Abstract
G protein-coupled receptors (GPCRs) constitute the largest family of membrane proteins and are important drug targets. The discovery of drugs targeting these receptors and their G protein signaling properties are based on assays mainly performed with modified receptors expressed in heterologous cells. However, GPCR responses may differ in their native environment. Here, by using highly sensitive Gi/o sensors, we reveal specific properties of Gi/o protein-mediated responses triggered by GABAB, α2 adrenergic and cannabinoid CB1 receptors in primary neurons, different from those in heterologous cells. These include different profiles in the Gi/o protein subtypes-mediated responses, and differences in the potencies of some ligands even at similar receptor expression levels. Altogether, our results show the importance of using biosensors compatible with primary cells for evaluating the activities of endogenous GPCRs in their native environment.
Similar content being viewed by others
Introduction
G protein-coupled receptors (GPCRs) are ubiquitously expressed in every cell type, and constitute the largest family of membrane proteins1. They participate in the regulation of a large variety of physiological functions2. Their dysregulation or malfunction can be the cause of numerous diseases3. Accordingly, controlling their activity with selective drugs can have multiple therapeutic effects, as illustrated by the large proportion of the clinical drugs targeting GPCRs4.
Most of the physiological functions of GPCRs are mediated through the coupling of G proteins2. Eventually, GPCRs are able to activate several G protein subtypes, such as Gi/o, Gs, Gq, and G12/13 family5,6,7,8, but this may vary depending on the use of ligands, a phenomenon called ligand biased effect9,10, or/and the cellular environment known as system bias11. This concept of functional selectivity11, that combined effect of ligand and system bias, has been largely used in the characterization of ligands12,13,14, but the signalling properties and identification of potential drug candidates targeting GPCRs are performed in heterologous cells. Since the cellular environment is critical, the analysis of G protein response profile of endogenous receptors in their native environment is therefore of much interest to validate drug candidates. Indeed, it becomes clear that many interacting proteins, post-translational modifications, localization in specific compartments, lipid membrane and ion environments largely influence GPCR signalling properties15,16. It is thus essential to examine the effects of drugs on the signalling properties of GPCRs in their native environments.
Functional analysis of endogenous GPCRs in their native environment is usually performed through the measurement of second messengers, such as cAMP, inositol phosphate or Ca2+17, or ion channel regulation2,18. Due to receptor reserves and signal amplification, such assays are not precise for estimating drug activity, potency and efficacy9. G proteins are first effectors of GPCRs, such that measuring their activation is less prone to signal amplification18. GTPγS binding assay can detect G protein activation in native cells or tissues19,20. However it requires the preparation of membranes and these assays are not specific for one G protein subtype, unless an immunoprecipitation of the G protein is added21. Nowadays, much hope in studying native GPCRs is based on the development of specific optical biosensors that must be sensitive enough to detect the activity of native receptors22 usually expressed at very low levels, but most of them are still not compatible with these endogenously expressed receptors13,23,24,25,26.
In the central nervous system, GPCRs are essential regulators of synaptic transmission, acting at both pre- and post-synaptic levels, as well as in glial cells27. More than half of neuronal GPCRs can couple to one or more of the various Gi/o protein subtypes5, but their expression profiles vary in different native tissues or cell types and specific cellular function depends on distinct Gi/o protein subtypes12,28,29. It is therefore essential to determine which G protein can be activated by a GPCR upon stimulation with various agonists in its native environment.
In the present study, we describe BRET-based sensors for each Gi/o protein subtypes that can be used to study endogenous GPCRs in living neurons. Our study focuses on three types of GPCRs that play important roles in modulating synaptic activity, the GABAB30, α2 adrenergic31,32 and cannabinoid CB1 receptors33. We could detect the activation of Gi/o proteins from a small number of neurons and monitor both the kinetics and dose-response of the effect mediated by various compounds including agonists, antagonists and positive allosteric modulators, in different types of neurons. Our data reveal differences in the profile of Gi/o protein subtypes mediated responses and agonist potencies between these recombinant and native receptors in HEK293 cells and neurons, respectively. In addition, our results show a major difference in Gi1 versus GoA protein activation induced by a CB1 receptor agonist in neurons, but not in HEK293 cells. Finally, different composition of Gγ subunits in the neurons can also lead to specific Gi/o protein responses. Altogether, our results reveal the importance of evaluating GPCR activities in their native environment and highlight the need for sensitive biosensors compatible with native receptors in their natural environment.
Results
Gi/o protein sensors for endogenous GPCRs in neurons
To measure the rearrangement or dissociation of the Gi/o proteins upon receptor activation in live cells and neurons, we used a series of BRET-based biosensors. They are based on the use of a luciferase as an energy donor inserted in the Gα subunit and the fluorescent protein Venus as an energy acceptor attached to the Gγ subunit, as reported for previous BRET- and FRET-based G protein sensors23,24,25. To monitor these sensors in conditions close to the physiological ones, in our approach, the cells were cotransfected only with the luciferase-tagged Gα subunit and Venus-tagged Gγ subunit (VenusGγ), while the endogenous Gβ subunits were used (Fig. 1a). In these experiments, the amount of Gβγ complexes that can produce a BRET signal with Gα is expected to be limited by the endogenous level of Gβ subunits (Supplementary Fig. 1a) that are able to form a complex with the VenusGγ. We used mostly the mouse cerebellum granule neurons (CGNs) that constitute the largest homogenous neuronal population of the mammalian brain34.
a Scheme of the BRET-based sensors to detect the activity of the endogenous GPCR in neurons. b Scheme of the Gαi/oNluc and VenusGγ2 constructs co-transfected in the neurons. Sequence of the linker (L) before and after Nluc was indicated. Gγ was fused with Venus in the N-terminal. c Composition of G protein sensors made of the different Gαi/oNluc and VenusGγ2. d Kinetics of the BRET signal between the indicated Gα constructs (Gαi1Nluc or Gαi1Rluc8) and VenusGγ2 in CGNs. Buffer or baclofen (100 μM) were injected at the indicated time (arrow). The data are representative of BRET ratios from five independent experiments. e Representative fluorescent images of Venus and DAPI in CGNs transfected with GαoANluc and VenusGγ2 or in HEK293 cells transfected with GαoANluc, Gβ1 and VenusGγ2 from three independent experiments. Scale bar: 20 μm. f Kinetics of the BRET signal between Gαi1Nluc and VenusGγ2 in CGNs. Buffer or baclofen (100 μM) or competitive antagonist CGP64213 (10 μM) were injected at the indicated time. The data are representative of the mean ± SEM of BRET ratios from three independent experiments. The raw data and p-values are available in source data provided as a Source Data file.
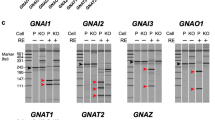
Our sensors rely on the use of a small luciferase, the Oplophorus nanoluciferase (Nluc; 19 kDa) that produces a more intense and sustained luminescence signal than other commonly used luciferases from Renilla luciferase (Rluc and Rluc8; 36 kDa)35. Nluc was inserted in the helical domain of different Gαi/o subunits similarly to previously reported G protein BRET sensors with Rluc23,24 and Nluc36, and the constructs were named GαNluc (Fig. 1b). The Nluc was inserted after residue 91 (for Gαi1, Gαi2, Gαi3, GαoA and GαoB), or 113 (for Gαz) (Fig. 1c). Of note, GαoA has 94% sequence identity with GαoB and around 70% with Gαi1, Gαi2 and Gαi337. To facilitate the insertion of Nluc, a short and flexible linker sequence (SGGGGS) was added at the N- and C-terminal ends of Nluc. For BRET signal measurement, Venus was fused to the N-terminus of Gγ2 subunit with the amino acid sequence KLGT serving as a linker, and the construct is named VenusGγ2 (Fig. 1b). To show the advantage of using Nluc over Rluc8 in measuring agonist-induced BRET change in neurons, Rluc8 was inserted at the same position as Nluc in Gαi1 (named Gαi1Rluc8) and GαoA (named GαoARluc8). These Gαi1Rluc8 and GαoARluc8 constructs are highly similar to the constructs reported for the Gα protein of TRUPATH BRET sensors (Supplementary Fig. 1b). The GαoANluc construct showed much higher luminescence intensity at emission 480 nm compared with the subunits GαRluc8 in CGNs, for the same amount of cDNA co-transfected with VenusGγ2 (Supplementary Fig. 1c). In addition, the basal BRET ratio of the constructs Gαi1Nluc and GαoANluc co-transfected with VenusGγ2 was also more stable over time, compared to the sensors that used subunits GαRluc8 (Fig. 1d and Supplementary Fig. 1d). We also verified that the co-expression of the constructs GαNluc + VenusGγ2 for the different Gα subunits did not change the expression level of endogenous Gβ subunits (Supplementary Fig. 1e). Altogether, the G protein Nluc-biosensors are expected to produce a much improved signal-to-noise ratio for BRET measurement in neurons.
