Abstract
Neural stem and progenitor cell (NSPC) maintenance is essential for ensuring that organisms are born with proper brain volumes and head sizes. Microcephaly is a disorder in which babies are born with significantly smaller head sizes and cortical volumes. Mutations in subunits of the DNA organizing complex condensin have been identified in microcephaly patients. However, the molecular mechanisms by which condensin insufficiency causes microcephaly remain elusive. We previously identified conserved roles for condensins in repression of retrotransposable elements (RTEs). Here, we show that condensin subunit knockdown in NSPCs of the Drosophila larval central brain increases RTE expression and mobility which causes cell death, and significantly decreases adult head sizes and brain volumes. These findings suggest that unrestricted RTE expression and activity may lead to improper brain development in condensin insufficient organisms, and lay the foundation for future exploration of causative roles for RTEs in other microcephaly models.
Similar content being viewed by others
Introduction
Brain development in both the fruit fly, Drosophila melanogaster, and mammals relies on the ability of neural stem and precursor cells (NSPCs) to asymmetrically divide (reviewed in refs. 1,2). This asymmetric division allows for self-renewal and the generation of additional cells which will differentiate into neurons or glia, thus allowing for a small number of stem cells to give rise to hundreds of cells that populate the adult brain and increase brain volume. Abnormal brain development, accompanied by changes in brain volume and head size, is a common characteristic of several neurodevelopmental disorders, including autism spectrum disorder (ASD), attention deficit hyperactivity disorder (ADHD), Rett syndrome, and microcephaly3,4,5,6,7,8,9,10,11. However, the underlying mechanisms responsible for the change in brain volume are not well understood.
Biallelic mutations and deletions of genes encoding subunits of the condensin complex I and condensin complex II have been identified in patients with microcephaly12,13,14,15,16. Furthermore, conditional, brain-specific knockout17 or mutation12 of condensin subunits caused reduced cortical volume and microcephaly in mice. While these studies revealed increased DNA damage and cell death in condensin-deficient neural progenitor cells, the exact causes of these phenotypes remain unclear. Condensins interact with DNA to promote efficient chromatin condensation and ensure that equal division of genetic material occurs during mitosis18,19,20,21,22,23. Our lab previously identified roles for condensin I and condensin II in the repression of retrotransposable elements (RTEs) in both human cells24,25 and Drosophila cells and tissues26. RTEs are DNA elements that can copy and paste themselves into other areas of the genome through the process of retrotransposition which involves the generation of cDNA by an RTE-encoded reverse transcriptase27. RTEs are present in most organisms, and they make up approximately 40% of the human genome28 and 30% of the Drosophila genome29,30. Interestingly, increased RTE expression and/or activity has been observed in tissues of patients with several different neurodevelopmental disorders. Rett syndrome patients harbor mutations in MECP2, which contributes to DNA methylation of Long Interspersed Element 1 (L1) RTE copies; L1 copies were increased in postmortem brain tissue from patients with Rett syndrome31,32. L1 expression was also increased in autism spectrum disorder (ASD) patient tissue33, and endogenous retrovirus expression was increased in attention deficit with hyperactivity disorder (ADHD) and ASD brains34,35. However, increased RTE expression and activity have not been investigated as a potential causative mechanism for the development of abnormal brain volumes associated with neurodevelopmental disorders.
In this study, we used a newly developed Drosophila model of microcephaly to investigate whether the unrestricted expression and activity of RTEs caused by condensin subunit insufficiency is responsible for the observed decreases in adult brain volume and head size. Our data show that significant loss of brain volume is first observed in condensin-insufficient pupae, and that condensin knockdown in NSPCs in the third instar larval central brain is necessary for the observed phenotypes. Excitingly, we show that RTE expression and mobility are increased in NSPCs in the third instar larval central brain. Like the previous studies in mammals17, our results also demonstrate increased cell death in these NSPCs. Interestingly, increased levels of retrotransposition and increased cell death are rescued when condensin-insufficient larvae are allowed to develop on food containing nucleoside reverse transcriptase inhibitors (NRTIs) which inhibit RTE activity36,37. Finally, we show that condensin-insufficient larvae which are fed NRTIs can develop into adults with head sizes that are not significantly different from controls. These findings suggest that unrestricted RTE expression and mobility in condensin-insufficient NSPCs of the developing Drosophila central brain can lead to high levels of cell death, resulting in significant loss of brain volume and the development of microcephaly. Our studies, therefore, identify RTE-mediated cell death as a potential cause of microcephaly in Drosophila, and lay the foundation for investigation of this mechanism in human patient tissue and mammalian models of neurodevelopmental disorders.
Results
Condensin knockdown results in microcephaly in Drosophila
Similar to humans with condensin mutations, mutation or conditional knockout of condensin proteins in the developing murine brain causes microcephaly12,17. To determine whether the knockdown of condensin proteins results in microcephaly in Drosophila melanogaster, the condensin II protein, Cap-d3, was targeted by expressing dsRNAs in the developing brain. Ubiquitous knockdown of Cap-d3 in the fly is lethal in the first instar larval stage, and antibodies that reliably work to detect Cap-d3 protein in immunofluorescence analyses performed on Drosophila tissue are unavailable. Therefore, to confirm that the dsRNA was effective in knocking down cap-d3, qRT-PCR analyses of cap-d3 transcript levels were performed in first instar larvae expressing cap-d3 dsRNA under the control of armGAL4, which drives expression in a ubiquitous manner during this early developmental stage. Indeed, results showed significant decreases in cap-d3 transcripts in cap-d3 dsRNA expressing larvae, as compared to control, GFP dsRNA expressing larvae (Fig. 1a). To drive expression of cap-d3 dsRNA in the developing brain, a combined eyGAL4, GMRGAL4 driver obtained from the Johnston lab at Johns Hopkins University was used. Both the Eyeless and GMR proteins are expressed in the developing brain38,39,40,41,42. Results demonstrated that adult female flies expressing cap-d3 dsRNA under the control of eyGAL4, GMRGAL4 exhibited visibly smaller heads in comparison to control GFP dsRNA expressing flies (Fig. 1b). Female flies are shown, since males expressing cap-d3 dsRNA died at the mid to late pupal stages. As a proxy for measuring adult head size, the distance between two macrochaetes positioned between the eyes of adult flies was measured (pictured in Fig. 1c). Results demonstrated that Cap-d3 insufficient, female, adult flies reared at either 25 °C (Fig. 1d) or 18 °C (Fig. 1e) exhibited significantly decreased head sizes. Measurement of dissected adult brain volumes from flies 1–3 days post-eclosure, revealed that Cap-d3 insufficient flies also possessed significantly smaller adult brains, as compared to controls (Fig. 1f).
a qRT-PCR analyses of cap-d3 transcripts were performed on cDNA generated from first instar larvae expressing control UAS-GFP dsRNA (pink) or UAS-cap-d3 dsRNA (red) under the control of armGAL4. Transcripts were normalized to the housekeeping gene, rp49. The chart shown includes technical replicates and is representative of two independent experiments. P values were determined by performing a two-tailed, unpaired t-test. b Adult female fly heads were dissected from flies expressing control UAS-GFP dsRNA (top row) or UAS-cap-d3 dsRNA (bottom row) under the control of eyGAL4, GMRGAL4. c–e The distance between two machrochaetes positioned between the eyes of adult flies was measured as a proxy for measuring adult head size. The region measured is shown in (c). Adult, female fly head sizes were measured on flies expressing control UAS-GFP dsRNA (pink) or UAS-cap-d3 dsRNA (red) driven by eyGAL4, GMRGAL4 at 25 °C (d) or at 18 °C (e). Charts shown in (d) and (e) are representative of two independent experiments; each data point represents a single fly. f Adult brain volumes were measured in female flies (1–3 days post-eclosure) expressing control UAS-GFP dsRNA (pink) or UAS-cap-d3 dsRNA (red) driven by eyGAL4, GMRGAL4 at 25 °C. The charts shown are representative of two independent experiments; each data point represents a single brain. g Adult head sizes were measured in female flies expressing different combinations of control UAS-GFP dsRNA or UAS-cap-d3 dsRNA and UAS-GFP or UAS-GFP-cap-d3 under the control of eyGAL4, GMRGAL4. The results shown are the combined results of two independent experiments; each data point represents a single fly. For experiments in (d–g), P values were determined by performing two-tailed Mann–Whitney analyses. For a, p ≤ 0.0001. For d, p = 0.0001. For e, p = 0.0121. For f, p = 0.0010. For g, **p = 0.0060, ***p = 0.0001, ****p ≤ 0.0001. NS not significant. For all experiments, error bars indicate standard deviations from the mean.
To determine whether the knockdown of Cap-d3 was directly responsible for the observed decreases in head size and brain volume, genetic rescue experiments were conducted. A UAS-GFP-cap-d3 allele was expressed in the background of cap-d3 insufficiency to bring Cap-d3 expression levels closer to wild-type levels during development (Fig. 1g). Results demonstrated that female flies expressing cap-d3 dsRNA in combination with GFP-Cap-d3 (blue) exhibited head sizes that were not significantly different from control flies (black, pink, and turquoise). They were, however, significantly larger than the head sizes of flies expressing cap-d3 dsRNA alone (dark purple) or in combination with GFP protein (light purple).
