of 18O-H2O/NO3− and 15N-NH4+/NO3− were added to 1.0 atom% 18O and 10 atom% 15N excess, respectively. The final N concentrations of both NH4+ and NO3− were the site N concentrations in the fresh sediments. All vials were sealed with plugs (Ochs Laborbedarf, Germany) and aluminum crimp cap (Agilent, USA) septum-equipped lids during the incubation period. Subsequently, the gas in the vials was evacuated and then adjusted to standard atmospheric pressure using high-purity Ar (99.99%). Subsequently, the O2 concentration was immediately adjusted by the injection of a known volume of high-purity O2 (99.99%) to the site O2 concentration after the same volume of Ar was evacuated from the headspace to balance the pressure. The treatments were incubated at 60 rpm for 36 h to guarantee complete exposure of the sediments to all substrates under site temperatures. At the end of the incubation, the gas samples were transferred to 12-mL vacuum exetainers (Labco, UK) to quantify N2O concentrations.
0.01% C2H2 inhibitor assay
Three treatments were applied to the sediments at each scale in triplicate: (i) no treatment (control), (ii) treatment with 0.01% C2H2 (v/v; ammonia oxidation inhibitor), and (iii) treatment with ZnCl2 (600 μl, 7 M; biotic process inhibitor). The incubation conditions were the same as those in the 15N-18O experiment. The headspace gas was sampled with a locked syringe equipped with a Luer lock valve (25.0 ml; Agilent) and injected into a 12.0 ml vacuumed glass serum vial after 0, 3, 12, 24, and 36 h.
N2O concentration, production rate, and contribution of 15N and 18O tracer assays
The N2O concentration was determined via a gas chromatograph (7890 A, Agilent, USA) with an autosampler (precisions ± 2.8%), while linear regression was carried out for the potential N2O production rate with incubation time, with coefficients of determination (R2) greater than 0.80.
The 15N and 18O signatures of N2O in gas samples were measured using an isotope-ratio mass spectrometer (IRMS and Precon, Delta V Advantage, Thermo Fisher Scientific, Bremen, Germany; precisions <0.04 ‰ δ15N and <0.07 ‰ δ18O, respectively). The relative contributions of microbial N2O production pathways (NH4+-derived pathways, NN, ND, and NCD; NO3−-derived pathway, HD) were calculated according to the methods of refs. 20,21.
RNA extraction and quantitative reverse-transcription (RT‒qPCR) assays
In the site-scale study, RNA was extracted from three parallel sediment cores from both the riparian zone and riverbed sediments using an RNeasy Power Microbiome RNA Isolation Kit (QIAGEN, Hilden, Germany) according to the manufacturer’s protocol. RNA quality and concentration were estimated using a NanoDrop 2000 Spectrophotometer (NanoDrop Technologies, Wilmington, DE, USA).
Reverse transcription was performed with a PrimeScriptTM RT Reagent Kit with gDNA Eraser (Perfect Real Time) (TaKaRa, Dalian, China). All RT-qPCR analyses were performed on a sequence detection system (ABI 7500; Applied Biosystems, Foster City, CA, USA) with SYBR-Green fluorescent dye (TaKaRa). The copy numbers of N2O-related genes were quantified by using the specific primers for the archaeal amoA gene with Arch-amoAF and Arch-amoAR, the bacterial amoA gene with amoA-1F and amoA-2R, the denitrifier and ammonia oxidizer nirK gene with nirK-876F and nirK-1040R, the nirS gene with nirS-F and nirS-R, the denitrifier and ammonia oxidizer norB gene with norB-F and norB-R, and the nosZ gene with nosZ-F and nosZ-R. All tests were conducted in triplicate with amplification efficiencies between 90% and 110% and correlation coefficients (R2) above 0.98. More details on the primers and thermal profiles are listed in Supplementary Table S4.