We then validated our different Gi/o protein sensors with the GABAB receptor as a prototype Gi/o-coupled receptor, which is abundantly expressed in many types of neurons38, including in CGNs39. Interestingly, despite a low percentage of transfected neurons compared with HEK293 cells (Fig. 1e), the GABAB receptor specific agonist baclofen largely decreased the BRET signal with the sensor Gαi1Nluc + VenusGγ2 (Fig. 1d, Supplementary Fig. 2a) and GαoANluc + VenusGγ2 (Supplementary Fig. 1d, 2b), while the change was barely detectable with Gαi1Rluc8 + VenusGγ2 (Fig. 1d) and GαoARluc8 + VenusGγ2 (Supplementary Fig. 1d). In kinetics experiments, baclofen induced a rapid and strong decrease of BRET signal in CGNs, that was reversed by the GABAB receptor competitive antagonist CGP64213 for both the Gi1 (Fig. 1f) and GoA (Supplementary Fig. 2c) sensors. As a control, no change of BRET signal was measured when Gαi1Nluc was expressed in the absence of VenusGγ2 or with Venus (Supplementary Fig. 2d). Similarly, no change of BRET signal was measured with the dominant negative mutants Gαi1Nluc-S47A40 or Gαi1Nluc-G202T41 (Supplementary Fig. 2e) when they are co-expressed with VenusGγ2. This is consistent with the unability of these mutants to exchange their GDP for GTP. We also verified that the co-expression of Gβ with the Gα and Gγ2 was not required for a large BRET change signal induced by baclofen (Supplementary Fig. 2f). Altogether, these results validated the use of the Gαi1Nluc and GαoANluc -based BRET sensors in neurons.
These Gi/o protein Nluc sensors were highly sensitive even though only about 2% of CGNs expressed the sensor as measured by the fluorescence of the transfected VenusGγ2. This low level of transfection is consistent with previous data reported with CGNs42, even though any transfected cell is expected to overexpress the sensors thanks to the strong promotor. The transfected neurons displayed normal morphology with dendrites and branches (Fig. 1e). It indicates that the Gαi/o protein Nluc-sensors were sensitive enough to monitor endogenous GPCR activation in a small number of neurons. In addition, they are compatible with measurements in 96-well plates. The amount of cDNA to be transfected in CGNs was optimized for each sensor (Supplementary Fig. 2g). Finally, our sensors are also compatible with the measurement of the activation of endogenous GPCRs expressed in the HEK293 cells such as the lysophosphatidic acid receptor (Supplementary Fig. 3a). In these experiments, the Gi2 showed a weak response than those generated by the other Gi/o sensors (Supplementary Fig. 3a), and this is unrelated to differences in expression levels of these sensors (Supplementary Fig. 3b).
We then compared the agonist-induced BRET signals obtained with the same Gαi/o sensor in the two different cell types, CGNs and HEK293 cells (Fig. 2a). Conditions were optimized such that the GABAB receptor expression level in HEK293 cells was similar to that in CGNs (Fig. 2b). Since the basal BRET signal in the absence of agonist can be different between the G protein sensors (Supplementary Fig. 3c), the signal was expressed as the agonist-induced change in BRET ratio expressed as percentage of the basal signal. The results showed that in CGNs, the BRET changes induced by baclofen are significantly different from those measured in HEK293 cells except for the two Go sensors (Fig. 2a). Indeed, a larger BRET signal is measured in CGNs with Gi1 and Gi3, and a large signal is observed with Gz in HEK293 cells while absent in CGNs. The differences observed are related neither to the expression levels of the GαNluc sensor components (Fig. 2c) nor to the endogenous Gαi/o mRNA expression levels (Fig. 2d). Though lower signal was observed in Gi2 in CGNs, the difference of baclofen response can be detected in dose-dependent manner (Supplementary Fig. 3d). Similar pEC50 values were measured in three Gi subtype sensors and two Go subtype sensors in CGNs upon baclofen activation (Supplementary Fig. 3d). Altogether, the data showed different abilities of the GABAB receptor to generate responses by the different Gi/o subtypes in transfected HEK293 cells and in CGNs, then revealing the importance of studying GPCRs in their native environment.
a Baclofen-induced change in BRET ratio between GαNluc and VenusGγ2 in CGNs or transfected HEK293 cells for the indicated Gi/o proteins, and expressed as percentage of the basal signal ((BRETbasal-BRETagonist/BRETbasal) x 100). CGNs were co-transfected with VenusGγ2 (50 ng) and the Nluc-tagged Gαi1 (25 ng), Gαi2 (75 ng), Gαi3 (75 ng), GαoA (25 ng), GαoB (25 ng) or Gαz (25 ng), while HEK293 cells were co-transfected with GB1 (9 ng), GB2 (12 ng), Gβ1 (10 ng), VenusGγ2 (10 ng) and the Nluc-tagged Gαi1 (1 ng), Gαi2 (3 ng), Gαi3 (3 ng), GαoA (1 ng), GαoB (1 ng) or Gαz (1.5 ng) per well in 96-well plate. b Expression of GB1 subunit of GABAB receptor in the cell membrane of CGNs, mock HEK293 cells, and HEK293 cells transfected with indicated amount of GB1 and GB2, detected by western blotting. Values are mean ± SEM normalized as fold of GB1 expression in CGNs from four independent experiments. c Expression of the Nluc-tagged Gα transfected in CGNs or in HEK293 cells in (a), measured by the luminescent signal at 480 nm. The values of individual well are shown. Values are mean ± SEM from four biologically independent experiments each performed in triplicates or quadruplicate in (a) and (c). Data in (a) are analysed using unpaired t-test (two-tailed) to determine significance. Data in (c) are analysed using one-way analysis of variance (ANOVA) with a Dunnett’s post-hoc multiple comparison test to determine significance (compared with Gi1 group). ****p < 0.0001, ***p < 0.001, **p < 0.01 and *p < 0.05 and not significant (ns). d Expression of the genes encoding indicated Gα in CGNs (mouse species: Gnai1, Gnai2, Gnai3, Gnao-1, Gnao-2, Gnaz) and HEK293 cells (human species: GNAI1, GNAI2, GNAI3, GNAO-1, GNAO-2, GNAZ). Values were determined by qRT-PCR and mean ± SEM normalized to Rplp0 (mouse) or RPLP0 (humans) from three biologically independent experiments each performed in quadruplicate. The values of individual sample are shown. The raw data and p-values are available in source data provided as a Source Data file.
Lower agonist potencies for the endogenous GABAB receptor in neurons
We have further validated these G protein sensors by measuring the potency of different orthosteric GABAB receptor ligands (Fig. 3a) in both CGNs and HEK293 cells. Similar potencies were obtained for the Gi1 and GoA for baclofen and other agonists, and for the antagonist CGP64213 in CGNs (Fig. 3b–f, Supplementary Table 1). The GABAB receptor agonists APPA and SKF 97541 were more potent than GABA and baclofen in both CGNs (Fig. 3c–f) and HEK293 cells (Supplementary Fig. 4a), which was consistent with previous findings using the GTPγS assay in brain tissues21. pEC50s obtained in CGNs were correlated with those measured in HEK293 cells (Fig. 3e, f), even though they were more than 10 times higher in HEK293 cells, except for baclofen where the difference is lower (Supplementary Table 1, Fig. 3e, f). Similar results were obtained in the hippocampal neurons and cortical neurons (Supplementary Fig. 4b–d and Supplementary Table 1). These differences in baclofen potencies observed between transfected HEK293 cells and native neurons were not due to the difference in receptor expression. Indeed, when the expression level GABAB receptors was similar to that in CGNs (Fig. 2b), the baclofen pEC50 was similar to that observed with higher expression levels, and still significantly different from that measured in CGNs (Fig. 3g).
a Scheme of the GABAB receptor, made of GB1 and GB2 subunits, in CGNs and its various specific ligands (agonist, red; antagonists, blue; PAM, black). b Change of BRET signal induced by the GABAB receptor antagonist CGP64213 in presence of 20 μM baclofen (EC80) or different doses of baclofen in CGNs measured by Gi1 or GoA sensor. c, d Change of BRET ratio induced by various GABAB receptor agonists measured by Gi1 (c) or GoA (d) sensor. e, f Correlation of the agonist potencies (pEC50) between HEK293 cells and CGNs determined by the Gi1 (e) or GoA (f) BRET sensors. Dotted line is the correlation of pEC50 determined with HEK293 cells. Red line is fit of pEC50 in CGNs with the same slope as the dotted line. g The pEC50 of baclofen in CGNs and transfected HEK293 cells with indicated amount of GB1 and GB2 measured by Gi1 or GoA sensor. Data are mean ± SEM from at least three biologically independent experiments each performed in triplicate and analysed using one-way ANOVA with a Dunnett’s post-hoc multiple comparison test to determine significance. The n number of Gi1 and GoA group for CGNs, and HEK293 cells transfected with the indicated amount of GB1 and GB2 are 10, 4, 3, and 4, 3, 3, respectively. ****p < 0.0001, **p < 0.01 and not significant (ns). h Change of BRET ratio induced by baclofen with or without indicated concentrations of the GABAB PAM Rac BHFF in transfected HEK293 cells and CGNs measured by Gi1 sensor. Data are normalized to maximum of baclofen response and mean ± SEM from at least three biologically independent experiments each performed in triplicate in (b–d) and (h). b n = 3; c n = 6; d n = 4; h CGNs n = 3, HEK293 cells n = 4. The raw data and p-values are available in source data provided as a Source Data file.
We have also validated our sensors for the analysis of positive allosteric modulators (PAMs). PAMs can synergistically enhance the activity of native neurotransmitter GPCRs in neurons43 and have a greater potential for drug development with less side effect44, including for the GABAB receptor45. It was recently demonstrated that these PAMs bind at the interface of the transmembrane domains of this dimeric receptor46,47,48 (Fig. 3a). Here we showed that Rac BHFF increased strongly the potency of baclofen-induced Gi1 (Fig. 3h) and GoA protein (Supplementary Fig. 5a) response in CGNs. However, in contrast to what is observed in HEK293 cells48,49 (Fig. 3h and Supplementary Fig. 5a–b), Rac BHFF alone (10 μM) did not increase the net BRET signal in CGNs (Fig. 3h and Supplementary Fig. 5a). It is nicely illustrated by concentration responses curves of the Rac BHFF obtained with transfected HEK293 cells but not so efficiently with neurons (Supplementary Fig. 5c). It suggests that the Rac BHFF agonist activity observed with the recombinant GABAB receptor is much higher than that measured for the receptor in its native environment. This slight agonist effect of Rac BHFF in CGNs is probably not due to the lower GABAB receptor expression in neurons compared to HEK293 cells. Indeed, the agonist activity of Rac BHFF can still be observed in HEK293 cells that expressed GABAB receptor at a similar level to that found in CGNs (Figs. 2b, 3h, Supplementary Fig. 5a).