Drosophila condensin I and condensin II both contain structural maintenance of chromosome proteins (Smc) Smc2 and Smc4, but differ in the chromosome-associated protein (Cap) subunits, with Cap-h, Cap-d2, and Cap-g comprising the Cap subunits of condensin I and Cap-h2 and Cap-d3 comprising the Cap subunits of condensin II (Fig. 2a). To determine whether decreasing the expression of other condensin subunits also results in microcephaly, head sizes of flies expressing dsRNAs targeting smc2, cap-d2, and cap-h were analyzed (Figs. 2b, c, e, g, i, k). The knockdown of the respective transcripts in first instar larvae were analyzed by qRT-PCR (Figs. 2d, f, h, j, l). Results confirmed that all dsRNAs efficiently targeted condensin subunits and demonstrated that decreasing each of the subunits also resulted in smaller female, pupal head sizes (knockdown was pupal lethal) (2c, g) or smaller female, adult head sizes (2e, i, k). Together, these results suggest that condensin I and condensin II proteins are necessary for proper brain development in Drosophila.
a Diagram of the proteins that comprise the condensin I and condensin II complexes in Drosophila melanogaster. b, c, g) Female pupae expressing (b) UAS-GFP dsRNA, (c) smc2 dsRNA, or (g) cap-d2 dsRNA under the control of eyGAL4, GMRGAL4 were dissected from pupal cases and imaged. e, i, k Adult, female fly head sizes were measured from flies expressing control (UAS-GFP or UAS-mCherry) dsRNA or dsRNA targeting (e) smc2, (i) cap-d2, and (k) cap-h under the control of eyGAL4, GMRGAL4. The charts shown are representative of two independent experiments; each data point represents a single fly. P values were determined by performing two-tailed Mann–Whitney analyses. For (e), p = 0.0010. For (i), p = 0.0376. For (k), p = 0.0067. d, f, h, j, l qRT-PCR analyses of d, f smc2, h, j cap-d2, and l cap-h transcript levels were performed on cDNA generated from first instar larvae expressing control (UAS-GFP or UAS-mCherry) dsRNA or dsRNAs targeting the respective condensin subunit mRNA under the control of armGAL4. Transcripts were normalized to the housekeeping gene, rp49. The charts shown include technical replicates of samples collected from four independent experiments (~200 larvae per experiment, combined into one sample per genotype). P values were determined by performing two-tailed, unpaired t-tests. For (d), p = 0.0021. For (f), p = 0.0421. For (h), p ≤ 0.0001. For all experiments, error bars indicate standard deviations from the mean.
Cap-d3 insufficiency leads to smaller pupal brain volumes
Given that smc2 and cap-d2 knockdown resulted in visibly smaller pupal heads, cap-d3 dsRNA expressing pupae were also examined. Comparisons between cap-d3 dsRNA expressing male pupae and controls (pupae were removed from their cases at 72–96 h post pupation) also revealed visibly smaller head sizes in Cap-d3 insufficient pupae (Fig. 3a). To determine whether larval brain volumes were also decreased by Cap-d3 insufficiency, the volumes of both female and male third instar larval brains expressing cap-d3 dsRNA were measured and compared to controls (Fig. 3b). Interestingly, no differences in brain volumes were observed for either sex, when compared to controls. These data suggest that microcephaly in condensin-insufficient brains first becomes apparent in the pupal stages, and the causative events most likely occur in the late larval stages.
a Larvae expressing control UAS-GFP dsRNA or UAS-cap-d3 dsRNA, under the control of eyGAL4, GMRGAL4 were allowed to pupate for 72–96 h, at which time males were dissected and imaged. b The volumes of brains dissected from third instar larvae expressing control UAS-GFP dsRNA or UAS-cap-d3 dsRNA, under the control of eyGAL4, GMRGAL4 were measured in males (left) and females (right). Results shown include larvae harvested from two independent experiments; each data point represents a single brain. P values were determined by performing two-tailed Mann–Whitney analyses. NS not significant. Error bars indicate standard deviations from the mean.
To identify the cell types in which eyGAL4, GMRGAL4 drives Cap-d3 insufficiency, subsequently leading to loss of brain volume and head size, immunofluorescence analyses were used to image the eyGAL4, GMRGAL4-driven expression pattern of GFP protein in third instar larval brains. Results demonstrated that the eyGAL4, GMRGAL4 drives expression in a large region of the third instar larval central brain as well as parts of the optic lobes (Fig. 4a). Since this region includes deadpan (Dpn) positive NSPCs, as well as post-mitotic neurons and glial cells, experiments were performed to test whether expression of cap-d3 dsRNA driven by different, cell type-specific GAL4 drivers could recapitulate the loss of adult brain volume and/or adult head size seen with eyGAL4, GMRGAL4. Cap-d3 dsRNA expression driven by repoGAL4, or by elavGAL4, which respectively drive expression in glial cells or in post-mitotic neurons, had no effect on adult head size (Supp. Fig. 1a–d). However, when two different GAL4 drivers, optixGAL4, and c253GAL4, were used to drive expression of cap-d3 dsRNA in regions of the central brain that contain Dpn+ NSPCs (Fig. 4b, c), adult male brain volumes were significantly decreased, as compared to controls expressing GFP dsRNA (Fig. 4d, e). Interestingly, optixGAL4 and c253GAL4-mediated Cap-d3 insufficiency did not affect adult female brain volumes (Supp. Fig. 2), again suggesting that microcephaly caused by Cap-d3 insufficiency in the developing brain, may be more severe in males. Additionally, while eyGAL4, GMRGAL4, and optixGAL4 all drive expression in developing eye disks, c253GAL4 does not43, suggesting that cap-d3 dsRNA expression in eye disks is not required to cause microcephaly. It should be noted, however, that impaired development of eye disks can affect head development44,45,46,47, and it is, therefore, possible that eyGAL4, GMRGAL4, and optixGAL4-driven expression of condensin dsRNAs in developing eye disks could contribute to the severity of the microcephaly observed. Finally, due to the fact that eyGAL, GMRGAL4, optixGAL4, and c253GAL4 expression patterns all include regions containing Dpn+ central brain NSPCs, as well as Dpn+ optic lobe NSPCs, a dpnGAL4 driver was used to drive expression of cap-d3 dsRNA primarily in optic lobe stem cells (Supp. Fig. 3a) and test whether Cap-d3 knockdown in these cells is sufficient to decrease brain volume. Results showed no significant differences in adult male brain volumes between flies expressing cap-d3 dsRNA and controls (Supp. Fig. 3b). Therefore, these combined data suggest that Cap-d3 insufficiency in larval central brain NSPCs may be sufficient to cause microcephaly.
a–c Third instar larval brains were dissected from male larvae expressing UAS-GFP under the control of a eyGAL4, GMRGAL4, b optixGAL4, or c c253GAL4, immunostained with antibodies to detect GFP (green) and stem cell marker, Dpn (red), and imaged using confocal microscopy. Maximum projections of z-stacks are shown. Nuclei are stained with DAPI (blue). All images were taken with 40x magnification. All images shown are representative of at least five biological replicates. Scale bar = 50 µm. d, e Adult brain volumes were measured in male flies expressing control UAS-GFP dsRNA or UAS-cap-d3 dsRNA driven by d optixGAL4 or e c253GAL4 at 25 °C; each data point represents a single brain. P values were determined by performing two-tailed Mann–Whitney analyses. For (d), p = 0.0156. For (e), p = 0.0278. Error bars indicate standard deviations from the mean. Results shown include adults harvested from two independent experiments.
Knockdown of Cap-d3 in the larval central brain increases NSPC death
Proliferation, differentiation, and programmed cell death are each required for the development of the brain in most organisms, including Drosophila, and must be tightly regulated to produce a normal, functioning adult organ (reviewed in refs. 1,48). To determine whether Cap-d3 insufficiency in third instar larval NSPCs affected any of these cellular processes, immunofluorescence analyses were first performed to detect markers of proliferation (phosphorylated Histone H3; PH3) and differentiation (Prospero; Pros). No differences in the number of Dpn+, PH3+ or Dpn+, Pros+ stem and progenitor cells were observed between cap-d3 dsRNA expressing brains and control brains (Fig. 5a, b). Total numbers of Dpn+ stem and progenitor cells present in the central brain also did not significantly change, although Cap-d3 insufficient brains trended toward a decrease (Fig. 5c). To assay for cell death, immunofluorescence analyses were performed to detect the effector caspase Death Caspase 1 (Dcp1) (Fig. 5d) in larvae expressing cap-d3 dsRNA or controls, GFP dsRNA or eyGMR (no RNAi expression). Results demonstrated significant increases in numbers of Dpn+, Dcp1+ stem and progenitor cells in brains expressing cap-d3 dsRNA (Fig. 5e). Furthermore, there was also an increase of Dcp1+, Dpn− cells positioned directly adjacent to the Dpn+ stem and progenitor cells were also observed in brains expressing cap-d3 dsRNA (Fig. 5f). Larval central brain neural stem cells (Type I and Type II neuroblasts) and the ganglionic mother cells produced through asymmetric division of Type I neuroblasts are Dpn+. Not all of the cells produced through asymmetric division of neuroblasts are Dpn+, however, since Type II neuroblasts generate Dpn− intermediate progenitor cells (INPs) that only re-express Dpn following a maturation period49,50. Mature INPs then divide asymmetrically to self-renew and produce additional Dpn− cells. Therefore, the Dpn- cells neighboring the Dpn+ cells could either be direct descendants of neuroblasts, or they could be non-progenitor cells. Together, these data suggest that Cap-d3 insufficiency in the third instar larval central brain increases the number of dying NSPCs and the surrounding neighbors. Finally, to determine whether the increases in cell death were responsible for the reduced head sizes in flies expressing cap-d3 dsRNA, the Drosophila inhibitor of apoptosis 1 (Diap1) was overexpressed in combination with cap-d3 dsRNA. Diap1 overexpression caused adult female head sizes of cap-d3 dsRNA expressing flies to resemble the sizes of control flies expressing GFP dsRNA and Diap1 (Fig. 5g, blue compared to purple), thus rescuing the significantly smaller head sizes observed in flies expressing cap-d3 dsRNA alone (Fig. 5g, blue compared to red).
