Featured
-
-
Article
| Open Access Neutron-encoded diubiquitins to profile linkage selectivity of deubiquitinating enzymes
Neutron-encoded diubiquitins to profile linkage selectivity of deubiquitinating enzymes
Most insights into deubiquitinase (DUB) substrate specificity originate from studies with isolated di-ubiquitins (diUb), but in cells diUbs with different linkage types coexist. Here, the authors develop a mass spectrometric DUB activity assay that can probe all diUbs simultaneously under substrate competition conditions.
- Bianca D. M. van Tol
- , Bjorn R. van Doodewaerd
- & Paul P. Geurink
-
Article
| Open Access Electrostatic and steric effects underlie acetylation-induced changes in ubiquitin structure and function
Electrostatic and steric effects underlie acetylation-induced changes in ubiquitin structure and function
Ubiquitin is not only a posttranslational modifier but itself is subject to modifications, such as acetylation. Characterization of distinct acetylated ubiquitin variants reveals that each acetylation site has a particular impact on ubiquitin structure and its protein-protein interaction properties.
- Simon Maria Kienle
- , Tobias Schneider
- & Martin Scheffner
-
Article
| Open Access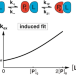 A litmus test for classifying recognition mechanisms of transiently binding proteins
A litmus test for classifying recognition mechanisms of transiently binding proteins
The authors provide a litmus test for the recognition mechanism of transiently binding proteins based on nuclear magnetic resonance and find a conformational selection binding mechanism through concentration-dependent kinetics of ubiquitin and SH3.
- Kalyan S. Chakrabarti
- , Simon Olsson
- & Christian Griesinger
-
Article
| Open Access The Ube2m-Rbx1 neddylation-Cullin-RING-Ligase proteins are essential for the maintenance of Regulatory T cell fitness
The Ube2m-Rbx1 neddylation-Cullin-RING-Ligase proteins are essential for the maintenance of Regulatory T cell fitness
Absence of regulatory T cells results in a severe inflammatory disease which leads to death in infancy in both human patients and in mouse models. Authors show here that in mice, conditional deletion of Rbx1, the RING component of Cullin-RING ligases in regulatory T cells causes a similar phenotype, due to the disrupted degradation of important regulatory proteins.
- Di Wu
- , Haomin Li
- & Yi Sun
-
Article
| Open Access A modular toolbox to generate complex polymeric ubiquitin architectures using orthogonal sortase enzymes
A modular toolbox to generate complex polymeric ubiquitin architectures using orthogonal sortase enzymes
Ubiquitin (Ub) and Ub-like modifiers (Ubls) can form chains of various topologies, but preparing defined chains for functional studies remains challenging. Here, the authors develop chemoenzymatic tools to tailormake Ub/Ubl chains and study the involvement of specific Ub/SUMO chains in DNA repair.
- Maximilian Fottner
- , Maria Weyh
- & Kathrin Lang
-
Article
| Open Access Structural basis for UFM1 transfer from UBA5 to UFC1
Structural basis for UFM1 transfer from UBA5 to UFC1
Ufmylation is a well-established ubiquitin-like protein modification, but its mechanism is largely unclear. Here, the authors present a crystal structure of the ufmylation-specific E1-E2 complex, revealing differences to the ubiquitination machinery and mechanistic details of the ufmylation process.
- Manoj Kumar
- , Prasanth Padala
- & Reuven Wiener
-
Article
| Open Access Reconstitution defines the roles of p62, NBR1 and TAX1BP1 in ubiquitin condensate formation and autophagy initiation
Reconstitution defines the roles of p62, NBR1 and TAX1BP1 in ubiquitin condensate formation and autophagy initiation
Misfolded proteins are ubquitinated and subsequently condensed by cargo receptors for selective autophagy. Here, the authors use in vitro reconstitution to elegantly dissect how the receptors p62/SQSTM1, NBR1 and TAX1BP1 contribute to p62-ubiquitin condensate formation and degradation by autophagy.