Finally, we took advantage of our sensors to measure the constitutive activity50, in the absence of agonist, of the GABAB receptor that is usually observed in recombinant systems47,49,51,52,53,54,55. This constitutive activity was reversed by the competitive antagonists known for its inverse agonist activity49,51,55,56. Accordingly, the competitive antagonist CGP54626 increased the basal BRET signal between Gα and Gβγ for both the Gi1 and GoA sensors in the HEK293 cells (Supplementary Fig. 5d), as previously reported49. It is consistent with the properties of this inverse agonist to favour the Gαβγ heterotrimer by stabilizing the inactive conformation of the receptor. Interestingly, this increase of BRET signal was not observed in the CGNs (Supplementary Fig. 5d), indicating the native GABAB receptor has no or a very low constitutive activity that cannot be observed with our sensor. It is consistent with the weak intrinsic agonist activity of Rac BHFF in neurons.
Altogether, our results show that GABAB receptor ligands have different potencies and efficacies in the cell lines compared to neurons, which cannot be explained by a difference in GABAB receptor expression levels between the two cell types.
Various efficacies of endogenous Gi/o-coupled GPCRs in neurons
We then examined the activities of other endogenous Gi/o-coupled GPCRs in the CGNs. First, the agonist CP 55940 that actives both the cannabinoid receptor type 1 (CB1) and 2 (CB2) induced a strong change in both Gi1 and GoA BRET signal as observed for the GABAB receptor (Fig. 4a). It was expected that it is due to the activation of CB1 since this receptor is highly expressed in cerebellum33 compared to CB2 that is mainly expressed at the periphery57. When testing other GPCR agonists, a strong or significant change of BRET signal was measured with different ligands used at saturation concentrations for their expected receptors: brimonidine (also named UK-14304), pramipexole, eletriptan and carbachol known to activate α2 adrenoceptor58, D2/4 receptor59, 5-hydroxytryptamine receptor 1B/1D60 and the M2/4 muscarinic receptors61, respectively. In contrast, agonists of the platelet-activating factor (PAF) receptor and δ/μ opioid receptor DAMGO failed to induce a BRET signal change (Fig. 4a). Dose-response of brimonidine, CP 55940, pramipexole and carbachol further confirmed the specific activation of these endogenous Gi/o-coupled receptors (Fig. 4b). Altogether, our data correspond to the receptors for which higher amount of mRNA were detected in these cultured CGNs at the developmental stage investigated62.
a Net BRET signal of the Gi1 and GoA sensors in CGNs induced by specific agonists of various Gi/o-coupled GPCRs using a saturating concentration of the indicated ligands (PAF, 10 μM; DAMGO, 100 μM; eletriptan, 100 μM; brimonidine, 10 μM; carbachol, 100 μM; pramipexole, 100 μM; CP 55940, 20 μM and baclofen, 100 μM). Values are mean ± SEM from at least biologically independent experiments each performed in triplicate. The n number of the indicated treatments in Gi1 and GoA are 6, 4, 5, 4, 6, 4, 6, 7, 8, and 8, 6, 4, 6, 6, 6, 6, 6, 7, 8, respectively. Data are analysed using one-way ANOVA with a Dunnett’s post-hoc multiple comparison test to determine significance (compared with no treated group). ****p < 0.0001, *p < 0.05 and not significant (ns). b Dose-response of the agonists brimonidine, CP 55940, carbachol and pramipexole for the α2 adrenoceptor, CB1, muscarinic M2/4 and dopamine D2/4 receptors, respectively, in CGNs co-transfected with Gαi1Nluc and VenusGγ2 or GαoANluc and VenusGγ2. Data are normalized to maximum response of each compound and mean ± SEM from at least three biologically independent experiments each performed in triplicate. Gi1, brimonidine (n = 3), CP 55940 (n = 4), carbachol (n = 4) and pramipexole (n = 4); GoA, brimonidine (n = 3), CP 55940 (n = 3), carbachol (n = 4) and pramipexole (n = 3). The raw data and p-values are available in source data provided as a Source Data file.
Next, using α2 adrenoceptor we show that our G protein sensors can determine ligand efficacies, by evaluating the full agonist brimonidine and several reported partial agonists (clonidine, oxymetazoline, tizanidine and xylazine)63. In contrast, brimonidine exhibited the highest response of Gi1 and GoA protein in CGNs, while others showed less response (Fig. 5a). However, the discrimination between brimonidine and other agonists was not significantly observed in HEK293 cells transfected with α2A adrenoceptor (α2AAR) (Fig. 5b). It was not due to the high level of expression of α2AAR, since no discrimination between brimonidine and other agonists was observed in HEK293 cells that expressed α2AAR at a similar level to that found in CGNs (Fig. 5c). In these conditions (15 ng of α2AAR cDNA/well), the brimonidine pEC50 was similar to that observed with higher expression levels (Supplementary Fig. 6a), and still significantly different from that measured in CGNs (Fig. 5d), as observed previously for the GABAB receptor (Fig. 3g). Finally, the data showed a significant difference in the G protein responses mediated by the different Gi/o subtypes triggered by the recombinant α2AAR in HEK293 cells and the endogenous one in CGNs (Fig. 5e and Supplementary Fig. 6b). Of note, similarly to experiments above with GABAB receptor (Fig. 2c), the expression of the different GαNluc proteins was within a factor of three in each type of cell (Supplementary Fig. 6c).
a Effect of the indicated α2AR agonists on the net BRET of the Gi1 and GoA sensors in CGNs co-transfected with Gαi1Nluc and VenusGγ2 or GαoANluc and VenusGγ2 (amounts of cDNA as in Fig. 2a). b Effect of the indicated α2AR agonists on the net BRET of the Gi1 and GoA sensors in HEK293 cells co-transfected with the mouse α2AAR (10 ng, 20 ng or 50 ng), and Gβ1, VenusGγ2 and Gαi1Nluc or GαoANluc as in Fig. 2a. Saturating concentrations of brimonidine (10 μM), oxymetazoline (100 μM), xylazine (50 μM), clonidine (100 μM), tizanidine (100 μM) were used in (a, b). c Detection of α2AR in cell membranes of CGNs, mock-transfected HEK293 cells and HEK293 cells transfected with indicated amount of α2AR, by western blotting. Values are mean ± SEM normalized as fold of α2AR expression in CGNs from four independent experiments. d The pEC50 of brimonidine in CGNs (n = 3) and transfected HEK293 cells (n = 4) with indicated amount of α2AR measured by Gi1 or GoA sensor. e Percentage of change in BRET ratio between GαNluc and VenusGγ2 induced by brimonidine between GαNluc and VenusGγ2 in HEK293 cells (n = 3) (α2AR: 15 ng/well per 96-well plate) or CGNs (n = 4) for the indicated Gαi1, Gαi2, Gαi3, GαoA, GαoB or Gαz sensors (amounts of cDNA as in Fig. 2a). Values are mean ± SEM from at least three biologically independent experiments each performed in triplicate or quadruplicate in (a, b, d, e). a, n = 4; b, n = 3; (d, e), CGNs, n = 3; HEK293 cells, n = 4. Data are analysed using one-way ANOVA with a Dunnett’s post-hoc multiple comparison test to determine significance (compared with brimonidine in a, b). Data are analysed using unpaired t-test (two-tailed) in (e). ****p < 0.0001, ***p < 0.001, **p < 0.01 and not significant (ns). The raw data and p-values are available in source data provided as a Source Data file.
Our results demonstrated that Gi/o protein Nluc-biosensors can be widely applied to different endogenous Gi/o-coupled GPCRs and evaluate the ligand efficacies and G protein coupling profile in neurons. They also further revealed the importance of studying a GPCR in its native environment.