a Third instar larval brains were dissected from male larvae expressing control UAS-GFP dsRNA or UAS-cap-d3 dsRNA, under the control of eyGAL4, GMRGAL4, immunostained with antibodies to detect stem cell marker, Dpn, and mitotic cell marker, PH3, and imaged using confocal microscopy. The percentage of cells that stained positive for both Dpn and PH3 were quantified. b Immunostaining for Dpn and Pros, a marker of differentiation, was performed in male third instar larval brains. The number of cells that stained positive for both Dpn and Pros were quantified. c The number of Dpn-positive cells per square micrometer of larval central brain tissue in larvae expressing cap-d3 dsRNA or control, gfp dsRNA was quantified. For a–c, results shown include larvae harvested from two independent experiments; each data point represents a single brain. P values were determined by performing two-tailed Mann–Whitney analyses. NS not significant. d Immunostaining to detect Dpn and the cell death marker, Dcp1 was performed in male third instar larval brains expressing control UAS-GFP dsRNA (dark blue) or UAS-cap-d3 dsRNA (light blue), under the control of eyGAL4, GMRGAL4. Images were taken with a confocal microscope. Maximum projections of z-stacks are shown. Nuclei are stained with DAPI (blue). All images were taken with 40x magnification. Scale bar = 50 µm. e The percentage of cells that stained positive for both Dpn and Dcp1 in (d) were quantified. Additionally, larval brains expressing eyGAL4 and GMRGAL4 (gray; no RNAi control) were imaged and quantified. Each data point represents a single brain. f Dcp1+, Dpn− cells located adjacent to Dpn-positive cells in male third instar larval brains (experiments described in d, e) were quantified. Each data point represents a single brain. g Adult female head sizes were measured in female flies expressing different combinations of control UAS-GFP dsRNA or UAS-cap-d3 dsRNA and the Drosophila inhibitor of apoptosis, UAS-diap1, under the control of eyGAL4, GMRGAL4. For e–g, results shown include larvae harvested from two independent experiments; each data point represents a single brain (e, f) or a single fly (g). P values were determined by performing two-tailed Mann–Whitney analyses. For (e), *p = 0.0212 and *#p = 0.0411. For (f), *p = 0.0013 and *#p = 0.0024. For (g), *p = 0.0422, ***p = 0.0004, ****p ≤ 0.0001, ****#p ≤ 0.0001, NS not significant. For all experiments, error bars indicate standard deviations from the mean.
Cap-d3 insufficiency increases retrotransposition in larval NSPCs
We previously published that condensin proteins repress RTE expression and mobility in Drosophila cells and tissues26. To determine whether Cap-d3 knockdown in third instar larval brains causes de-repression of RTE transcripts, qRT-PCR was performed to detect transcript levels of three separate RTE families, mdg1, mdg4 (formerly known as gypsy), and X-element in third instar larval brains of larvae that were transheterozygous for a hypomorphic mutant cap-d3 allele and a chromosomal deficiency spanning the region encompassing the cap-d3 locus (cap-d3c07081/Df(2L)Exel7023). Results demonstrated significant increases in transcript levels of all three RTE families in mutants, as compared to w1118 controls (Fig. 6a). Since RTE expression is the first step in the process of retrotransposition, we used the gypsyCLEVR reporter system51 to test whether mdg4 retrotransposition, in addition to expression, was also increased in cap-d3 dsRNA expressing larval brains. This reporter causes cells to express a membrane-localized GFP protein and a nuclear mCherry protein under the control of a UAS, immediately following retrotransposition of a mdg4 RTE which is under the control of its own promoter51. To detect cells in which mdg4 retrotransposition events had occurred, immunofluorescence analyses were performed on third instar larval brains with anti-mCherry antibody (pseudocolored green). Brains were also immunostained with antibody to Dpn (pseudocolored red). Unexpectedly, many mCherry-positive cells were observed in the central brains of control, GFP dsRNA expressing larval brains, and the majority of these cells either stained positive for Dpn or were adjacent to Dpn+ cells (Fig. 6b). Furthermore, cap-d3 dsRNA expressing brains exhibited significant increases in the number of cells in which mdg4 retrotransposition events occurred (Fig. 6b, c), and the majority of these cells were also either Dpn+ or a neighbor of a Dpn+ cell. These increases were not a result of general RNAi pathway activation, since larval brains expressing the gypsyCLEVR reporter and GAL4 driver, without dsRNA, exhibited similar numbers of Dpn+, mCherry+ cells, as compared to larval brains expressing GFP dsRNA. As expected, no mCherry-positive cells were observed in larval brains expressing a gypsyCLEVR mutant51 which was used as a negative control (Supp. Fig. 4). These data suggest that RTE expression and mobility is increased in Cap-d3 insufficient NSPCs, as well as in the neighboring cells. Interestingly, the increased percentages of cells exhibiting retrotransposition events in Cap-d3 insufficient larvae were returned to control levels when larvae were allowed to develop on food containing the Nucleoside Reverse Transcriptase Inhibitor (NRTI), Azidothymidine (AZT) (Fig. 6d). Results also revealed that AZT does not completely abrogate retrotransposition events in developing control or cap-d3 insufficient brains. The most likely explanation for these results is that the increased retrotransposition events observed in Cap-d3 insufficient brains occur after the development of the first instar larval mouth hooks. Once the mouth hooks develop and larvae ingest AZT, this inhibits future retrotransposition events. Remaining retrotransposition events, detected by the gyspyCLEVR reporter in both Cap-d3 insufficient and control brains, would have occurred prior to this developmental stage, thus causing them to be unaffected by the addition of NRTIs to the food.
a qRT-PCR analyses of RTE (mdg1, mdg4, and X-element) transcripts were performed with cDNA generated from third instar larval brains dissected from w1118 controls or cap-d3 transheterozygous mutant larvae (150 brains per genotype; mixed sexes). Transcripts were normalized to the housekeeping gene, rp49. Results show the averages of three independent biological replicates. P values were determined by performing two-tailed, unpaired t-tests. For mdg1, **p = 0.0054, for mdg4, **p = 0.0015, and for x-element, **p = 0.0067. b Third instar larval brains were dissected from male larvae expressing UAS-gypsyCLEVR51 and UAS-GFP dsRNA or UAS-cap-d3 dsRNA under the control of eyGAL4, GMRGAL4, immunostained with antibodies to mCherry, a reporter of gypsy retrotransposition (pseudocolored green), and stem cell marker, Dpn (red), and imaged using confocal microscopy. Maximum projections of z-stacks are shown. Nuclei are stained with DAPI (blue). All images were taken with 40x magnification. Scale bar = 50 µm. c The percentage of cells staining positive for both mCherry and Dpn were quantified from experiments described in (b). Male larval brains expressing eyGAL4 and GMRGAL4-driven UAS-gypsyCLEVR were also analyzed as “no RNAi” controls. Results shown include larvae harvested from two independent experiments; each data point represents a single brain. d Third instar larval brains were dissected from male larvae expressing UAS-gypsyCLEVR and control UAS-GFP dsRNA (green circles) or UAS-cap-d3 dsRNA (orange squares), under the control of eyGAL4, GMRGAL4, immunostained with antibodies to detect mCherry and Dpn, and imaged using confocal microscopy. The percentage of cells that stained positive for both Dpn and Dcp1 were quantified. Flies were developed on food containing DMSO as a control (closed circles and squares), or on food containing 5 µM AZT (open circles and squares). Results shown include larvae harvested from two independent experiments; each data point represents a single brain. For experiments in (c, d), P values were determined by performing two-tailed Mann–Whitney analyses. For (c), **p = 0.0029, ****p ≤ 0.0001, NS not significant. For (d), **p = 0.0037, **#p = 0.0039, NS not significant. Error bars indicate standard deviations from the mean.