- Eleonora Turco
- , Adriana Savova
- & Sascha Martens
-
Article
| Open Access Traceless native chemical ligation of lipid-modified peptide surfactants by mixed micelle formation
Traceless native chemical ligation of lipid-modified peptide surfactants by mixed micelle formation
Sequestration of reactants in lipid vesicles is a strategy prevalent in biological systems to raise the rate and specificity of chemical reactions. Here, the authors show that micelle-assisted reactions facilitate native chemical ligation between a peptide-thioester and a Cys-peptide modified by a lipid-like moiety.
- Shuaijiang Jin
- , Roberto J. Brea
- & Neal K. Devaraj
-
Article
| Open Access A deubiquitylase with an unusually high-affinity ubiquitin-binding domain from the scrub typhus pathogen Orientia tsutsugamushi
A deubiquitylase with an unusually high-affinity ubiquitin-binding domain from the scrub typhus pathogen Orientia tsutsugamushi
Many pathogens manipulate ubiquitin-mediated signaling to evade host cell defense. Here, the authors characterize the structure and enzymatic activity of a deubiquitylase domain from the causative pathogen of scrub typhus, providing evidence for a distinct mechanism of ubiquitin chain selectivity.
- Jason M. Berk
- , Christopher Lim
- & Mark Hochstrasser
-
Article
| Open Access UBB pseudogene 4 encodes functional ubiquitin variants
UBB pseudogene 4 encodes functional ubiquitin variants
Ubiquitin pseudogenes are present in many organisms but whether they encode functional proteins has remained unclear. Here, the authors show that human UBB pseudogene 4 produces ubiquitin variants with amino acid compositions and cellular functions that are distinct from canonical ubiquitin.
- Marie-Line Dubois
- , Anna Meller
- & François-Michel Boisvert
-
Article
| Open Access Structural insights into ubiquitin recognition and Ufd1 interaction of Npl4
Structural insights into ubiquitin recognition and Ufd1 interaction of Npl4
The Lys48-linked polyubiquitin-mediated proteasomal degradation in yeast depends on Cdc48 and its cofactors Ufd1 and Npl4. Here, the authors present crystal structures of Npl4 bound to Lys48-linked diubiquitin and the Npl4-binding motif of Ufd1, providing insights into the reaction mechanism of the Cdc48- Ufd1/Npl4 complex.
- Yusuke Sato
- , Hikaru Tsuchiya
- & Shuya Fukai
-
Article
| Open Access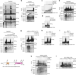 K27-linked ubiquitination of BRAF by ITCH engages cytokine response to maintain MEK-ERK signaling
K27-linked ubiquitination of BRAF by ITCH engages cytokine response to maintain MEK-ERK signaling
BRAF drives MEK/ERK activation to facilitate tumorigenesis. Here, the authors show that in response to pro-inflammatory cytokines, ITCH mediates a non-proteolytic ubiquitination and activation of BRAF, which in turn sustains MEK/ERK signaling to facilitate melanoma cell growth.
- Qing Yin
- , Tao Han
- & Lixin Wan
-
Article
| Open Access UFL1 promotes histone H4 ufmylation and ATM activation
UFL1 promotes histone H4 ufmylation and ATM activation
Ufmylation is a ubiquitylation-like protein modification but only a few ufmylation substrates and functions have been discovered so far. Here, the authors demonstrate ufmylation of histone H4 upon DNA damage and show that this modification is involved in the amplification of ATM activation.
- Bo Qin
- , Jia Yu
- & Zhenkun Lou
-
Article
| Open Access Rpn11-mediated ubiquitin processing in an ancestral archaeal ubiquitination system
Rpn11-mediated ubiquitin processing in an ancestral archaeal ubiquitination system
Ubiquitin modification also occurs in archaea. Here, the authors characterize an archaeal ancestral ubiquitination system, present the crystal structure of the archaeal deubiquitinase Rpn11 from Caldiarchaeum subterraneum bound to ubiquitin and provide insights into evolutionary relationships.
- Adrian C. D. Fuchs
- , Lorena Maldoner
- & Jörg Martin
-
Article
| Open Access Structure of ubiquitylated-Rpn10 provides insight into its autoregulation mechanism
Structure of ubiquitylated-Rpn10 provides insight into its autoregulation mechanism
Ubiquitin (Ub) receptors are responsible for the recognition of ubiquitylated proteins. Here the authors describe the crystal structure of the ubiquitylated form of the Ub-receptor Rpn10, which suggest that ubiquitylation of Rpn10 promotes its dissociation from the proteasome.