Difference between Gi and Go proteins revealed for the endogenous CB1 receptor
Finally, we took advantage of the major change in BRET signal observed for the cannabinoid receptor to investigate the G protein response in CGNs. While level of expression of the CB1 receptor in the two systems was similar in western blotting experiments (Fig. 6a), a higher CB1 receptor-mediated G protein response was observed with Gi1 and Gi3 only, with the native receptor in CGNs compared to the recombinant one in HEK293 cells (Fig. 6b, Supplementary Fig. 7a). Similar responses were observed with the other Gi/o subtype sensors between two types of cells. Again, as with the GABAB receptor (Fig. 2b) and α2AAR (Supplementary Fig. 6c), the expression of the different GαNluc proteins was within a factor of three in each type of cell (Supplementary Fig. 7b). And a small overexpression of the CB1 receptor by transfection of CGNs with the recombinant CB1 receptor gave similar results (Supplementary Fig. 7c, d). Interestingly, our G protein sensors revealed a higher potency of CP 55940, Win 55,212-2 and Bay 59-3074 in CGNs compared with transfected HEK293 cells for both Gi1 and GoA response (Fig. 6c, d and Supplementary Table 2). It is different from GABAB and α2AR, which showed higher agonist potencies in transfected HEK293 cells. Of note, when taking Win 55,212-2 as a reference ligand, Bay 59-3074 showed GoA bias in both CGNs and HEK293 cells with a bias factor of 1.6 and 1.3 respectively. CP 55940 showed a Gi1 bias in CGNs and a GoA bias in HEK293 cells with a bias factor of 1.6 and 1.4 respectively, indicating a difference between Gi1 and GoA proteins revealed by CP 55940 (Fig. 6c, d and Supplementary Tables 2 and 3).
a Detection of CB1 receptor in cell membranes of CGNs, mock-transfected HEK293 cells and HEK293 cells transfected with mouse CB1 cDNA (80 ng/well in 96-well plate), by western blotting. Values are mean ± SEM normalized as fold of CB1 expression in CGNs from three independent experiments. b Percentage of change in BRET ratio between GαNluc and VenusGγ2 induced by CP 55940 in HEK293 cells (CB1 cDNA; 80 ng/well per 96-well plate) or CGNs for the indicated Gαi1, Gαi2, Gαi3, GαoA, GαoB or Gαz sensors (amounts of cDNA as in Fig. 2a). Values are mean ± SEM from four biologically independent experiments each performed in triplicate. Data are analysed using unpaired t-test (two-tailed) to determine significance. ****p < 0.0001, *p < 0.05 and not significant (ns). c, d Change of BRET signal between GαNluc and VenusGγ2 induced by the indicated CB1 receptor agonists in HEK293 cells and CGNs measured by Gi1 and GoA sensors. The transfection was the same as in (b). Inset, correlation of the agonist potencies (pEC50) between HEK293 cells and CGNs determined by the G protein sensors. Dotted lines are the correlation of pEC50 determined with HEK293 cells. Red lines are the fit of pEC50 in CGNs with the same slope as the dotted lines. Data are normalized to maximum CP 55940 response and are mean ± SEM from at least three biologically independent experiments each performed in triplicate (HEK293 cells: Gi1, CP 55940 (n = 5), Win 55,212-2 (n = 5) and Bay 59-3074 (n = 4); GoA, CP 55940 (n = 5), Win 55,212-2 (n = 4) and Bay 59-3074 (n = 4). CGNs: Gi1, CP 55940 (n = 4), Win 55,212-2 (n = 4) and Bay 59-3074 (n = 4); GoA, CP 55940 (n = 4), Win 55,212-2 (n = 4) and Bay 59-3074 (n = 3)). The raw data and p-values are available in source data provided as a Source Data file.
Major influence of the Gγ subunit on the G protein coupling in CGNs
The influence of Gβγ subunit composition on G protein coupling64,65 was largely reported even though the underlying molecular mechanism is largely unknown. Therefore, we have compared sensors composed by GαNluc and different Gγ including VenusGγ2, VenusGγ8 or VenusGγ9, which are well-used in the previously reported G protein BRET sensors25,36. VenusGγ8 or VenusGγ9 belong to two different groups of Gγ subunits (Supplementary Fig. 8). Gγ8, like Gγ2, is well expressed in some regions of the brain, in contrast to Gγ9 that seems restricted to retina66. Interestingly, a major difference in G protein response profile of the GABAB receptor and α2 adrenoceptor was measured depending on the Gγ subunits used (Fig. 7a–c and Supplementary Fig. 9a–d), that clearly shows the impact of the Gγ subunit in these GPCR-mediated responses.
a, b Percentage of change in BRET ratio between GαNluc and VenusGγ8 (a) or VenusGγ9 (b) induced by baclofen, brimonidine and CP 55940 in CGNs for the indicated Gi/o proteins. CGNs were co-transfected with the indicated VenusGγ and Nluc-tagged Gαi1, Gαi2, Gαi3, GαoA, GαoB or Gαz. Values are mean ± SEM from biologically independent experiments (baclofen and brimonidine, n = 5; CP 55940, n = 3) each performed in triplicate. c Scheme illustrating the difference in Gi/o (Gi1, Gi2, Gi3, GoA, GoB and Gαz) and indicated Gγ protein (Gγ2, Gγ8 and Gγ9) responses in CGNs for the receptors GABAB, CB1 and α2AR in presence of the indicated agonists. For each receptor and Gi/o sensor, the size of the circle is the percentage of change in BRET ratio upon the agonist stimulation, for GABAB (Figs. 2a and a, b), α2AR (Figs. 5e and a, b) and CB1 (Figs. 6b and a, b) normalized to the Gi/o sensor that has the highest response: for Gγ2, Gi1 for both the GABAB and CB1, and Gαz for α2AR; for Gγ8, GoA for GABAB and α2AR and Gi1 for CB1; and for Gγ9, Gi1 for GABAB and CB1, and Gαz for α2AR. For Gγ8, the empty circle indicates an increase in the BRET ratio in contrast to other conditions where a decrease BRET ratio is measured. The raw data and p-values are available in source data provided as a Source Data file.
Discussion
Our study compares the pharmacological and G protein responses of native, versus recombinant Gi/o-coupled receptors in primary neurons and in heterologous cells, respectively. Such an analysis was made possible thanks to the sensitive BRET-based Gi/o protein sensors. These sensors allow a simple analysis of different types of GPCR ligands on each Gi/o protein subtypes in primary neurons in a medium throughput format (in 96-well plate). For the same G protein sensor, our study reveals important differences in agonist potencies between neurons and heterologous HEK293 cells (Fig. 8a). GABAB and α2 adrenergic receptor agonists displayed higher potencies in cell lines versus neurons even at similar receptor expression levels, in contrast to CB1 agonist. And when comparing the profile of response of different G protein sensors in the same cellular system, major differences of potencies can be observed, such as for a CB1 agonist (CP 55940) that shows high potency in mediating Gi1 response compared to GoA in neurons. These data highlight the importance of evaluating the activities of endogenous GPCRs in their native environment, and of developing biosensors allowing such analyses.
a Scheme illustrating the difference in agonist or PAM potencies for the GABAB and CB1 receptors between HEK293 cells and CGNs for the Gi1 sensor. For each cell and receptor, the relative size of the triangles is according to the pEC50 values of the indicated agonists. b Scheme illustrating the difference in Gi/o protein response (Gi1, Gi2, Gi3, GoA, GoB and Gz) between CGNs and HEK293 cells for the receptors GABAB, CB1 and α2AR in presence of the indicated agonists, when Gγ2 is used for all these Gi/o sensors. For each cell and receptor, the size of the circle is the percentage of change in BRET ratio upon the agonist stimulation for GABAB (Fig. 2a), α2AR (Fig. 5e) and CB1 (Fig. 6b) normalized to the Gi/o sensor that has the highest response: in HEK293 cells, GoA for GABAB, α2AR and Gi1 for CB1, and in CGNs, Gi1 for both the GABAB and CB1, and Gz for α2AR.
In order to be able to compare the properties of recombinant and native Gi/o-coupled GPCRs, we needed sensors for the various Gi/o proteins compatible with measurements of their activity in primary neurons. Here we showed that Nluc-based BRET sensors, could do so, most likely due to the high emission of Nluc, compared to Rluc8 used previously24,25, since the efficacy of transfection of these constructs in primary neurons remains very low (around 2%). The principle of our optical biosensors is not novel, since the rearrangement between Gα and Gβγ upon receptor activation using resonance energy transfer techniques was previously reported by FRET23, BRET24,25 including with the use of Nluc36. However, these biosensors have never been tested in primary neurons. Interestingly, other kind of biosensors compatible with the detection of endogenous G protein activity were recently reported22,36. Among them, the unimolecular “BERKY” sensors were used in live primary cells but they required the use of a viral vector. In addition, these sensors were not able to discriminate between different G protein subtypes, such as between Gi1, Gi2, Gi3, GoA, GoB and Gz since it is a unimolecular sensor using a peptide recognizing the GTP form of these different G proteins. Accordingly, they cannot be used to analyse the influence of the different Gβγ subunits as our sensors and the previous BRET sensors24,25,36 do. Instead, our sensors nicely revealed specific responses of each Gi/o protein triggered by some native GPCRs in primary neurons, which are different from what can be observed in heterologous cells. They work well for the most abundant neuronal GPCRs, but would likely need to be further improved for receptors expressed at lower levels.
We have to be cautious when comparing the profile of response of different G protein sensors in the same cellular system (Figs. 7 and 8b). Indeed, each G protein BRET sensor might have its own properties, with its specific range of signal, even though they were constructed similarly, and Gαi/o subtypes have a high sequence identity37. G protein sensor response can result either from a complete dissociation between Gα and Gβγ but also from a repositioning of Gβγ relative to Gα without a real dissociation23. Within the Gi/o protein family, using similar FRET-based G protein sensors between Gα and Gβγ, Frank et al. have proposed that receptor-activated Gi1, Gi2, Gi3 and Gz undergo subunit rearrangement rather than subunit dissociation, whereas Go proteins either rearrange with a very distinct pattern or dissociate during activation67. But even though there is good evidence that the activated Gα and Gβγ subunits actually loose affinity to each other and exchange faster depending of the G proteins, there is no evidence that the subunits separate completely in intact cells68.