NRTIs rescue microcephaly induced by Cap-D3 insufficiency
RTE expression and mobilization can lead to cell death in human cell lines52, and in a cortical organoid-derived model of the early onset inflammatory disorder, Aicardi-Goutières syndrome53. De-repression/overexpression of retrotransposons in adult neurons of Drosophila models of neurodegenerative disease can also lead to cell death54,55. To determine whether inhibition of RTE activity could prevent the increased NSPC death in cap-d3 dsRNA-expressing larval brains, male larvae were raised in food containing 5 µM AZT or DMSO. Cap-d3 dsRNA expressing larval brains dissected from larvae developed on AZT, exhibited similar numbers of Dpn+, Dcp1+ NSPCs as compared to GFP dsRNA expressing larval brains dissected from larvae developed on AZT or DMSO (Fig. 7a). This was also true for Dpn− cells neighboring the Dpn+ cells (Fig. 7b). Additionally, cap-d3 transheterozygous mutants developed on DMSO also exhibited increased percentages of Dcp1+ NSPCs, and these percentages were decreased to levels resembling wild-type controls when larvae were developed on AZT (Fig. 7c).
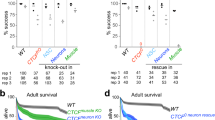
a Third instar larval brains were dissected from female larvae expressing control UAS-GFP dsRNA (green circles) or UAS-cap-d3 dsRNA (orange squares), under the control of eyGAL4, GMRGAL4, immunostained with antibodies to detect stem cell marker, Dpn, and cell death marker, Dcp1, and imaged using confocal microscopy. The percentage of cells that stained positive for both Dpn and Dcp1 were quantified. Larvae were developed on food containing DMSO as a control (closed circles and squares), or on food containing 5 µM AZT (open circles and squares). b Dcp1+, Dpn− cells located adjacent to Dpn-positive cells in female third instar larval brains (experiments described in (a)) were quantified. Results shown in (a, b) include larvae harvested from two independent experiments; each data point represents a single brain. c Experiments described in (a) were performed on brains dissected from cap-d3 transheterozygous mutant (blue squares) male larvae or wild-type (green circles) male larvae. Results shown include larvae harvested from two independent experiments; each data point represents a single brain. d, e Adult female fly head sizes were measured on flies expressing control UAS-GFP dsRNA (green circles) or UAS-cap-d3 dsRNA (orange squares) driven by eyGAL4, GMRGAL4 at 25 °C. Flies were developed on food containing DMSO as a control (closed circles and squares), or on food containing (d) 5 µM AZT or (e) 5 µM d4T (open circles and squares). f Adult male head sizes were measured on wild-type flies (green circles), on cap-d3 transheterozygotes (sky blue squares) or cap-d3 heterozygotes (light blue and dark blue squares). Flies were developed on food containing DMSO as a control (closed circles and squares), or on food containing 5 µM AZT (open circles and squares). Results shown in (d–f) include adults harvested from two independent experiments; each data point represents a single fly. P values were determined by performing two-tailed Mann–Whitney analyses. For (a), ****p ≤ 0.0001 and ****# p ≤ 0.0001. For (b), *p = 0.0434, **p = 0.0079, NS not significant. For (c), **p = 0.0013, ***p = 0.0007, NS not significant. For (d), *p = 0.0208, *#p = 0.0336, **p = 0.0014. For (e), *p = 0.038, **p = 0.0015, **#p = 0.0084, NS not significant. For (f), *p = 0.0181, **p = 0.0018, ****p ≤ 0.0001, ****#p ≤ 0.0001, NS not significant. For all experiments, error bars indicate standard deviations from the mean.
The involvement of RTE expression and activity in causing reduced head sizes in flies expressing cap-d3 dsRNA under the control of eyGAL4, GMRGAL4, was also analyzed (Fig. 7d). Results demonstrated that raising cap-d3 dsRNA expressing flies on food containing AZT prevented the significant decreases in adult head size observed in cap-d3 dsRNA expressing flies raised on DMSO, causing head size measurements to more closely resemble control GFP dsRNA expressing flies developed on DMSO or AZT (Fig. 7d). This was also true for flies developed on stavudine (d4T), another NRTI (Fig. 7e), and for cap-d3 transheterozygous or heterozygous mutants developed on AZT, when compared to wild-type (w1118) controls (Fig. 7f). Together, these data suggest that unrestricted RTE expression may be responsible for the increased numbers of dying cells in Cap-d3 insufficient third instar larval central brains, and NRTI-mediated inhibition of RTEs can prevent both the increased cell death, as well as the resulting decreases in brain volumes and head size.
To determine whether the general upregulation of RTEs could result in similar phenotypes, ago2 mutants were analyzed for levels of NSPC death in the larval central brain and for the development of microcephaly in the absence or presence of NRTIs. AGO2 is required in somatic tissues to repress RTE expression56,57,58,59. The ago2454 allele was shown to carry a deletion of ~6 kb that removes the PAZ and PIWI domains of AGO2, resulting in a null allele60. The ago2Df(3L)BSC558 allele is a deficiency allele harboring a much larger chromosomal deletion which has been mapped but is uncharacterized in relation to ago2 expression. To confirm that flies expressing the ago2Df(3L)BSC558 exhibit decreased ago2 transcript levels, qRT-PCR was performed on RNA harvested from dissected intestines of ago2Df(3L)BSC558 adults that were reared on food containing DMSO or AZT. Results demonstrated reduced ago2 transcript levels, in comparison to wild-type, w1118 controls (Supp. Fig. 5a), regardless of which food they were developed on. Interestingly, ago2 transheterozygous larval brains exhibited increased percentages of dying NSPCs, and this was rescued by allowing larvae to develop on food containing AZT (Supp. Fig. 5b). Furthermore, ago2 mutants also possessed smaller head sizes, as compared to wild-type flies, and this could also be rescued by ingestion of AZT (Supp. Fig. 5c). Combined, these results suggest that general upregulation of RTE expression and activity, independent of genetic background, may impair brain development and cause microcephaly in Drosophila.
Discussion
Collectively, our results suggest: (1) Tissue-specific knockdown of condensin I and condensin II proteins in the central brain results in significant decreases in adult brain volume and adult head sizes, thus modeling microcephaly observed in patients with condensin loss of function mutations; (2) RTE expression and activity are upregulated in NSPCs of third instar larval central brains expressing decreased levels of condensin proteins, leading to increased numbers of dying cells; (3) The ingestion of NRTIs during development inhibits RTE activity, allowing larvae to develop into adults with normal head sizes. These findings suggest a working model where condensin I and condensin II restrict RTE expression and mobilization in NSPCs, thus preventing an overabundance of cell death in the third instar larval brain which would lead to decreased brain volumes, starting in the pupal stage, and ultimately cause reduced adult head sizes (Fig. 8).
Condensin I and II restrict RTE expression and retrotransposition in neuroblasts and intermediate neural precursor cells in the developing larval brain. This, in turn, restricts cell death of stem and precursor cells, as well as cells adjacent to stem/precursor cells, thereby allowing the brain to achieve proper volume by the late pupal stages.
The tissue-specific knockdown of Cap-d3 and other condensin subunits in the developing Drosophila brain, driven by eyGAL4 and GMRGAL4, results in significant decreases in adult brain volumes and head sizes (Figs. 1, 2). Similarly, cap-d3 mutants also exhibit decreased brain volumes and head sizes (Fig. 7), in addition to decreases in overall body weight61. This suggests that decreasing condensin subunit expression can have the universal effect of decreasing organ size in Drosophila, regardless of the cell type. cap-h2 mutant mice also exhibit overall decreases in body weight, in addition to microcephaly12. Interestingly, germline condensin mutations identified in human microcephaly patients caused some, but not all, of the patients to exhibit significantly decreased stature, or even dwarfism12. It is perplexing that some germline mutations that cause decreased expression of condensin proteins, which function in every cell type, cause reductions in brain volume in some patients without affecting overall body size. LINE-1 RTEs can mobilize within mammalian neural stem cells31,62,63, and LINE-1 and Alu insertions have been identified in adult human neurons and glia64. However, these insertions were not observed in fibroblasts from the same organism/individual31,64. Additionally, whole-genome sequencing of brain tissues revealed that the majority of somatic LINE-1 retrotransposition events resulted in the integration of the LINE-1 sequence into a preexisting L1 sequence65. Therefore, we hypothesize that RTE expression and activity may be higher in developing human brain tissues than in other somatic tissues within the body, and thus, the depletion of condensin protein increases RTE expression and activity to very high levels that cannot be tolerated in the brain. This may not happen in other organs where baseline RTE expression/activity levels are much lower, such that condensin insufficiency still results in higher RTE expression/activity, but it remains below a threshold that can still be tolerated by the cells. It is also possible that in some organisms, the mechanisms that regulate condensin expression/ activity may become dysregulated, specifically in the brain. The identity of the major mechanisms regulating condensin expression still remains largely unknown, and future studies aimed at elucidating these mechanisms in the brain will be important for understanding the tissue-specific vs. whole-body effects of condensin insufficiency.
Cap-d3 knockdown results in decreased pupal brain volumes, which in males, causes pupal heads to almost completely disappear and results in lethality (Figs. 2, 3). Females, however, do eclose, and exhibit reduced brain volumes and head sizes, but their heads are more fully formed than their male counterparts (Fig. 1b vs 3a). Why do condensin-insufficient males exhibit more severe phenotypes? Different RTE families are expressed at different levels in males and females66,67,68,69. It is possible that the upregulation of a few specific RTE families in condensin-insufficient Drosophila brains could be responsible for the observed phenotypes. Therefore, if those families naturally display higher expression levels in males, then further upregulation could have more severe effects. It will be interesting to determine whether condensin insufficiency causes a global upregulation of all RTEs or specific RTE families in both males and females, and whether the upregulation is to an equal extent in both sexes.