DNA extraction, metagenomic library sequencing, N2O-related genome binning, taxonomic classification, functional annotation, and phylogenetic analyses
DNA was extracted from 100 regional-scale sediment cores by using the FastDNA Spin Kit for Soil (MP Biomedicals, Solon, OH, USA) according to the manufacturer’s protocol. DNA quality and concentration were estimated using a NanoDrop 2000 Spectrophotometer (NanoDrop Technologies). Approximately 1.5 μg of extracted DNA (per sample) was used for metagenomic library preparation and subsequent sequencing on the Illumina PE150 platform (150-bp paired-end) with a sequencing depth of 10 G. Clean data were generated from raw metagenomic reads after low-quality nucleotides and reads with any ambiguous base calls were filtered out using Kneaddata (github.com/biobakery/kneaddata), and the data were subsequently quality-checked using FastQC (Babraham Bioinformatics, Babraham Institute, Cambridge, UK) (Supplementary Data S5). The quality-controlled clean data were used to obtain contigs using Megahit46, while Bowtie247 and Samtools48 were used for comparison and format conversion, respectively.
Metagenome binning was performed with contigs above 1500 bp using MetaWRAP (v 1.2.1)49. The obtained metagenome-assembled genomes (MAGs) were purified with RefineM50 to remove contaminating contigs (Supplementary Data S6). The MAGs were quality-checked by using CheckM51 and were dereplicated with dRep software52, and 198 MAGs with a degree of completion greater than 75% and a degree of contamination less than 15% were selected for subsequent analysis52. The Quant_bins module in MetaWRAP (salmon algorithm)53 was used to calculate the average relative abundance. The taxonomic affiliation of the MAGs was determined by GTDB-Tk v2.3.054. Functional gene and protein annotation for the 198 MAGs was performed against the KEGG, NCyc55, COG, and GO databases at an e value <1e−5. According to taxonomy affiliation and functional gene and protein annotation, the MAGs containing amo, hao, or nxrAB genes and those containing the nirKS56,57 gene were identified to belong to nitrifying and denitrifying bacteria, respectively. A phylogenetic tree of high-quality MAGs was generated by the maximum likelihood statistical method with the nearest-neighbor-interchange (NNI) ML heuristic method in MEGA 11 software58.
Analysis of environmental variables
The sediment NH4+, NO3−, NO2−, TOM, TN, and TP concentrations were determined59,60. The oxygen concentration and temperature in sediments were measured in situ by using a Pocket Oxygen Meter (FireStingGO2, PyroScience GmbH, Germany). The details are shown in the Supplementary Information.
Statistical analysis
The mean values, t-tests, Spearman’s correlations, and linear regression analyses were performed by using Statistical Product and Service Solutions 18.0 software (SPSS Inc., USA). The significance level was to α = 0.05 (p value ≤0.05).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data generated in this study are provided within the article, Supplementary Information, and Supplementary Data files. Metagenomic sequencing data and metagenome-assembled genomes are available in the NCBI Sequence Read Archive (SRA) under the accession codes PRJNA943572 and PRJNA1031250, respectively.
Change history
28 May 2024
A Correction to this paper has been published: https://doi.org/10.1038/s41467-024-49020-7
References
Canadell J. G. et al. In Climate Change 2021: The Physical Science Basis. Contribution of Working Group I to the Sixth Assessment Report of the Intergovernmental Panel on Climate Change (eds Brovkin V. & Feely R. A.) Ch. 5 (Cambridge Univ. Press, 2021).
Tian, H. Q. et al. A comprehensive quantification of global nitrous oxide sources and sinks. Nature 586, 248–256 (2020).
Yao, Y. Z. et al. Increased global nitrous oxide emissions from streams and rivers in the Anthropocene. Nat. Clim. Change. 10, 138–139 (2020).
IPCC. in 2019 Refinement to the 2006 IPCC Guidelines for National Greenhouse Gas Inventories. (2019).
Maavara, T. et al. Nitrous oxide emissions from inland waters: Are IPCC estimates too high? Global Change Biol. 25, 473–488 (2019).
Beaulieu, J. J. et al. Nitrous oxide emission from denitrification in stream and river networks. Proc. Natl Acad. Sci. USA 108, 214–219 (2011).