When analysing the same G protein sensor, our study reveals major differences in G protein responses and agonist potencies between neurons and heterologous HEK293 cells. The difference in the profiles of G protein response using different Gγ subunits (Gγ2, Gγ8 or Gγ9) (Fig. 7) shows clearly the impact of Gγ in CGNs. It shows that not only the Gα subunit is important, but also the Gβ and Gγ, consistent with other recent studies65,66. Therefore, it brings to our attention that the G protein-mediated response also depends detected on the biosensor components used, which is the limitation of biosensors that only detect the response of a well-defined G protein. Meanwhile, the difference can be due to the intrinsic properties of the system under study. For example, the association of these receptors with specific protein partners that may influence their coupling to some Gαi subtypes, present in neurons but not in HEK293 cells. Indeed, specific proteins interacting with the GABAB receptor were identified in brain tissues15,69, while they are not expressed in HEK293 cells. For example, the soluble form of APP, AJAP-1 and PIANP bind to the extracellular sushi domain of GABAB70,71, TRPV1 channels interact in the membrane72, and the intracellular KCTDs and 14-3-3 proteins73 modulate GABAB receptor downstream signalling. The KCTD proteins are constitutively associated with GABAB and they control its kinetics of activation73.
Our sensors showed a difference between Gi and Go proteins revealed by a CB1 agonist CP 55940, which has a higher potency in generating Gi1 than GoA responses in neurons, relative to the other agonists tested. Indeed, such biased signalling between Gi and Go proteins have already been reported upon agonist stimulation12,13,74, by allosteric modulators49 or as a result of genetic variation75. In our study, the molecular bases of this difference in agonist effect is unclear. CP 55940 and Win 55212-2 (Ki 1.1 nM and 62.3 nM for CB1 receptor, respectively)76,77 have different scaffolds. Their differential effects might be explained by the ligand-binding kinetics78, by specific active conformation stabilized by the agonist79, or due to specific components associated to the receptor in these neurons, better stabilizing the CP 55940/CB1/Gi complex.
Our sensors also revealed larger differences in some agonist potencies in neurons compared to transfected HEK293 cells. This is nicely illustrated for the CB1 and GABAB agonists CP 55940 and APPA, respectively, that show a higher potency than the other agonists in neurons (Fig. 8a). In addition and unexpectedly, no intrinsic agonist activity of the PAM Rac BHFF was detected with the native GABAB receptor in contrast to the recombinant one in HEK293 cells. Finally, the important differences in GABAB receptor agonist potencies that are higher in transfected HEK293 cells compared to neurons are most probably not due to the higher expression in the transfected cells in our study. Indeed, a 10-times higher affinity for GABA in neurons compared to HEK cells80 and similar agonist potencies in GTPγS binding experiments in the two systems81 were previously reported. Possible explanations might come from the environment of the receptor in neurons, such as its compartmentalization and its interaction with extracellular and intracellular proteins15,50, including other endogenous receptors.
Our sensors also revealed specific ligand-independent G protein response in neurons compared to HEK293 cells. Indeed, no detectable constitutive activity of the GABAB receptor was observed in CGNs in contrast to transfected cell lines49,51,52. As discussed above, this lack of constitutive activity might be explained by the environment of the receptor in neurons. The importance of the intracellular protein partners for the constitutive activity of a GPCR was nicely illustrated for the postsynaptic metabotropic glutamate receptor mGlu5. Its constitutive activity was maintained low in neurons by the long isoform of Homer, while it is revealed when the short isoform of Homer is expressed82.
In conclusion, our study reveals the importance of evaluating the pharmacological and G protein response properties of native GPCRs in primary cells to better understand their signalling, identify and characterize ligands expected to have therapeutic effects. It also reveals the powerfulness of using G protein sensors compatible with native cells, though biosensors compatible with single cell analysis and imaging will certainly help revealing the potential cell-to-cell or sub-compartment heterogeneity in GPCR and G protein coupling83,84. In the future, methods measuring the activation of a specific endogenous G protein in live cells should be envisioned. Regardless of the criticism of our approach, our study clearly reveals that data generated in recombinant systems should be taken with caution before going further into pre-clinical and clinical development of drugs candidates only characterized this way.
Methods
Materials
GABA (Cat. A2129), CP 55940 (Cat. C1112) and carbachol (PHR1511) were purchased from Sigma-Aldrich (Shanghai, China). R-baclofen (Cat. 0796), SKF 97581 (Cat. 0379) and CGP54626 (Cat. 1088) were obtained from Tocris Biosciences (Shanghai, China). 3-APPA (Cat. ab120329) was purchased from Abcam (Shanghai, China). Win 55,212-2 (T4458), Bay 59-3074 (T3699), DAMGO (T4351), eletriptan HBr (T0216), pramipexole 2HCl monohydrate (T6951), brimonidine tartrate (T6422), tizanidine hydrochloride (T0290), oxymetazoline hydrochloride (T0252), xylazine (T7046), clonidine hydrochloride (T1247), and 1-oleoyl lysophosphatidic acid sodium (T21654) were purchased from TargetMol (Shanghai, China). PAF C-16 (74389-68-7) was purchased from Santa Cruz Biotechnology (Shanghai, China). CGP64213 was a gift from Prof. Fajun Nan (Shanghai Institute of Materials Medicine, China). Coelenterazine h (Cat. S2001) and furimazine (Cat. N1120) were purchased from Promega (Madison, WI, USA).
Plasmids
The coding sequences of Nluc and Rluc8 were PCR-amplified and inserted into the coding sequence of human Gαi1, Gαi2, Gαi3, GαoA and GαoB (between residues 91 and 92) and Gαz (between residues 113 and 114) in pcDNA3.1, using the flexible linkers SGGGGS and SGGGEF, before and after the sequence of Nluc, as shown in Supplementary Figs. 10c, d and 12. The pRK5 plasmid encoding the rat GABAB subunits (HA-tagged GABAB1 (GB1) and Flag-tagged GABAB2 (GB2)) and pcDNA3.1 encoding the human Gβ1 and human VenusGγ2 were provided by the Institut de Génomique Fonctionnelle (Montpellier, France). The cDNA of mouse α2AAR, mouse CB1, human Gγ8 and human Gγ9 were bought from Miaoling Bio (Wuhan, China) and inserted in pcDNA3.1. Venus was inserted at the N-terminus of Gγ2, Gγ8 and Gγ9 between the HindIII and KpnI restriction enzymatic sites and the linker KLGT between Venus and the Gγ subunits was used. The schematic representation of the constructs (Supplementary Fig. 10) and their protein sequences (Supplementary Figs. 11–13) were shown.
HEK293 cell culture and transfection
HEK293 cells (ATCC, CRL-1573, lot: 3449904) were cultured in DMEM supplemented with 10% FBS at 37 °C in a humidified incubator containing 5% CO2. For transfections, cells were suspended and transfected using Lipofectamine 2000 with the appropriate expression constructs as previously described49. The cDNA amounts used per well in 96-well plate were as following unless indicated in figure legends: rat GABAB receptor, GB1 (3 ng, 6 ng, 9 ng, 30 ng) and GB2 (4 ng, 8 ng, 12 ng, 40 ng), mouse α2AAR (10 ng, 15 ng, 20 ng, 50 ng), mouse CB1 receptor (80 ng), Gαi1Nluc (1 ng), Gαi2Nluc (3 ng), Gαi3Nluc (3 ng), GαoANluc (1 ng), GαoBNluc (1 ng), GαzNluc (1.5 ng), Gβ1 (10 ng) and VenusGγ2 (10 ng). The ratio of DNA to Lipofectamine 2000 ratio was 1:2. After a 24 h culture in 96-well plates, the cells were ready for the bioluminescence resonance energy transfer (BRET) experiments.
Primary neuron culture and transfection
For the primary culture of neurons, all experiments were specifically designed to minimize the number of animals used and were approved by the Animal Experimentation Ethics Committee of the College of Life Science and Technology, Huazhong University of Science and Technology, Wuhan, China. Kunming mice were obtained from the Center for Disease Control and Prevention of Hubei Province. The mice were raised in a specific pathogen free (SPF) environment with an ambient temperature of 18–22 °C, a humidity of 50%-60%, and a 12 h light-dark cycle.
Primary cerebellar granule neuronal cultures were prepared from one-week-old newborn Kunming mice as previously described39. The dissected tissue was gently triturated after Versene (15040066; Gibco, Shanghai, China) treatment for 5 min at 37 °C, and the homogenate was centrifuged at 170 g for 5 min. In the meantime, mixtures of the DNA and Lipofectamine 2000 (Ref 11668019; Thermo Fisher Scientific, Shanghai, China) in a 1:3 ratio in an Opti-minimal essential medium (Opti-MEM) (Ref 31985070; Thermo Fisher Scientific, Shanghai, China) were prepared following the manufacturer’s protocols and as previously described42. The pellet was re-suspended in culture medium DMEM-F12 (Ref 11320-033; Gibco) supplemented with 2 mM glutamine, 30 mM KCl, 100 U/mL penicillin, 100 μg/mL streptomycin, and 10% foetal bovine serum (FBS) and seeded into 96-well plates (100 μL/well) previously coated with poly-L-ornithine (Sigma-Aldrich). Then, the mixtures of DNA and Lipofectamine 2000 (50 μL/well) were added to the wells. BRET experiments were performed three or four days after transfection. BRET experiments were performed at DIV3 or DIV4. The cDNA amounts used per well in 96-well plate were as following: Gαi1Nluc (25 ng), Gαi2Nluc (75 ng), Gαi3Nluc (75 ng), GαoANluc (25 ng), GαoBNluc (25 ng), GαzNluc (25 ng), VenusGγ2 (50 ng), VenusGγ8 (50 ng) and VenusGγ9 (50 ng).