Our results demonstrated that mdg4 retrotransposition was permissible in control brains expressing GFP dsRNA or no RNAi (Fig. 6b, c). These mdg4 retrotransposition events were primarily observed in Dpn+ cells or Dpn− cells positioned within clusters surrounding the Dpn+ cells, which are, most likely, precursor cells (INPs) that directly descended from the Dpn+ neuroblast49,50,70. The fact that eyGAL4 and GMRGAL4 drives in a large area of the larval central brain (Fig. 4a), but retrotransposition events were only observed in NSPCs, suggests that these cell types may be the only cell types that permit RTE mobilization in the developing brain. It is also possible that eyGAL4 and GMRGAL4 drive expression at higher levels in NSPCs than in other retrotransposition-permissible cells in the central brain, potentially making it difficult to observe the reporter identifying retrotransposition events in these other cell types. Interestingly, NRTI ingestion did not prevent retrotransposition in developing control brains, although it did prevent the increased retrotransposition in cap-d3 insufficient brains (Fig. 6d). We hypothesized that this was because the retrotransposition events that occurred in the control brains happened prior to the development of first instar larval mouth hooks. If this is true, then RTEs may play essential roles during early Drosophila brain development. RTEs can modulate mammalian gene expression through multiple mechanisms, including the introduction of transcription factor binding sites, regulation of 3D genome architecture, production of ncRNAs, and generation of a silenced chromatin environment (reviewed in ref. 71). In fact, lineage-specific enhancers that are enriched in insertions of a specific family of SINEs play important roles in mammalian neural development72,73. Recently, an elegant study determined that L1s are expressed in apical and basal progenitor cells and early-born neurons present in developing human fetal forebrain; silencing of a specific L1 insertion in cerebral organoids developed from these cells resulted in smaller size74. Therefore, it is possible that RTEs also regulate gene expression programs that control early brain development in Drosophila, and future experiments involving scRNA-seq of wild type, condensin mutant, and ago2 mutants may shed light on these questions.
Interestingly, mdg4 retrotransposition events detected using the gypsyCLEVR reporter in the adult Drosophila brain, were observed at a much lower frequency than what we observed in the larval brain51. This suggests that cells which have undergone retrotransposition events during development must largely disappear during metamorphosis. Similar observations were recently reported in the Drosophila hindgut75. Our data show that unrestricted RTE expression and activity in condensin-insufficient or ago2 deficient NSPCs results in increased cell death (Fig. 7 and Supp. Fig. 5). Combined with the possibility that cells harboring mdg4 retrotransposition events seem to disappear during metamorphosis, we hypothesize that RTE expression and activity may contribute to the balance of proliferation and cell death that is necessary for the proper development of the brain. Programmed cell death is, in fact, necessary to eliminate post-mitotic neurons of specific neuroblast lineages which are generated during larval development and eliminated in early pupal stages76, and it will be interesting to determine whether RTEs may play a role in these processes. Finally, RTE insertions in mammalian neural precursor cells occur frequently in genes associated with neuron-specific functions and can regulate the expression of those genes to drive neuronal somatic mosaicism63,77. Recent evidence demonstrated that the knockdown of condensin I subunit Cap-g in post-mitotic neurons affected the expression of both neuronal and non-neuronal genes78. It is, therefore, conceivable that retrotransposition of RTEs could promote somatic mosaicism in larval central brain NSPCs. However, if the majority of larval brain cells exhibiting retrotransposition events die during metamorphosis (as discussed above), then the effects of mosaic gene expression in larval neurons generated from the NSPCs would most likely be very transient.
While our findings suggest that increased RTE expression and activity is a potential mechanism driving condensin-insufficient NSPC death, the molecular events that activate apoptotic pathways downstream of RTE expression/activity are unresolved. Condensin protein insufficiency leads to increased levels of DNA damage in Drosophila cells, specifically within loci containing RTEs26. Decreased condensin expression in Drosophila larvae also results in increased transcription of antimicrobial peptides, which are major effectors of innate immune responses in Drosophila79. Both DNA damage, and increased immune signaling can be caused by the failure to repair DNA breaks at the site of RTE insertions and this can activate cell death pathways in mammalian cells52,80,81,82,83,84,85,86,87. Cytoplasmic DNAs generated by increased RTE expression and activity can also activate cGAS-STING and IFN signaling in mammalian cells82,83, which could lead to cell death. While Drosophila possesses cGAS-STING homologs88,89, their functions in innate immune signaling are still being studied, and we currently do not know whether they are required for the increased cell death observed in condensin-insufficient or ago2 deficient larval brains. Likewise, it will be interesting to determine in future studies whether RTE insertion or simply the expression of RTEs are required for the loss of brain volumes and smaller adult head sizes observed in these microcephaly models.
Finally, our studies invoke the question of whether the upregulation of RTEs is a common mechanism that could lead to microcephaly development in other genetic backgrounds. Interestingly, other microcephaly proteins (proteins encoded by genes identified to be mutated in microcephaly patients) have been associated with RTE expression or repression of virus90,91,92,93. Additionally, the microcephaly protein MCPH1 has been shown to physically and genetically interact with condensin proteins94,95,96,97. Thus, a more extensive investigation of potential roles for all known microcephaly proteins in repressing RTEs in the brain will be required to fully elucidate the generality of RTE-mediated mechanisms of microcephaly.
Methods
Fly maintenance
All Drosophila melanogaster stocks and crosses were maintained at 25 °C on standard dextrose medium, unless otherwise noted. To prepare food containing Nucleoside Reverse Transcriptase Inhibitors (NRTIs), 500 mL of standard dextrose medium was poured into four 1 L flasks. After allowing the media to cool below 60 °C, 50 µl of 50 mM AZT (Zidovudine)(Selleckchem), or d4T (Stavudine) (Selleckchem), dissolved in dimethyl sulfoxide (DMSO) were dispensed into flasks and mixed thoroughly, so that the final concentration of each drug in the food was 5 µM. About 50 µl of DMSO alone were also dispensed into flasks to prepare control media. Media was divided into 6 oz bottles and allowed to cool. NRTI-containing media was stored at 18 °C for no longer than 2 weeks before use.
Fly stocks
The following wild-type, mutant, and deficiency stocks were used in these studies
w[1118] (Bloomington Stock Center 6326), w[1118]; dCap-D3c07081/c07081 (Harvard Exelixis collection generated in the w[1118] line98, w[1118]; Df(2 L)Exel7023/CyO (Bloomington Stock Center 7797), w[1118]; Df(3 L)BSC558/TM6C, Sb[1] (Bloomington Stock Center 25120), w[1118]; AGO2[454]/TM3, Sb[1] Ser[1] (Bloomington Stock Center 36512)
The following RNAi stocks were used in these studies
UAS-GFP dsRNA (P{UAS-GFP.dsRNA.R}142; Bloomington Stock Center 9330), UAS-cap-d3 dsRNA (w1118; P{GD913}; VDRC stocks 29657 and 9402), UAS-smc2-dsRNA (P{KK100466}VIE-260B; VDRC stock 103406 and P{TRiP.GL00440}attP40; Bloomington Stock Center 35602), UAS-cap-d2 dsRNA (P{KK101115}VIE-260B; VDRC stock 108289 and w1118; P{GD9677}v33424; VDRC stock 33424), UAS-cap-h dsRNA (y1 sc* v1 sev21; P{TRiP.HMS00049}attP2; Bloomington Stock Center 34068).
The following GAL4 and UAS stocks were used in these studies
armadillo GAL4 (w[*]; P{w[+mW.hs]=GAL4-arm.S}4a P{w[+mW.hs]=GAL4-arm.S}4b; Bloomington Stock Center 1561 that has lost the balancer), eyGAL4, GMRGAL4 (generous gift from Dr. Bob Johnston at Johns Hopkins University), UAS-GFP.nls (P{UAS-GFP.nls}14; Bloomington Stock 4775), UAS-GFP-CAP-D361, tubPGAL4 (Bloomington Stock 5138), repoGAL4 (generous gift from Dr. Heather Broihier at Case Western Reserve University), elavGAL4 (Bloomington Stock Center 8765), c253GAL4 (w[1118]; P{w[+mW.hs]=GawB}C253; Bloomington Stock Center 6980), dpnGAL4 (Bloomington Stock Center 47456), optixGAL4 (w[1118]; P{y[+t7.7] w[+mC]=GMR30D11-GAL4}attP2; Bloomington Stock Center 48098), UAS-gypsyCLEVR and UAS-gypsy-CLEVRΔPBS51.
RNA analysis by quantitative reverse transcriptase PCR (qRT-PCR)
Tissues were dissected, submerged in Trizol (Life Technologies) in a microcentrifuge tube, and homogenized using a Pellet pestle cordless motor (Kimble) and disposable Kontes pestle. For the first instar larvae collection, crosses (30 virgins and 15 males) were set up in bottles, and females were allowed to lay eggs onto grape agar plates, overnight. Grape agar plates were stored in the 25 °C incubator for 24 h, after which time, a paintbrush was used to collect larvae into a 100 µm nylon mesh cell strainer (Fisher). Larvae in cell strainers were submerged in 1X PBS, twice, and then submerged in 7% bleach in distilled water for 3 min, at room temperature. Larvae in cell strainers were then submerged in fresh 1X PBS for 30 s, and this was repeated four times. Larvae were transferred to microcentrifuge tubes, the remaining PBS was removed, and 50 µl Trizol was added to the tube. RNA was extracted during the standard Trizol protocol, according to the manufacturer.