- Tal Keren-Kaplan
- , Lee Zeev Peters
- & Gali Prag
-
Article
| Open Access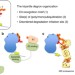 Tripartite degrons confer diversity and specificity on regulated protein degradation in the ubiquitin-proteasome system
Tripartite degrons confer diversity and specificity on regulated protein degradation in the ubiquitin-proteasome system
Degrons are determinants within proteins that direct programmed degradation by the ubiquitinproteasome system. Here, the authors propose a three-part degron architecture which contains an E3-ligase recognition motif, a ubiquitination site(s), and a disordered site to initiate degradation.
- Mainak Guharoy
- , Pallab Bhowmick
- & Peter Tompa
-
Article
| Open Access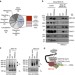 A non-proteolytic role for ubiquitin in deadenylation of MHC-I mRNA by the RNA-binding E3-ligase MEX-3C
A non-proteolytic role for ubiquitin in deadenylation of MHC-I mRNA by the RNA-binding E3-ligase MEX-3C
mRNA deadenylation, the first step in regulated degradation, is mediated by the action of the CCR4-NOT and PAN2-PAN3 complexes. Here the authors show that the RNA-binding E3 ubiquitin-ligase MEX-3C associates with the CCR4-NOT complex and ubiquitinates the catalytic subunit CNOT7 to regulate its deadenylation activity.
- Florencia Cano
- , Radu Rapiteanu
- & Paul J. Lehner
-
Article
| Open Access Functional genomics identifies negative regulatory nodes controlling phagocyte oxidative burst
Functional genomics identifies negative regulatory nodes controlling phagocyte oxidative burst
Phagocytes employ multiple bactericidal mechanisms to kill microorganisms, including the generation of toxic superoxide and other reactive oxygen species. Here the authors utilize a multi-omics approach to identify and characterize new regulatory nodes implicated in mucosal immunity that control phagocyte oxidative burst.
- Daniel B. Graham
- , Christine E. Becker
- & Ramnik J. Xavier
-
Article
| Open Access The unexpected role of polyubiquitin chains in the formation of fibrillar aggregates
The unexpected role of polyubiquitin chains in the formation of fibrillar aggregates
Ubiquitin is a stable and soluble protein, but it is commonly found in inclusion bodies in neurodegenerative disorders and cancer. Here, Morimoto et al. report that increasing ubiquitin chain length leads to the formation of amyloid-like fibrils, which are degraded by an autophagy mechanism.
- Daichi Morimoto
- , Erik Walinda
- & Masahiro Shirakawa
-
Article
| Open Access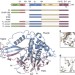 The DUSP–Ubl domain of USP4 enhances its catalytic efficiency by promoting ubiquitin exchange
The DUSP–Ubl domain of USP4 enhances its catalytic efficiency by promoting ubiquitin exchange
Ubiquitin-specific protease USP4 regulates several cellular signalling pathways. Here, Clerici et al.show that the DUSP–Ubl domain of USP4 is required for full catalytic activity, by enhancing the release of ubiquitin from the catalytic site after substrate hydrolysis.
- Marcello Clerici
- , Mark P. A. Luna-Vargas
- & Titia K. Sixma
-
Article
| Open Access Arabidopsis nitrate reductase activity is stimulated by the E3 SUMO ligase AtSIZ1
Arabidopsis nitrate reductase activity is stimulated by the E3 SUMO ligase AtSIZ1
Posttranslational modification of proteins by small ubiquitin-related modifier is a response to stress signalling in plants. Here, theArabdiposisprotein SIZ1 is shown to cause SUMOylation of nitrate reductases 1 and 2 and to increase their activity, suggesting that SIZ1 controls nitrate uptake via SUMOylation.
- Bong Soo Park
- , Jong Tae Song
- & Hak Soo Seo

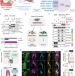 A ubiquitin-based effector-to-inhibitor switch coordinates early brain, craniofacial, and skin development
A ubiquitin-based effector-to-inhibitor switch coordinates early brain, craniofacial, and skin development