Primary cortical and hippocampal neuronal cultures were prepared from embryonic day 17.5 mice as previously described85. The cortex or hippocampi were digested with trypsin, and cells were seeded in neurobasal medium (Ref 21103049; Gibco) supplemented with 2% B27 (Ref 17504-044; Gibco), 4 mM GlutaMAX, 25 μM glutamic acid, 100 U/mL penicillin, 100 μg/mL streptomycin and 10% FBS in 96-well plates. After three days in culture (DIV3), the culture medium was supplemented with 5 μM cytosine β-D-arabinofuranoside hydrochloride (C6645; Sigma-Aldrich) and incubated overnight. Then, 75% of the medium was replaced by neurobasal medium supplemented with B27, GlutaMAX and antibiotics. Neurons were then transfected with Gαi1Nluc or GαoANluc (50 ng/well), together with VenusGγ2 (50 ng/well) using Lipofectamine 2000 on DIV7. The ratio of DNA to Lipofectamine 2000 ratio was 1:2. BRET experiments were performed on DIV10 or DIV11.
Fluorescent imaging
Cerebellar granule neurons (CGNs) and transfected HEK293 cells were fixed with 4% formaldehyde and blocked with 2% bovine serum albumin (BSA) and 0.1% Triton X-100 in phosphate-buffered saline (PBS). The CGNs were incubated with primary GFP antibody (1:200; ab1218, Abcam, Shanghai, China) at 4 °C overnight. After washing three times with PBS, the cells were incubated with secondary anti-mouse antibody Alexa Fluor® 488 AffiniPure Donkey Anti-Mouse IgG (H + L) (1:500, 715-545-150, Jackson ImmunoResearch, Shanghai, China) at 25 °C for 2 h. The CGNs were then stained with DAPI for 15 min. The cells were washed with PBS and mounted with FluorSave reagent (AR1109, Boster Biological Technology Co. Ltd., Wuhan, China). Images were obtained with an Olympus FV1000 laser scanning confocal microscope (60 x objective for HEK293 cells and 40x objective for CGNs, Olympus Corporation, Tokyo, Japan) equipped with appropriate fluorescence and filters (FITC: 488/530 nm; DAPI: 405/449 nm). The images were digitized and saved in TIFF format.
BRET measurement
CGNs were starved in HEPES-buffered saline (HBS) containing 10 mM HEPES pH 7.4, 140 mM NaCl, 4 mM KCl, 2 mM MgSO4, and 1 mM KH2PO4. Cortical and hippocampal neurons were starved with artificial cerebrospinal fluid (aCSF) buffer containing 140 mM NaCl, 2 mM CaCl2, 3 mM KCl, 10 mM HEPES, and 10 mM D-glucose, at 37 °C for 1 h before the BRET experiments. BRET measurements were performed as previously described using the Mithras LB 940 multimode microplate reader (Berthold Technologies, Bad Wildbad, Germany)49 with the programme MikroWin (Version 4.41) or PHERAstar FS (BMG Labtech, USA)86 with the programme PHERAstar control (Version 4.00 R4). The signals emitted by the donor (460–500 nm band-pass filter, Em 480 nm) and the acceptor (510–550 nm band-pass filter, Em 530 nm) were recorded by Mithras LB 940 after the addition of 10 μM furimazine. All measurements were performed at 37 °C. The BRET signal was determined by calculating the ratio between the emission of acceptor and donor (Em 530 nm / Em 480 nm). The basal BRET ratio (BRETbasal) of cells was recorded before the stimulation with drugs or buffer. The change in BRET ratio (net BRET) was obtained by subtracting the BRET ratio after agonist treatment from the basal BRET (BRETbasal-BRETagonist). Agonist-induced change in BRET ratio for the different Gi/o sensors was expressed as percentage of the basal signal ((BRETbasal-BRETagonist/BRETbasal) x100). To study the kinetics, the BRET was measured in real time with a counting time of 0.5 s, and the drugs were injected using the Mithras LB 940 injectors at the indicated time.
Quantitative reverse transcription PCR
Total RNA was extracted using standard methods (Trizol, Invitrogen) from CGNs and HEK293 cells. Quantitative reverse transcription PCR (qRT-PCR) was carried out with SYBR Green (Vazyme Biotechnology, Nanjing, China) according to the manufacturer’s protocol as reported87. Rplp0 was used as an internal reference for normalization, and the ΔΔCt method was adopted to analyse quantitative PCR data. The primer sets used were referred to previous references75,88. For mouse (CGNs), Gnai1 (NM_010305), Forward (Fw): 5’-AAGCTGACTCGCCTTCCCAG-3’, Reverse (Rv): 5’-GTAGTTTACAGTTCTCCACACG-3’; Gnai2 (NM_008138), Fw: 5’-TGCCTTGAGTGTGTCTGCGTG-3’, Rv: 5’-CTCAGTGACGTTGGCAGTTG-3’; Gnai3 (NM_010306), Fw: 5’-GTGCAGTCCGTGTACAAGAG-3’, Rv: 5’-GATGAATGGATCCGAGCCAC-3’; Gnao1 transcript isoform A (NM_010308) Fw: 5’-AGGAAGACGGACTCCAAGATG-3’, Rv: 5’-AGTCGAAGAGCATGAGAGAC-3’; Gnao1 transcript isoform B (NM_001113384) Fw: 5’-AGGAAGACGGACTCCAAGATG-3’, Rv: 5’-AGATGTGTCTGTGAACCACTTG-3’; Gnaz (NM_010311), Fw: 5’-CAGCCGTGCTTAGAAACATCG-3’, Rv: 5’-TCTAGTGACACTCCACCTCC-3’ and Rplp0 (NM_007475), Fw: 5’-CTCACTGAGATTCGGGATATG-3’, Rv: 5’-CTCCCACCTTGTCTCCAGTC-3’. For human (HEK293 cells), GNAI1 (NM_002069), Fw: 5’-CATCTCTGACCTTGTTTCAGC-3’, Rv: 5’-CTTCAACCCAGTGACAACACG-3’; GNAI2 (NM_002070), Fw: 5’-ACTCCGTGCCTTGAGTGTG-3’, Rv:5’-TTGTCTGGAACAGCCCTTGG-3’; GNAI3 (NM_010306), Fw: 5’-GGAAAGTTACGTTCACTTCAACC-3’, Rv: 5’-TTGGACCCCAAAAGGCACTG-3’; GNAO1 transcript isoform A (NM_020988) Fw: 5’-AGAAAGGCTGACGCCAAGAT-3’, Rv: 5’-AGTCGAAGAGCATGAGAGAC-3’; GNAO1 transcript isoform B (NM_138736) Fw: 5’-AGAAAGGCTGACGCCAAGAT-3’, Rv: 5’-TGGACGTGTCTGTGAACCAT-3’; GNAZ (NM_002073), Fw: 5’-CTACGAGGATAACCAGAC-3’, Rv: 5’-TACGTGTTCTGGCCCTTG-3’ and RPLP0 (NM_053275), Fw: 5’-ATGCAGCAGATCCGCATGT-3’, Rv: 5’-TTGCGCATCATGGTGTTCTT-3’.
Cell membrane preparations
The cell membranes were prepared as reported89. HEK293 cells or CGNs were washed three times with PBS, then scraped and collected by centrifugation at 170 g for 5 min. Cells were resuspended in 500 μl Tris buffer (50 mM Tris pH 7.4, 50 mM NaCl) with cOmplete protease inhibitor cocktail (Roche) and crushed through a 26 gauge 5/8 inch needle attached to a syringe for 30 passages. After centrifugation at 860 g at 4 °C for 5 min, liquid supernatants were transferred to high speed centrifuge tubes and centrifuged at 44000 g at 4 °C for 20 min by Optima MAX-TL ultracentrifuge (Beckman Coulter, Brea, CA, USA). The precipitated membranes were diluted gently in 50 μl Tris-NaCl buffer. The protein amount was further determined using BCA protein assay kit and 20 μg protein was loaded for western blotting detection.
Western blotting analysis
Cells were treated using RIPA lysis buffer (50 mM Tris pH 7.4, 150 mM NaCl, 1% NP-40, 0.5% sodium deoxycholate, 0.1% SDS, sodium orthovanadate, sodium fluoride, EDTA, leupeptin; Beyotime Bio., Cat. P0013C, China) and protein concentrations were determined using the BCA protein assay kit. Equal amounts of protein (20 μg) from total cell lysis or cell membranes were separated by SDS–polyacrylamide gel electrophoresis (PAGE) on 10 to 12% gels. Proteins were transferred to nitrocellulose membranes and washed in blocking buffer (5% nonfat dry milk in Tris-buffered saline and 0.1% Tween 20) for 2 h at 25 °C. The blots were incubated with the primary monoclonal antibodies anti-GB1 mAb (1:1000, ab55051, Abcam, Shanghai, China), anti-Gβ2 rabbit (1:1000, A9643, ABclonal Technology, Wuhan, China), anti-β-actin (1:3000, KM9001T, Sungene Biotech., Tianjin, China), the rabbit polyclonal antibodies anti-CB1 (1:1000, A1447, ABclonal, Wuhan, China), anti-α2AAR (1:1000, A2809, ABclonal, Wuhan, China), anti-Gβ1 (1:1000, A1867, ABclonal, Wuhan, China), anti-Gβ3 (1:1000, A1387, ABclonal, Wuhan, China) and anti-Gβ5 (1:1000, A4447, ABclonal, Wuhan, China). The primary antibodies were at the relevant dilution overnight at 4 °C and then incubated with DyLight 800 4 X PEG conjugated secondary antibodies (1:20,000, anti-mouse IgG, #5257; 1:20,000, anti-rabbit IgG, #5151, Cell Signaling Technology, Shanghai, China) for 2 h at 25 °C. Membranes were imaged using an Odyssey infrared scanner (LI-COR Biosciences, Lincoln, NE, USA) at 700 nm.