Extracted RNAs were then treated with RNase-free DNase in buffer RDD (Qiagen) prior to further purification using an RNeasy Mini Kit (Qiagen). An aliquot of cDNA was generated from 0.5 to 2 µg of total RNA using the TaqMan Reverse Transcription Reagents (Applied Biosystems) and an oligo-dT(16) primer (Invitrogen). Quantitative RT-PCR was performed using the Roche Lightcycler 480 to amplify 15 µL reactions containing .5 µL of cDNA, 0.5 µL of a 10 µM primer mix, and 7.5 µL of Fast Start SYBR Green Master Mix (Roche). Each reaction was performed in triplicate. Crossing point (Cp/ Ct) values were determined using the Roche LightCycler 480 Absolute Quantification Second Derivative Analysis software. Relative quantitation of transcript levels was then performed using the delta-delta Ct method (2−ΔΔCt), where the Ct values of a reference gene (rp49) in each sample are subtracted from the Ct values of the gene of interest to create a ΔCt value for each sample. The ΔCt is compared to a control sample to generate a ΔΔCt value for each sample. Following the calculation of 2-ΔΔCt for each sample, triplicates were averaged. The sequences of oligos used in the qRT-PCR studies are listed below:
dCap-D3 FW: 5′-GGCGAATCATCAGCACCCTGC-3′
dCap-D3 RV: 5′-CAGGCATCGGTAGCCATGGAC-3′
dSMC2 FW: 5′-CGCGTAAGGTGCGTGGTTTG-3′
dSMC2 RV: 5′-GGGAATCAAGTGACACGACGCTG-3′
dCap-D2 FW: 5′-GGGAATCAAGTGACACGACGCTG-3′
dCap-D2 RV: 5′-CCATGCGAAGCGGCAAAAGTTCG-3′
dCap-H FW: 5′-GGCCTCACCCAGATGAACGCC-3′
dCap-H RV: 5′-TCGTCTCCTCCGGCACAGCG-3′
rp49 FW: 5′-TACAGGCCCAAGATCGTGAAG-3′
rp49 RV: 5′-GACGCACTCTGTTGTCGATACC-3′
ago2 FW: 5′-CCCCAACTCCATTGTCTGAACGTTG-3′
ago2 RV: 5′-CTTGCGCTTTCGCACGTTCGTC-3′
mdg4 FW: 5′-GTTCATACCCTTGGTAGTAGC-3′
mdg4 RV: 5′-CAACTTACGCATATGTGAGT-3′
mdg1 FW: 5′-AACAGAAACGCCAGCAACAGC-3′
mdg1 RV: 5′-TTTCTGATCTTGGCAGTGGA-3′
X-element FW: 5′-GCCAGCCTGCAACAGGTTGAAG-3′
X-element RV: 5′-CTCTGGCGCACAATGACTTCGG-3′
Drosophila adult head measurements
Adult Drosophila were anesthetized with CO2 and a razor blade was used to separate heads from bodies. Heads were stored at 4 °C until ready for imaging, at which point they were transferred to slides covered with double-sided sticky tape. A Leica MZ 16FA stereoscope with Leica imaging software was used to obtain images. Adobe Photoshop was used to measure the distance between two machrochaetes positioned between the eyes, depicted in Fig. 1c. This pair of machrochaetes was used in measurements for all figures. Pixels were converted to µm, based on the magnification used.
Drosophila adult and larval brain measurements
To dissect adult brains from flies that were between 1 and 3 days post-eclosure, Drosophila heads were separated as described above and placed in cold PBS in a well of a nine-well, glass, dissection plate (Corning). Forceps were used to peel the cuticle and eye tissue away from the brain. Brains were fixed in 4% Paraformaldehyde/ TritonX-100 solution (4% paraformaldehyde made from 16% paraformaldehyde aqueous solution, EM grade (Electron Microscopy Sciences), 0.25 M EGTA, pH 8.0, 1X PBS, 0.5% TritonX-100) for 25 min at room temperature. Brains were then incubated in Vectashield with DAPI (Vector Laboratories) overnight, and placed onto slides containing tiny shards of crushed coverslips and vacuum grease to prevent squashing of brain tissue when coverslips were placed over the top of the slides. Brains were imaged with a Leica SP5 confocal/multi-photon microscope. Z stacks were processed using Leica Imaging software. Volocity imaging software (Quorum Technologies) was used to measure the volumes of individual brains; 3D images were analyzed using the Volocity “measurements” and “find objects” features, with the threshold adapted to use “intensity”, and a minimum object size of 1000,000 µm3. The population of Interest was selected for measurement.
To dissect larval brains, third instar larvae were placed into wells of a nine-well, glass, dissection plate (Corning). Larval brains containing the central brain, optic lobes, and ventral nerve cords were removed from the larvae using forceps. Fixation, slide preparation, and imaging were performed as described above for adult brains.
Immunostaining of Drosophila larval brains
All tissues were fixed in 4% Paraformaldehyde/ TritonX-100 solution (described above) for 25 min, with rocking, at room temperature. Tissues were washed three times with 0.1% PBS-Triton and then incubated in blocking buffer (0.1% PBS-Triton/1%BSA) for 1 h at room temperature, with rocking. Tissues were then incubated with primary antibodies in a blocking buffer, overnight at 4 °C, with rocking. The following day, tissues were washed three times with 1X PBS at room temperature, with rocking. Tissues were then incubated with secondary antibodies for 1–2 h at room temperature, with rocking. Tissues were mounted in Vectashield with DAPI (Vector Laboratories), prior to imaging. Brains were imaged with a Leica SP5 confocal/multi-photon microscope. Z stacks were processed using Leica Imaging software. Cells positive for DPN, PH3, DCP1, and/or Pros were counted by hand. The total surface area of larval central brain tissues was measured using Image J.
Antibodies
Primary antibodies used included anti-deadpan (Abcam ab195174; 1:100), anti-GFP (Invitrogen A10262; 1:200), anti-DCP1 (Cell Signaling 9578; 1:200), anti-phospho-histone H3 ser10 (Cell Signaling 9701; 1:500), anti-prospero (Developmental Studies Hybridoma Bank MR1A; 1:1000, and anti-mCherry (Cell signaling 43590; 1:200). All secondary antibodies were used at a dilution of 1:500 and included Goat anti-Chicken IgY (H + L) Secondary Antibody, Alexa Fluor™ 488 (Thermo Fisher A-11039), Goat anti-Rabbit IgG (H + L) Cross-Adsorbed Secondary Antibody, Alexa Fluor™ 568 (Thermo Fisher A-11011), Donkey anti-Rabbit IgG (H + L) Highly Cross-Adsorbed Secondary Antibody, Alexa Fluor™ 647 (Thermo Fisher A-31573), Donkey anti-Rabbit IgG (H + L) Highly Cross-Adsorbed Secondary Antibody, Alexa Fluor™ 488 (Thermo Fisher A-21206), Goat anti-Mouse IgG (H + L) Cross-Adsorbed Secondary Antibody, Alexa Fluor™ 488 (Thermo Fisher A-11001), Goat anti-Mouse IgG (H + L) Cross-Adsorbed Secondary Antibody, Alexa Fluor™ 568 (Thermo Fisher A-11004), and Goat anti-Mouse IgG (H + L) Cross-Adsorbed Secondary Antibody, Alexa Fluor™ 647 (Thermo Fisher A-21235).
Statistics and data analysis
All Data analysis and statistical calculations were performed using GraphPad Prism software, version 10.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Further information and requests for resources and reagents should be directed to and will be fulfilled by the corresponding author, Michelle Longworth (longwom@ccf.org). Source data are provided with this paper.
References
Homem, C. C., Repic, M. & Knoblich, J. A. Proliferation control in neural stem and progenitor cells. Nat. Rev. Neurosci. 16, 647–659 (2015).
Homem, C. C. & Knoblich, J. A. Drosophila neuroblasts: a model for stem cell biology. Development 139, 4297–4310 (2012).
Hobbs, K. et al. A retrospective fetal ultrasound study of brain size in autism. Biol. Psychiatry 62, 1048–1055 (2007).
Sacco, R., Gabriele, S. & Persico, A. M. Head circumference and brain size in autism spectrum disorder: a systematic review and meta-analysis. Psychiatry Res. 234, 239–251 (2015).
Gilmore, E. C. & Walsh, C. A. Genetic causes of microcephaly and lessons for neuronal development. Wiley Interdiscip. Rev. Dev. Biol. 2, 461–478 (2013).
Carter, J. C. et al. Selective cerebral volume reduction in Rett syndrome: a multiple-approach MR imaging study. Am. J. Neuroradiol. 29, 436–441 (2008).
Naidu, S. et al. Neuroimaging studies in Rett syndrome. Brain Dev. 23, S62–S71 (2001).
