Featured
-
-
Article
| Open Access A toxin-deformation dependent inhibition mechanism in the T7SS toxin-antitoxin system of Gram-positive bacteria
A toxin-deformation dependent inhibition mechanism in the T7SS toxin-antitoxin system of Gram-positive bacteria
Antimicrobial toxins are secreted by bacteria to kill rival species. Here the authors report the mechanism of inhibition of EsaD, a toxin secreted by some S. aureus strains to kill competitors that lack the antitoxin EsaG, showing marked mechanistic differences to other Type II toxin-antitoxin systems.
- Yongjin Wang
- , Yang Zhou
- & Zhi-Min Zhang
-
Article
| Open Access Proteolytic processing induces a conformational switch required for antibacterial toxin delivery
Proteolytic processing induces a conformational switch required for antibacterial toxin delivery
Contact-dependent growth inhibition (CDI) is an important mechanism of bacterial competition. Here, Bartelli et al. show that proteolytic processing of a CDI toxin induces a conformational switch required for translocation into target bacteria.
- Nicholas L. Bartelli
- , Victor J. Passanisi
- & Christopher S. Hayes
-
Article
| Open Access Mre11-Rad50 oligomerization promotes DNA double-strand break repair
Mre11-Rad50 oligomerization promotes DNA double-strand break repair
The Mre11-Rad50 (MR) complex has key functions in the detection, signaling and repair of DNA breaks. Here the authors use transmission electron microscopy to show MR oligomerization is governed by a small beta-sheet protruding from the head domain of Rad50 at the base of the MR structure, and reveal MR head domain oligomerization is required for efficient DNA end resection.
- Vera M. Kissling
- , Giordano Reginato
- & Matthias Peter
-
Article
| Open Access Towards plant resistance to viruses using protein-only RNase P
Towards plant resistance to viruses using protein-only RNase P
New approaches to plant disease control are important for pathogens that are difficult to control by existing methods. Here, the authors report a potential strategy to combat plant viruses by cytosolic expressed protein-only RNase P and show its ability for in vitro cleavage of tRNA-like structures existing in many plant viruses.
- Anthony Gobert
- , Yifat Quan
- & Philippe Giegé
-
Article
| Open Access Target preference of Type III-A CRISPR-Cas complexes at the transcription bubble
Target preference of Type III-A CRISPR-Cas complexes at the transcription bubble
Type III CRISPR-Cas systems are able to target transcriptionally active DNA sequences in phages and plasmids. Here, the authors reveal the mechanism of the target nucleic acid preference of Type III-A CRISPR-Cas complexes at the transcription bubble by a combination of structural and biochemical approaches.
- Tina Y. Liu
- , Jun-Jie Liu
- & Jennifer A. Doudna
-
Article
| Open Access Dynamic coordination of two-metal-ions orchestrates λ-exonuclease catalysis
Dynamic coordination of two-metal-ions orchestrates λ-exonuclease catalysis
Metal ions at the active site of an enzyme act as cofactors and their dynamic fluctuations might influence enzyme activity. Here authors use single-molecule FRET to study λ-exonuclease and find that metal-ion-coordination is correlated with enzymatic reaction-steps.
- Wonseok Hwang
- , Jungmin Yoo
- & Gwangrog Lee
-
Article
| Open Access Characterizing a thermostable Cas9 for bacterial genome editing and silencing
Characterizing a thermostable Cas9 for bacterial genome editing and silencing
CRISPR-Cas9 genome engineering tools have found wide application in a range of species, however they are unsuitable for applications at elevated temperatures. Here the authors characterise ThermoCas9 from which is functional from 20°C to 70°C.