Statistical analysis
Results are presented as the mean ± SEM of at least three independent experiments. Statistical analysis was performed using GraphPad Prism 9.5.1 software (GraphPad Software Inc., San Diego, CA, USA). Dose-response experiments were analysed using nonlinear curve fitting for the log (agonist) vs. response (three parameters) curves. Statistical analysis was performed using the Ordinary one-way ANOVA with a Dunnett’s post-hoc multiple comparison test or unpaired t-test (two-tailed) or paired t-test. P < 0.05 was considered to be statistically significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data generated in this study are provided in the main text, Supplementary information and source data files. The raw data and p-values for all Figures and Supplementary Figs. are available in Source Data file accompanying this paper. Source data are provided with this paper.
References
Hauser, A. S., Attwood, M. M., Rask-Andersen, M., Schioth, H. B. & Gloriam, D. E. Trends in GPCR drug discovery: new agents, targets and indications. Nat. Rev. Drug Discov. 16, 829–842 (2017).
Wettschureck, N. & Offermanns, S. Mammalian G proteins and their cell type specific functions. Physiol. Rev. 85, 1159–1204 (2005).
Heng, B. C., Aubel, D. & Fussenegger, M. An overview of the diverse roles of G-protein coupled receptors (GPCRs) in the pathophysiology of various human diseases. Biotechnol. Adv. 31, 1676–1694 (2013).
Sriram, K. & Insel, P. A. G protein-coupled receptors as targets for approved drugs: how many targets and how many drugs? Mol. Pharmacol. 93, 251–258 (2018).
Inoue, A. et al. Illuminating G-protein-coupling selectivity of GPCRs. Cell 177, 1933–1947 e1925 (2019).
Hauser, A. S. et al. Common coupling map advances GPCR-G protein selectivity. eLife 11, e74107 (2022).
Avet, C. et al. Effector membrane translocation biosensors reveal G protein and beta arrestin coupling profiles of 100 therapeutically relevant GPCRs. eLife 11, e74101 (2022).
Okashah, N. et al. Variable G protein determinants of GPCR coupling selectivity. Proc. Natl Acad. Sci. USA 116, 12054–12059 (2019).
Kenakin, T. Biased receptor signaling in drug discovery. Pharmacol. Rev. 71, 267–315 (2019).
Galandrin, S., Oligny-Longpre, G. & Bouvier, M. The evasive nature of drug efficacy: implications for drug discovery. Trends Pharmacol. Sci. 28, 423–430 (2007).
Kolb, P. et al. Community guidelines for GPCR ligand bias: IUPHAR review 32. Br. J. Pharmacol. 179, 3651–3674 (2022).
Busnelli, M. et al. Functional selective oxytocin-derived agonists discriminate between individual G protein family subtypes. J. Biol. Chem. 287, 3617–3629 (2012).
Masuho, I. et al. Distinct profiles of functional discrimination among G proteins determine the actions of G protein-coupled receptors. Sci. Signal. 8, ra123 (2015).
Tran, K. et al. Size-reduced macrocyclic analogues of [Pyr(1)]-apelin-13 showing negative Galpha12 bias still produce prolonged cardiac effects. J. Med. Chem. 65, 531–551 (2022).
Rosenbaum, M. I., Clemmensen, L. S., Bredt, D. S., Bettler, B. & Stromgaard, K. Targeting receptor complexes: a new dimension in drug discovery. Nat. Rev. Drug Discov. 19, 884–901 (2020).
Wingler, L. M. & Lefkowitz, R. J. Conformational basis of G protein-coupled receptor signaling versatility. Trends Cell. Biol. 30, 736–747 (2020).
Zhang, R. & Xie, X. Tools for GPCR drug discovery. Acta Pharmacol. Sin. 33, 372–384 (2012).
Kenakin, T. Drug efficacy at G protein-coupled receptors. Annu. Rev. Pharmacol. Toxicol. 42, 349–379 (2002).
Milligan, G. Principles: extending the utility of [35S]GTP gamma S binding assays. Trends Pharmacol. Sci. 24, 87–90 (2003).
Strange, P. G. Use of the GTPgammaS ([35S]GTPgammaS and Eu-GTPgammaS) binding assay for analysis of ligand potency and efficacy at G protein-coupled receptors. Br. J. Pharmacol. 161, 1238–1249 (2010).
Mannoury la Cour, C., Herbelles, C., Pasteau, V., de Nanteuil, G. & Millan, M. J. Influence of positive allosteric modulators on GABA(B) receptor coupling in rat brain: a scintillation proximity assay characterisation of G protein subtypes. J. Neurochem. 105, 308–323 (2008).
Maziarz, M. et al. Revealing the activity of trimeric G-proteins in live cells with a versatile biosensor design. Cell 182, 770–785 e716 (2020).
Bunemann, M., Frank, M. & Lohse, M. J. Gi protein activation in intact cells involves subunit rearrangement rather than dissociation. Proc. Natl Acad. Sci. USA 100, 16077–16082 (2003).
Gales, C. et al. Probing the activation-promoted structural rearrangements in preassembled receptor-G protein complexes. Nat. Struct. Mol. Biol. 13, 778–786 (2006).
Olsen, R. H. J. et al. TRUPATH, an open-source biosensor platform for interrogating the GPCR transducerome. Nat. Chem. Biol. 16, 841–849 (2020).
Wright, S. C. & Bouvier, M. Illuminating the complexity of GPCR pathway selectivity - advances in biosensor development. Cur. Opin. Struct. Biol. 69, 142–149 (2021).
Hyman, S. E. Neurotransmitters. Cur. Biol. 15, R154–R158 (2005).
Robishaw, J. D. & Berlot, C. H. Translating G protein subunit diversity into functional specificity. Cur. Opin. Cell. Biol. 16, 206–209 (2004).
Busnelli, M., Peverelli, E., Mantovani, G., Spada, A. & Chini, B. Deciphering the specific role of G(alphai/o) isoforms: functional selective oxytocin ligands and somatostatin SST5 receptor mutants. Biochem. Soc. Trans. 41, 166–171 (2013).
Gassmann, M. & Bettler, B. Regulation of neuronal GABA(B) receptor functions by subunit composition. Nat. Rev. Neurosci. 13, 380–394 (2012).
Saunders, C. & Limbird, L. E. Localization and trafficking of alpha2-adrenergic receptor subtypes in cells and tissues. Pharmacol. Ther. 84, 193–205 (1999).
Langer, S. Z. alpha2-Adrenoceptors in the treatment of major neuropsychiatric disorders. Trends Pharmacol. Sci. 36, 196–202 (2015).
Howlett, A. C. et al. International Union of Pharmacology. XXVII. Classification of cannabinoid receptors. Pharmacol. Rev. 54, 161–202 (2002).
Contestabile, A. Cerebellar granule cells as a model to study mechanisms of neuronal apoptosis or survival in vivo and in vitro. Cerebellum 1, 41–55 (2002).
England, C. G., Ehlerding, E. B. & Cai, W. NanoLuc: A small luciferase is brightening up the field of bioluminescence. Bioconjug. Chem. 27, 1175–1187 (2016).
Schihada, H., Shekhani, R. & Schulte, G. Quantitative assessment of constitutive G protein-coupled receptor activity with BRET-based G protein biosensors. Sci. Signal. 14, eabf1653 (2021).
Jiang, M. & Bajpayee, N. S. Molecular mechanisms of Go signaling. Neurosignals 17, 23–41 (2009).
Bettler, B., Kaupmann, K., Mosbacher, J. & Gassmann, M. Molecular structure and physiological functions of GABA(B) receptors. Physiol. Rev. 84, 835–867 (2004).
Tu, H. et al. GABAB receptor activation protects neurons from apoptosis via IGF-1 receptor transactivation. J. Neurosci. 30, 749–759 (2010).
Ramachandran, S. & Cerione, R. A. A dominant-negative Galpha mutant that traps a stable rhodopsin-Galpha-GTP-betagamma complex. J. Biol. Chem. 286, 12702–12711 (2011).
Liu, P. et al. The structural basis of the dominant negative phenotype of the Galphai1beta1gamma2 G203A/A326S heterotrimer. Acta Pharmacol. Sin. 37, 1259–1272 (2016).
Ango, F. et al. A simple method to transfer plasmid DNA into neuronal primary cultures: functional expression of the mGlu5 receptor in cerebellar granule cells. Neuropharmacology 38, 793–803 (1999).
Urwyler, S. Allosteric modulation of family C G-protein-coupled receptors: from molecular insights to therapeutic perspectives. Pharmacol. Rev. 63, 59–126 (2011).
Filip, M. et al. GABAB receptors as a therapeutic strategy in substance use disorders: focus on positive allosteric modulators. Neuropharmacology 88, 36–47 (2015).
Pin, J. P. & Prezeau, L. Allosteric modulators of GABA(B) receptors: mechanism of action and therapeutic perspective. Cur. Neuropharmacol. 5, 195–201 (2007).
Mao, C. et al. Cryo-EM structures of inactive and active GABAB receptor. Cell. Res. 30, 564–573 (2020).
Shen, C. et al. Structural basis of GABAB receptor-Gi protein coupling. Nature 594, 594–598 (2021).
Liu, L. et al. Allosteric ligands control the activation of a class C GPCR heterodimer by acting at the transmembrane interface. eLife 10, e70188 (2021).
Lecat-Guillet, N. et al. FRET-based sensors unravel activation and allosteric modulation of the GABAB receptor. Cell. Chem. Biol. 24, 360–370 (2017).
Berg, K. A. & Clarke, W. P. Making sense of pharmacology: inverse agonism and functional Selectivity. Int. J. Neuropsychopharmacol. 21, 962–977 (2018).