Subramaniam, B., Naidu, S. & Reiss, A. L. Neuroanatomy in Rett syndrome: cerebral cortex and posterior fossa. Neurology 48, 399–407 (1997).
Reiss, A. L. et al. Neuroanatomy of Rett syndrome: a volumetric imaging study. Ann. Neurol. 34, 227–234 (1993).
Hoogman, M. et al. Subcortical brain volume differences in participants with attention deficit hyperactivity disorder in children and adults—Authors’ reply. Lancet Psychiatry 4, 440–441 (2017).
Dark, C., Homman-Ludiye, J. & Bryson-Richardson, R. J. The role of ADHD associated genes in neurodevelopment. Dev. Biol. 438, 69–83 (2018).
Martin, C. A. et al. Mutations in genes encoding condensin complex proteins cause microcephaly through decatenation failure at mitosis. Genes Dev. 30, 2158–2172 (2016).
Khan, T. N. et al. Mutations in NCAPG2 cause a severe neurodevelopmental syndrome that expands the phenotypic spectrum of condensinopathies. Am. J. Hum. Genet. 104, 94–111 (2019).
Perche, O. et al. Combined deletion of two Condensin II system genes (NCAPG2 and MCPH1) in a case of severe microcephaly and mental deficiency. Eur. J. Med. Genet. 56, 635–641 (2013).
Ji, T., Wu, Y., Wang, H., Wang, J. & Jiang, Y. Diagnosis and fine mapping of a deletion in distal 11q in two Chinese patients with developmental delay. J. Hum. Genet. 55, 486–489 (2010).
Chen, C. P. et al. A de novo 7.9 Mb deletion in 22q13.2->qter in a boy with autistic features, epilepsy, developmental delay, atopic dermatitis and abnormal immunological findings. Eur. J. Med. Genet. 53, 329–332 (2010).
Nishide, K. & Hirano, T. Overlapping and non-overlapping functions of condensins I and II in neural stem cell divisions. PLoS Genet. 10, e1004847 (2014).
Piazza, I. et al. Association of condensin with chromosomes depends on DNA binding by its HEAT-repeat subunits. Nat. Struct. Mol. Biol. 21, 560–568 (2014).
Frosi, Y. & Haering, C. H. Control of chromosome interactions by condensin complexes. Curr. Opin. Cell Biol. 34, 94–100 (2015).
Ono, T., Fang, Y., Spector, D. L. & Hirano, T. Spatial and temporal regulation of Condensins I and II in mitotic chromosome assembly in human cells. Mol. Biol. Cell 15, 3296–3308 (2004).
Ono, T. et al. Differential contributions of condensin I and condensin II to mitotic chromosome architecture in vertebrate cells. Cell 115, 109–121 (2003).
Hartl, T. A., Smith, H. F. & Bosco, G. Chromosome alignment and transvection are antagonized by condensin II. Science 322, 1384–1387 (2008).
Naumova, N. et al. Organization of the mitotic chromosome. Science 342, 948–953 (2013).
Ward, J. R. et al Condensin I and condensin II proteins form a LINE-1 dependent super condensin complex and cooperate to repress LINE-1. Nucleic Acids Res. 50, 10680–10694 (2022).
Ward, J. R. et al. Condensin II and GAIT complexes cooperate to restrict LINE-1 retrotransposition in epithelial cells. PLoS Genet. 13, e1007051 (2017).
Schuster, A. T., Sarvepalli, K., Murphy, E. A. & Longworth, M. S. Condensin II subunit dCAP-D3 restricts retrotransposon mobilization in Drosophila somatic cells. PLoS Genet. 9, e1003879 (2013).
Kemp, J. R. & Longworth, M. S. Crossing the LINE toward genomic instability: LINE-1 retrotransposition in cancer. Front. Chem. 3, 68 (2015).
Lander, E. S. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Hoskins, R. A. et al. Heterochromatic sequences in a Drosophila whole-genome shotgun assembly. Genome Biol. 3, RESEARCH0085 (2002).
Smith, C. D., Shu, S., Mungall, C. J. & Karpen, G. H. The Release 5.1 annotation of Drosophila melanogaster heterochromatin. Science 316, 1586–1591 (2007).
Muotri, A. R. et al. L1 retrotransposition in neurons is modulated by MeCP2. Nature 468, 443–446 (2010).
Yu, F., Zingler, N., Schumann, G. & Stratling, W. H. Methyl-CpG-binding protein 2 represses LINE-1 expression and retrotransposition but not Alu transcription. Nucleic Acids Res. 29, 4493–4501 (2001).
Shpyleva, S., Melnyk, S., Pavliv, O., Pogribny, I. & Jill James, S. Overexpression of LINE-1 retrotransposons in autism brain. Mol. Neurobiol. 55, 1740–1749 (2018).
Balestrieri, E. et al. Human endogenous retroviruses and ADHD. World J. Biol. Psychiatry 15, 499–504 (2014).
Balestrieri, E. et al. Transcriptional activity of human endogenous retrovirus in Albanian children with autism spectrum disorders. N. Microbiol. 39, 228–231 (2016).
Dai, L., Huang, Q. & Boeke, J. D. Effect of reverse transcriptase inhibitors on LINE-1 and Ty1 reverse transcriptase activities and on LINE-1 retrotransposition. BMC Biochem. 12, 18 (2011).
Jones, R. B. et al. Nucleoside analogue reverse transcriptase inhibitors differentially inhibit human LINE-1 retrotransposition. PLoS ONE 3, e1547 (2008).
Li, W. Z., Li, S. L., Zheng, H. Y., Zhang, S. P. & Xue, L. A broad expression profile of the GMR-GAL4 driver in Drosophila melanogaster. Genet. Mol. Res. 11, 1997–2002 (2012).
Kammermeier, L. et al. Differential expression and function of the Drosophila Pax6 genes eyeless and twin of eyeless in embryonic central nervous system development. Mech. Dev. 103, 71–78 (2001).
Callaerts, P. et al. Drosophila Pax-6/eyeless is essential for normal adult brain structure and function. J. Neurobiol. 46, 73–88 (2001).
Wang, Y. C. et al. Drosophila intermediate neural progenitors produce lineage-dependent related series of diverse neurons. Development 141, 253–258 (2014).
Bayraktar, O. A. & Doe, C. Q. Combinatorial temporal patterning in progenitors expands neural diversity. Nature 498, 449–455 (2013).
Hrdlicka, L. et al. Analysis of twenty-four Gal4 lines in Drosophila melanogaster. Genesis 34, 51–57 (2002).
Heslip, T. R., Theisen, H., Walker, H. & Marsh, J. L. Shaggy and dishevelled exert opposite effects on wingless and decapentaplegic expression and on positional identity in imaginal discs. Development 124, 1069–1078 (1997).
Ma, C. & Moses, K. Wingless and patched are negative regulators of the morphogenetic furrow and can affect tissue polarity in the developing Drosophila compound eye. Development 121, 2279–2289 (1995).
Royet, J. & Finkelstein, R. hedgehog, wingless and orthodenticle specify adult head development in Drosophila. Development 122, 1849–1858 (1996).
Baonza, A. & Freeman, M. Control of Drosophila eye specification by wingless signalling. Development 129, 5313–5322 (2002).
Pinto-Teixeira, F., Konstantinides, N. & Desplan, C. Programmed cell death acts at different stages of Drosophila neurodevelopment to shape the central nervous system. FEBS Lett. 590, 2435–2453 (2016).
Bayraktar, O. A., Boone, J. Q., Drummond, M. L. & Doe, C. Q. Drosophila type II neuroblast lineages keep prospero levels low to generate large clones that contribute to the adult brain central complex. Neural Dev. 5, 26 (2010).
Wallace, K., Liu, T. H. & Vaessin, H. The pan-neural bHLH proteins DEADPAN and ASENSE regulate mitotic activity and cdk inhibitor dacapo expression in the Drosophila larval optic lobes. Genesis 26, 77–85 (2000).
Chang, Y. H., Keegan, R. M., Prazak, L. & Dubnau, J. Cellular labeling of endogenous retrovirus replication (CLEVR) reveals de novo insertions of the gypsy retrotransposable element in cell culture and in both neurons and glial cells of aging fruit flies. PLoS Biol. 17, e3000278 (2019).
Ardeljan, D. et al. Cell fitness screens reveal a conflict between LINE-1 retrotransposition and DNA replication. Nat. Struct. Mol. Biol. 27, 168–178 (2020).
Thomas, C. A. et al. Modeling of TREX1-dependent autoimmune disease using human stem cells highlights L1 accumulation as a source of neuroinflammation. Cell Stem Cell 21, 319–331.e318 (2017).
Sun, W., Samimi, H., Gamez, M., Zare, H. & Frost, B. Pathogenic tau-induced piRNA depletion promotes neuronal death through transposable element dysregulation in neurodegenerative tauopathies. Nat. Neurosci. 21, 1038–1048 (2018).
Krug, L. et al. Retrotransposon activation contributes to neurodegeneration in a Drosophila TDP-43 model of ALS. PLoS Genet. 13, e1006635 (2017).
Czech, B. et al. An endogenous small interfering RNA pathway in Drosophila. Nature 453, 798–802 (2008).