Zhu, X., Burger, M., Doane, T. A. & Horwath, W. R. Ammonia oxidation pathways and nitrifier denitrification are significant sources of N2O and NO under low oxygen availability. Proc. Natl Acad. Sci. USA 110, 6328–6333 (2013).
Biddulph, M. In Geomorphological Techniques (eds Clarke, L. & Nield, J.) Ch 3.11.1 (British Society for Geomorphology, 2015).
Kiel, B. A. & Cardenas, M. B. Lateral hyporheic exchange throughout the Mississippi River network. Nat. Geosci. 7, 413–417 (2014).
Boulton, A. J., Findlay, S., Marmonier, P., Stanley, E. H. & Valett, H. M. The functional significance of the hyporheic zone in streams and rivers. Annu. Rev. Ecol. Syst. 29, 59–81 (1998).
Reay, D. S. et al. Global agriculture and nitrous oxide emissions. Nat. Clim. Change. 2, 410–416 (2012).
Tian, H. Q. et al. Food benefit and climate warming potential of nitrogen fertilizer uses in China. Environ. Res. Lett. 7, 044020 (2012).
Wang, J., Chen, N., Yan, W., Wang, B. & Yang, L. Effect of dissolved oxygen and nitrogen on emission of N2O from rivers in China. Atmos. Environ. 103, 347–356 (2015).
Kumar, A., Yang, T. & Sharma, M. P. Greenhouse gas measurement from chinese freshwater bodies: a review. J. Clean. Prod. 233, 368–378 (2019).
Tang, M.-Y. et al. Diffusive fluxes and controls of N2O from coastal rivers in Tianjin City. Environ. Sci. 43, 1481–1491 (2022).
Gong, J.-W. Temporal and spatial distribution of CO2、CH4 and N2O concentration and emission rate in the Chaobai River Basin. Beijing Jiaotong University. https://doi.org/10.26944/d.cnki.gbfju.2022.000376 (2022).
Hirayama, J., Eda, S., Mitsui, H. & Minamisawa, K. Nitrate-dependent N2O emission from intact soybean nodules via denitrification by Bradyrhizobium japonicum bacteroids. Appl. Environ. Microbiol. 77, 8787–8790 (2011).
Hefting, M. M., Bobbink, R. & Janssens, M. P. Spatial variation in denitrification and N2O emission in relation to nitrate removal efficiency in a n-stressed riparian buffer zone. Ecosystems 9, 550–563 (2006).
Wrage, N., van Groenigen, J. W., Oenema, O. & Baggs, E. M. A novel dual-isotope labelling method for distinguishing between soil sources of N2O. Rapid Commun. Mass Spectrom. 19, 3298–3306 (2005).
Kool, D. M. et al. Nitrifier denitrification can be a source of N2O from soil: a revised approach to the dual-isotope labelling method. Eur. J. Soil Sci. 61, 759–772 (2010).
Kool, D. M., Dolfing, J., Wrage, N. & Van Groenigen, J. W. Nitrifier denitrification as a distinct and significant source of nitrous oxide from soil. Soil Biol. Biochem. 43, 174–178 (2011).
Wang, H. J., Wang, W. D., Yin, C. Q., Wang, Y. C. & Lu, J. W. Littoral zones as the “hotspots” of nitrous oxide (N2O) emission in a hyper-eutrophic lake in China. Atmos. Environ. 40, 5522–5527 (2006).
Wang, H. J., Yang, L. Y., Wang, W. D., Lu, J. W. & Yin, C. Q. Nitrous oxide (N2O) fluxes and their relationships with water-sediment characteristics in a hyper-eutrophic shallow lake, China. J. Geophys. Res. Biogeosci. 112, G01005 (2007).
Woodward, K. B., Fellows, C. S., Conway, C. L. & Hunter, H. M. Nitrate removal, denitrification and nitrous oxide production in the riparian zone of an ephemeral stream. Soil Biol. Biochem. 41, 671–680 (2009).
Qin, Y., Wang, S. Y., Wang, X. M., Liu, C. L. & Zhu, G. B. Contribution of ammonium-induced nitrifier denitrification to N2O in paddy fields. Environ. Sci. Technol. 57, 2970–2980 (2023).