Grunewald, S. et al. Importance of the gamma-aminobutyric acid(B) receptor C-termini for G-protein coupling. Mol. Pharmacol. 61, 1070–1080 (2002).
Rajalu, M. et al. Pharmacological characterization of GABAB receptor subtypes assembled with auxiliary KCTD subunits. Neuropharmacology 88, 145–154 (2015).
Shaye, H. et al. Structural basis of the activation of a metabotropic GABA receptor. Nature 584, 298–303 (2020).
Papasergi-Scott, M. M. et al. Structures of metabotropic GABAB receptor. Nature 584, 310–314 (2020).
Park, J. et al. Structure of human GABAB receptor in an inactive state. Nature 584, 304–309 (2020).
Vuillaume, M. L. et al. A novel mutation in the transmembrane 6 domain of GABBR2 leads to a Rett-like phenotype. Ann. Neurol. 83, 437–439 (2018).
Onaivi, E. S. et al. Discovery of the presence and functional expression of cannabinoid CB2 receptors in brain. Ann. N. Y. Acad. Sci. 1074, 514–536 (2006).
Burke, J. & Schwartz, M. Preclinical evaluation of brimonidine. Surv. Ophthalmol. 41, S9–S18 (1996).
Mierau, J. et al. Pramipexole binding and activation of cloned and expressed dopamine D2, D3 and D4 receptors. Eur. J. Pharmacol. 290, 29–36 (1995).
Goadsby, P. J. & Hoskin, K. L. Differential effects of low dose CP122,288 and eletriptan on Fos expression due to stimulation of the superior sagittal sinus in cat. Pain 82, 15–22 (1999).
Manuel, I., Lombardero, L., LaFerla, F. M., Giménez-Llort, L. & Rodríguez-Puertas, R. Activity of muscarinic, galanin and cannabinoid receptors in the prodromal and advanced stages in the triple transgenic mice model of Alzheimer’s disease. Neuroscience 329, 284–293 (2016).
Maurel, B., Le Digarcher, A., Dantec, C. & Journot, L. Genome-wide profiling of G protein-coupled receptors in cerebellar granule neurons using high-throughput, real-time PCR. BMC Genomics. 12, 241 (2011).
Jasper, J. R. et al. Ligand efficacy and potency at recombinant alpha2 adrenergic receptors: agonist-mediated [35S]GTPgammaS binding. Biochem. Pharmacol. 55, 1035–1043 (1998).
Robishaw, J. D. Preferential assembly of G-alphabetagamma complexes directed by the gamma subunits. Subcell. Biochem. 63, 181–191 (2012).
Masuho, I., Skamangas, N. K., Muntean, B. S. & Martemyanov, K. A. Diversity of the Gbetagamma complexes defines spatial and temporal bias of GPCR signaling. Cell Syst. 12, 324–337 e325 (2021).
Kankanamge, D., Tennakoon, M., Karunarathne, A. & Gautam, N. G protein gamma subunit, a hidden master regulator of GPCR signaling. J. Biol. Chem. 298, 102618 (2022).
Frank, M., Thumer, L., Lohse, M. J. & Bunemann, M. G Protein activation without subunit dissociation depends on a Galpha(i)-specific region. J. Biol. Chem. 280, 24584–24590 (2005).
Digby, G. J., Lober, R. M., Sethi, P. R. & Lambert, N. A. Some G protein heterotrimers physically dissociate in living cells. Proc. Natl Acad. Sci. USA 103, 17789–17794 (2006).
Pin, J. P. & Bettler, B. Organization and functions of mGlu and GABAB receptor complexes. Nature 540, 60–68 (2016).
Dinamarca, M. C. et al. Complex formation of APP with GABAB receptors links axonal trafficking to amyloidogenic processing. Nat. Commun. 10, 1331 (2019).
Rice, H. C. et al. Secreted amyloid-beta precursor protein functions as a GABABR1a ligand to modulate synaptic transmission. Science 363, eaao4827 (2019).
Hanack, C. et al. GABA blocks pathological but not acute TRPV1 pain signals. Cell 160, 759–770 (2015).
Schwenk, J. et al. Native GABA(B) receptors are heteromultimers with a family of auxiliary subunits. Nature 465, 231–235 (2010).
Von Moo, E. et al. Ligand-directed bias of G protein signaling at the dopamine D2 receptor. Cell. Chem. Biol. 29, 226–238 e224 (2022).
Peverelli, E. et al. Specific roles of G(i) protein family members revealed by dissecting SST5 coupling in human pituitary cells. J. Cell. Sci. 126, 638–644 (2013).
Melvin, L. S. et al. Structure-activity relationships for cannabinoid receptor-binding and analgesic activity: studies of bicyclic cannabinoid analogs. Mol. Pharmacol. 44, 1008–1015 (1993).
Felder, C. C. et al. Comparison of the pharmacology and signal transduction of the human cannabinoid CB1 and CB2 receptors. Mol. Pharmacol. 48, 443–450 (1995).
Klein Herenbrink, C. et al. The role of kinetic context in apparent biased agonism at GPCRs. Nat. Commun. 7, 10842 (2016).
Suomivuori, C. M. et al. Molecular mechanism of biased signaling in a prototypical G protein-coupled receptor. Science 367, 881–887 (2020).
Kaupmann, K. et al. Expression cloning of GABA(B) receptors uncovers similarity to metabotropic glutamate receptors. Nature 386, 239–246 (1997).
Galvez, T. et al. Ca(2+) requirement for high-affinity gamma-aminobutyric acid (GABA) binding at GABA(B) receptors: involvement of serine 269 of the GABA(B)R1 subunit. Mol. Pharmacol. 57, 419–426 (2000).
Bockaert, J., Perroy, J. & Ango, F. The complex formed by group I metabotropic Glutamate Receptor (mGluR) and Homer1a plays a central role in metaplasticity and homeostatic synaptic scaling. J. Neurosci. 41, 5567–5578 (2021).
Chavez-Abiega, S., Gronloh, M. L. B., Gadella, T. W. J., Bruggeman, F. J. & Goedhart, J. Single cell imaging of ERK and Akt activation dynamics and heterogeneity induced by G protein-coupled receptors. J. Cell. Sci. 135, jcs259685 (2022).
Sebastianutto, I. et al. D1-mGlu5 heteromers mediate noncanonical dopamine signaling in Parkinson’s disease. J. Clin. Invest. 130, 1168–1184 (2020).
Goyet, E., Bouquier, N., Ollendorff, V. & Perroy, J. Fast and high resolution single-cell BRET imaging. Sci. Rep. 6, 28231 (2016).
Hauser, A. S. et al. Pharmacogenomics of GPCR Drug Targets. Cell 172, 41–54 e19 (2018).
Zhang, B. et al. Environmental temperature differentially modulates C. elegans longevity through a thermosensitive TRP channel. Cell. Rep. 11, 1414–1424 (2015).
Doi, M. et al. Gpr176 is a Gz-linked orphan G-protein-coupled receptor that sets the pace of circadian behaviour. Nat. Commun. 7, 10583 (2016).
Liu, H. et al. Illuminating the allosteric modulation of the calcium-sensing receptor. Proc. Natl Acad. Sci. USA 117, 21711–21722 (2020).
Acknowledgements
We thank Dr Julie Perroy (IGF) for providing us Nluc cDNA. This work was supported by grants from the National Natural Science Foundation of China (NSFC) (grant number 32271198 to C.X.), the National Key R&D Programme of China (grant number 2022YFA1302901 to C.X.), the National Natural Science Foundation of China (NSFC) (grant number 32330049 and 82320108021 to J.L.), National Key R&D Programme of China (grant number 2022YFE0116600 and 2021ZD0203302 to J.L.), interdisciplinary Research Programme of HUST (grant number 2023JCYJ006 to C.X.), the Agence Nationale de la Recherche (ANR 18-CE11-0004-01 to J.-P.P.), and the Fondation Recherche Médicale (DEQ 20170336747 to J.-P.P. and EQU202303016470 to P.R.). P.R. and J.-P.P. were supported by the Institut National de la Santé et de la Recherche Médicale (INSERM; International Research Programme «Brain Signal») and the Franco-Chinese Joint Scientific and Technological Commission (CoMix) from the French Embassy in China.
Author information
Authors and Affiliations
Contributions
C.X., J.L., P.R. and J.-P.P. designed the experiments. C.X. initiated the project, designed the sensors, set up the protocols and screened different agonists; C.X., Y.Z. and Y.L. detected the Gi/o protein response in HEK293 cells and CGNs; Y.L. optimized the transfection amount in HEK293 cells and prepared cell membranes; Y.L. and Y.Z. performed all the western blotting experiments; L.L. performed the dose-response in HEK293 cells treated with GABAB receptor agonists; Y.L., L.L. and Z.X. produced constructs. P.L. and X.W. helped with the plasmid constructs and neuron preparations. C.X., Y.Z. and Y.L. collected and analysed the data. C.X., J.L., J.-P.P. and P.R. wrote the manuscript with input from all of the authors.
Corresponding authors
Ethics declarations
Competing interests
P.R. and J.-P.P. are involved in a collaborative team between the CNRS and Cisbio Bioassays, Revvity group. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Moritz Bünemann, Laura Humphrys, Bryan Roth and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Xu, C., Zhou, Y., Liu, Y. et al. Specific pharmacological and Gi/o protein responses of some native GPCRs in neurons. Nat Commun 15, 1990 (2024). https://doi.org/10.1038/s41467-024-46177-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-024-46177-z
This article is cited by
-
Absence of calcium-sensing receptor basal activity due to inter-subunit disulfide bridges
Communications Biology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.