Kawamura, Y. et al. Drosophila endogenous small RNAs bind to Argonaute 2 in somatic cells. Nature 453, 793–797 (2008).
Ghildiyal, M. et al. Endogenous siRNAs derived from transposons and mRNAs in Drosophila somatic cells. Science 320, 1077–1081 (2008).
Sala, L. et al. AGO2 silences mobile transposons in the nucleus of quiescent cells. Nat. Struct. Mol. Biol. 30, 1985–1995 (2023).
Hain, D. et al. Natural variation of the amino-terminal glutamine-rich domain in Drosophila argonaute2 is not associated with developmental defects. PLoS ONE 5, e15264 (2010).
Deutschman, E. et al. Comparing and contrasting the effects of Drosophila condensin II subunit dCAP-D3 overexpression and depletion in vivo. Genetics 210, 531–546 (2018).
Erwin, J. A. et al. L1-associated genomic regions are deleted in somatic cells of the healthy human brain. Nat. Neurosci. 19, 1583–1591 (2016).
Muotri, A. R. et al. Somatic mosaicism in neuronal precursor cells mediated by L1 retrotransposition. Nature 435, 903–910 (2005).
Zhu, X. et al. Machine learning reveals bilateral distribution of somatic L1 insertions in human neurons and glia. Nat. Neurosci. 24, 186–196 (2021).
Jacob-Hirsch, J. et al. Whole-genome sequencing reveals principles of brain retrotransposition in neurodevelopmental disorders. Cell Res. 28, 187–203 (2018).
Pasyukova, E., Nuzhdin, S., Li, W. & Flavell, A. J. Germ line transposition of the copia retrotransposon in Drosophila melanogaster is restricted to males by tissue-specific control of copia RNA levels. Mol. Gen. Genet. 255, 115–124 (1997).
Borie, N., Maisonhaute, C., Sarrazin, S., Loevenbruck, C. & Biemont, C. Tissue-specificity of 412 retrotransposon expression in Drosophila simulans and D. melanogaster. Heredity 89, 247–252 (2002).
Lankenau, S., Corces, V. G. & Lankenau, D. H. The Drosophila micropia retrotransposon encodes a testis-specific antisense RNA complementary to reverse transcriptase. Mol. Cell Biol. 14, 1764–1775 (1994).
Haoudi, A. et al. Developmental expression analysis of the 1731 retrotransposon reveals an enhancement of Gag-Pol frameshifting in males of Drosophila melanogaster. Gene 196, 83–93 (1997).
Jiang, Y. & Reichert, H. Drosophila neural stem cells in brain development and tumor formation. J. Neurogenet. 28, 181–189 (2014).
Fueyo, R., Judd, J., Feschotte, C. & Wysocka, J. Roles of transposable elements in the regulation of mammalian transcription. Nat. Rev. Mol. Cell Biol. 23, 481–497 (2022).
Sasaki, T. et al. Possible involvement of SINEs in mammalian-specific brain formation. Proc. Natl Acad. Sci. USA 105, 4220–4225 (2008).
Notwell, J. H., Chung, T., Heavner, W. & Bejerano, G. A family of transposable elements co-opted into developmental enhancers in the mouse neocortex. Nat. Commun. 6, 6644 (2015).
Garza, R. et al. LINE-1 retrotransposons drive human neuronal transcriptome complexity and functional diversification. Sci. Adv. 9, eadh9543 (2023).
Wang, L. et al. Retrotransposon activation during Drosophila metamorphosis conditions adult antiviral responses. Nat. Genet. 54, 1933–1945 (2022).
Kumar, A., Bello, B. & Reichert, H. Lineage-specific cell death in postembryonic brain development of Drosophila. Development 136, 3433–3442 (2009).
Baillie, J. K. et al. Somatic retrotransposition alters the genetic landscape of the human brain. Nature 479, 534–537 (2011).
Hassan, A. et al. Condensin I subunit Cap-G is essential for proper gene expression during the maturation of post-mitotic neurons. Elife https://doi.org/10.7554/eLife.55159 (2020).
Longworth, M. S. et al. A shared role for RBF1 and dCAP-D3 in the regulation of transcription with consequences for innate immunity. PLoS Genet. 8, e1002618 (2012).
Gasior, S. L., Wakeman, T. P., Xu, B. & Deininger, P. L. The human LINE-1 retrotransposon creates DNA double-strand breaks. J. Mol. Biol. 357, 1383–1393 (2006).
Belgnaoui, S. M., Gosden, R. G., Semmes, O. J. & Haoudi, A. Human LINE-1 retrotransposon induces DNA damage and apoptosis in cancer cells. Cancer Cell Int. 6, 13 (2006).
De Cecco, M. et al. L1 drives IFN in senescent cells and promotes age-associated inflammation. Nature 566, 73–78 (2019).
Simon, M. et al. LINE1 derepression in aged wild-type and SIRT6-deficient mice drives inflammation. Cell Metab. 29, 871–885.e875 (2019).
Heinrich, M. J. et al. Endogenous double-stranded Alu RNA elements stimulate IFN-responses in relapsing remitting multiple sclerosis. J. Autoimmun. 100, 40–51 (2019).
Cuellar, T. L. et al. Silencing of retrotransposons by SETDB1 inhibits the interferon response in acute myeloid leukemia. J. Cell Biol. 216, 3535–3549 (2017).
Tunbak, H. et al. The HUSH complex is a gatekeeper of type I interferon through epigenetic regulation of LINE-1s. Nat. Commun. 11, 5387 (2020).
Zhao, K. et al. LINE1 contributes to autoimmunity through both RIG-I- and MDA5-mediated RNA sensing pathways. J. Autoimmun. 90, 105–115 (2018).
Martin, M., Hiroyasu, A., Guzman, R. M., Roberts, S. A. & Goodman, A. G. Analysis of Drosophila STING reveals an evolutionarily conserved antimicrobial function. Cell Rep. 23, 3537–3550.e3536 (2018).
Holleufer, A. et al. Two cGAS-like receptors induce antiviral immunity in Drosophila. Nature 597, 114–118 (2021).
Watanabe, T. et al. Involvement of host cellular multivesicular body functions in hepatitis B virus budding. Proc. Natl Acad. Sci. USA 104, 10205–10210 (2007).
Theisen, D. J. et al. WDFY4 is required for cross-presentation in response to viral and tumor antigens. Science 362, 694–699 (2018).
McCormick, D., Lin, Y. T. & Grey, F. Identification of host factors involved in human cytomegalovirus replication, assembly, and egress using a two-step small interfering RNA screen. mBio https://doi.org/10.1128/mBio.00716-18 (2018)
Vazquez, B. N. et al. SIRT7 mediates L1 elements transcriptional repression and their association with the nuclear lamina. Nucleic Acids Res. 47, 7870–7885 (2019).
Trimborn, M., Schindler, D., Neitzel, H. & Hirano, T. Misregulated chromosome condensation in MCPH1 primary microcephaly is mediated by condensin II. Cell Cycle 5, 322–326 (2006).
Wood, J. L., Liang, Y., Li, K. & Chen, J. Microcephalin/MCPH1 associates with the Condensin II complex to function in homologous recombination repair. J. Biol. Chem. 283, 29586–29592 (2008).
Yamashita, D. et al. MCPH1 regulates chromosome condensation and shaping as a composite modulator of condensin II. J. Cell Biol. 194, 841–854 (2011).
Houlard, M. et al. MCPH1 inhibits Condensin II during interphase by regulating its SMC2-Kleisin interface. Elife https://doi.org/10.7554/eLife.73348 (2021).
Thibault, S. T. et al. A complementary transposon tool kit for Drosophila melanogaster using P and piggyBac. Nat. Genet. 36, 283–287 (2004).
Acknowledgements
We thank members of the Longworth laboratory for their insightful discussions and advice during the performance of experiments and the preparation of the manuscript. We thank Dr. Judy Drazba in the Lerner Research Institute Imaging Core for assistance with imaging. We thank Dr. Josh Dubnau (Stonybrook University), Dr. Bob Johnston (Johns Hopkins University), Dr. Heather Broihier (Case Western Reserve University), and Dr. Pam Vanderzalm (John Carroll University) for their generous donations of fly stocks. Stocks obtained from the Bloomington Drosophila Stock Center (NIH P40OD018537) were used in this study. Transgenic fly stocks were also obtained from the Vienna Drosophila Resource Center (VDRC, www.vdrc.at). This work was supported by a Lisa Dean Moseley Foundation Award to M.S.L. and by a postdoctoral fellowship award F32HD114499 from NIH/NICHD.
Author information
Authors and Affiliations
Contributions
B.I.C., M.J.T. and M.S.L. conceived and designed the study and experiments. B.I.C., M.J.T., J.Ri., J.Ru., S.T., T.W. and M.S.L. performed experiments. B.I.C., M.J.T., J.Ri., J.Ru. and M.S.L. analyzed data. B.I.C., M.J.T. and M.S.L. interpreted the data. M.J.T. and M.S.L. wrote and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Tony Southall and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Crawford, B.I., Talley, M.J., Russman, J. et al. Condensin-mediated restriction of retrotransposable elements facilitates brain development in Drosophila melanogaster. Nat Commun 15, 2716 (2024). https://doi.org/10.1038/s41467-024-47042-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-024-47042-9
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.