- Ioannis Mougiakos
- , Prarthana Mohanraju
- & John van der Oost
-
Article
| Open Access Real-space and real-time dynamics of CRISPR-Cas9 visualized by high-speed atomic force microscopy
Real-space and real-time dynamics of CRISPR-Cas9 visualized by high-speed atomic force microscopy
CRISPR RNA-guided endonuclease Cas9 recognizes and cleaves the double-stranded DNA complementary to the RNA guide. Here the authors use high-speed atomic force micropcopy (HS-AFM) to visualize the conformational dynamics of Cas9 during its DNA targeting and cleavage processes.
- Mikihiro Shibata
- , Hiroshi Nishimasu
- & Osamu Nureki
-
Article
| Open Access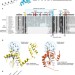 Structural insights into the function of ZRANB3 in replication stress response
Structural insights into the function of ZRANB3 in replication stress response
ZRANB3 (Zinc-finger, RAN-Binding domain containing 3) is a structure-specific endonuclease that is recruited to DNA breaks and stressed replication forks. Here the authors present the crystal structure of the ZRANB3 endonuclease domain and analyse how ZRANB3 is regulated by the DNA clamp PCNA.
- Marek Sebesta
- , Christopher D. O. Cooper
- & Dragana Ahel
-
Article
| Open Access MNase titration reveals differences between nucleosome occupancy and chromatin accessibility
MNase titration reveals differences between nucleosome occupancy and chromatin accessibility
Nucleosome positioning and chromatin accessibility are important contributors to the regulation of gene expression. Here the authors describe a method that allows the simultaneous measurement of nucleosome occupancy and chromatin accessibility in the same assay, revealing new features of chromatin organization linked to gene regulation.
- Jakub Mieczkowski
- , April Cook
- & Michael Y. Tolstorukov
-
Article
| Open Access The exonuclease activity of DNA polymerase γ is required for ligation during mitochondrial DNA replication
The exonuclease activity of DNA polymerase γ is required for ligation during mitochondrial DNA replication
Mitochondrial DNA (mtDNA) polymerase γ has a 3′–5′ exonuclease proofreading activity. Here, the authors show it is required for creating ligatable ends during mtDNA replication, and inactivation of the activity in mice causes strand-specific nicks in DNA and the formation of linear mtDNA fragments.
- Bertil Macao
- , Jay P. Uhler
- & Maria Falkenberg
-
Article |
 Structural insights into 5′ flap DNA unwinding and incision by the human FAN1 dimer
Structural insights into 5′ flap DNA unwinding and incision by the human FAN1 dimer
FAN1 is a structure-specific nuclease that plays a major role in eliminating highly cytotoxic interstrand DNA crosslinks. Here, Zhao et al. present several crystal structures of FAN1 in complex with DNA substrates and biochemical analyses that establish how FAN1 functions to resolve interstrand DNA crosslinks.
- Qi Zhao
- , Xiaoyu Xue
- & Yong Xiong
-
Article
| Open Access Endonuclease V cleaves at inosines in RNA
Endonuclease V cleaves at inosines in RNA
Bacterial endonuclease V enzymes are characterized as DNA repair proteins. Here the authors show that human endonuclease V is an inosine-specific ribonuclease, indicating a role for this enzyme in normal RNA metabolism rather than DNA repair.
- Erik Sebastian Vik
- , Meh Sameen Nawaz
- & Ingrun Alseth
-
Article
| Open Access Human endonuclease V is a ribonuclease specific for inosine-containing RNA
Human endonuclease V is a ribonuclease specific for inosine-containing RNA
In Escherichia coli, the highly conserved enzyme endonuclease V has a role in DNA repair. Here the authors show that human endonuclease V is an inosine 3' endoribonuclease and that Tudor Staphylococcal nuclease enhances this activity, suggesting a role for human endonuclease V in RNA metabolism.
- Yoko Morita
- , Toshihiro Shibutani
- & Isao Kuraoka

 Single-exonuclease nanocircuits reveal the RNA degradation dynamics of PNPase and demonstrate potential for RNA sequencing
Single-exonuclease nanocircuits reveal the RNA degradation dynamics of PNPase and demonstrate potential for RNA sequencing