Zhang, G. L. et al. Distribution of concentration and stable isotopic composition of N2O in the shelf and slope of the Northern South China Sea: implications for production and emission. J. Geophys. Res. Oceans 124, 6218–6234 (2019).
Yuan, D. D. et al. Nitrifiers cooperate to produce nitrous oxide in Plateau wetland sediments. Environ. Sci. Technol. 57, 810–821 (2023).
Cao, Y., Wang, X., Zhang, X., Misselbrook, T. & Ma, L. Nitrifier denitrification dominates nitrous oxide production in composting and can be inhibited by a bioelectrochemical nitrification inhibitor. Bioresour. Technol. 341, 125851 (2021).
Wells, N. S. & Eyre, B. D. Flow regulates biological NO3- and N2O production in a turbid sub-tropical stream. Geochim. Cosmochim. Acta 306, 124–142 (2021).
Daebeler, A. et al. Rapid nitrification involving comammox and canonical Nitrospira at extreme pH in saline-alkaline lakes. Environ. Microbiol. 25, 1055–1067 (2023).
Blaszczyk, M. K. Comparison of dentrification by Paracoccus denitrificans, Pseudomonas stutzeri and Pseudomonas aeruginosa. Acta Microbiol. Pol. 41, 203–210 (1992).
Kowalchuk, G. A. & Stephen, J. R. Ammonia-oxidizing bacteria: a model for molecular microbial ecology. Annu. Rev. Microbiol. 55, 485–529 (2001).
Wang, B. et al. Differential contributions of ammonia oxidizers and nitrite oxidizers to nitrification in four paddy soils. ISME J. 9, 1062 (2015).
Wang, B. et al. Expansion of Thaumarchaeota habitat range is correlated with horizontal transfer of ATPase operons. ISME J. 13, 3067–3079 (2019).
Wang, S., Wang, Y., Feng, X., Zhai, L. & Zhu, G. Quantitative analyses of ammonia-oxidizing archaea and bacteria in the sediments of four nitrogen-rich wetlands in China. Appl. Microbiol. Biot. 90, 779–787 (2011).
Moir, J. W. & Wood, N. J. Nitrate and nitrite transport in bacteria. Cell Mol. Life Sci. 58, 215–224 (2001).
Fukuda, M. et al. Structural basis for dynamic mechanism of nitrate/nitrite antiport by NarK. Nat. Commun. 6, 7097 (2015).
Kim, S. W., Miyahara, M., Fushinobu, S., Wakagi, T. & Shoun, H. Nitrous oxide emission from nitrifying activated sludge dependent on denitrification by ammonia-oxidizing bacteria. Bioresour. Technol. 101, 3958–3963 (2010).
Peng, Y. Z. & Zhu, G. B. Biological nitrogen removal with nitrification and denitrification via nitrite pathway. Appl. Microbiol. Biotechnol. 73, 15–26 (2006).
Lu, X. et al. Significant production of nitric oxide by aerobic nitrite reduction at acidic pH. Water Res. 230, 119542 (2023).
Liu, B. et al. High nitrite concentration accelerates nitrite oxidizing organism’s death. Water Sci. Technol. 77, 2812–2822 (2018).
Stein, L. Y. et al. Whole-genome analysis of the ammonia-oxidizing bacterium, Nitrosomonas eutropha C91: implications for niche adaptation. Environ.Microbiol. 9, 2993–3007 (2007).
Kuypers, M., Marchant, H. & Kartal, B. The microbial nitrogen-cycling network. Nat. Rev. Microbiol. 16, 263–276 (2018).
Shaw, L. J. et al. Nitrosospira spp. can produce nitrous oxide via a nitrifier denitrification pathway. Environ. Microbiol. 8, 214–222 (2006).
Lu, M. et al. A cultivated planet in 2010 – Part 1: the global synergy cropland map. Earth Syst. Sci. Data 12, 1913–1928 (2020).
Li, D., Liu, C., Luo, R., Sadakane, K. & Lam, T. MEGAHIT: an ultra-fast single-node solution for large and complex metagenomics assembly via succinct de bruijn graph. Bioinformatics 31, 1674–1676 (2015).
Langmead, B. & Salzberg, S. Fast gapped-read alignment with Bowtie 2. Nat. Methods. 9, 357–359 (2012).
Li, H. et al. The sequence alignment/map format and SAM tools. Bioinformatics 25, 2078–2079 (2009).
Uritskiy, G. V., DiRuggiero, J. & Taylor, J. MetaWRAP-A flexible pipeline for genome-resolved metagenomic data analysis. Microbiome 6, 1–103 (2018).
Parks, D. H. et al. Recovery of nearly 8,000 metagenome-assembled genomes substantially expands the tree of life. Nat. Microbiol. 2, 1533–1542 (2017).
Parks, D. H., Imelfort, M., Skennerton, C. T., Hugenholtz, P. & Tyson, G. W. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome. Res. 25, 1043–1055 (2015).
Olm, M. R., Brown, C. T., Brooks, B. & Banfield, J. F. Drep: a tool for fast and accurate genomic comparisons that enables improved genome recovery from metagenomes through de-replication. ISME J. 11, 2864–2868 (2017).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods. 14, 417–419 (2017).
Parks, D. H. et al. A standardized bacterial taxonomy based on genome phylogeny substantially revises the tree of life. Nat. Biotechnol. 36, 996–1004 (2018).
Tu, Q., Lin, L., Cheng, L., Deng, Y. & He, Z. NCycDB: a curated integrative database for fast and accurate metagenomic profiling of nitrogen cycling genes. Bioinformatics 35, 1040–1048 (2019).
Braker, G., Zhou, J., Wu, L., Devol, A. H. & Tiedje, J. M. Nitrite reductase genes (nirK and nirS) as functional markers to investigate diversity of denitrifying bacteria in pacific northwest marine sediment communities. Appl. Environ. Microbiol. 66, 2096–2104 (2000).
Braker, G., Fesefeldt, A. & Witzel, K. P. Development of PCR primer systems for amplification of nitrite reductase genes (nirK and nirS) to detect denitrifying bacteria in environmental samples. Appl. Environ. Microbiol. 64, 3769–3775 (1998).
Hall, B. G. Building phylogenetic trees from molecular data with MEGA. Mol. Biol. Evol. 30, 1229–1235 (2013).
Wang, S. et al. Microbial nitrogen cycle hotspots in the plant-bed/ditch system of a constructed wetland with N2O mitigation. Environ. Sci. Technol. 52, 6226–6236 (2018).
Zhu, G. B. et al. Hotspots of anaerobic ammonium oxidation at land-freshwater interfaces. Nat. Geosci. 6, 103–107 (2013).
Acknowledgements
This research is financially supported by the Strategic Priority Research Program of the Chinese Academy of Sciences (Grant No. XDB0750400), the National Natural Science Foundation of China (42177063 and 92251304), and the Special Project on eco-environmental technology for peak carbon dioxide emissions and carbon neutrality (RCEES-TDZ-2021-20). The authors, Shanyun Wang and Guibing Zhu, gratefully acknowledge the Program of the Youth Innovation Promotion Association of the Chinese Academy of Sciences. The authors would like to thank all the persons for sampling.
Author information
Authors and Affiliations
Contributions
The project was conceived and led by G.Z. and S.W. S.W., B.L., L.Y., M.X., L.J., Y.Q., Y.J., Y.Z., G.A., J.M., M.W., and Y.Z. contributed to the sample chemical, molecular, and isotopic analysis. G.Z. and S.W. wrote the manuscript, and M.S.M.J., H.T., and Y.-G.Z. substantially contributed by commenting upon and revising it. All authors discussed and interpreted the results and contributed to the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Qingling Fu and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, S., Lan, B., Yu, L. et al. Ammonium-derived nitrous oxide is a global source in streams. Nat Commun 15, 4085 (2024). https://doi.org/10.1038/s41467-024-48343-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-024-48343-9
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.