Article
|
Open Access
Featured
-
-
Article
| Open Access Altered chromatin landscape and enhancer engagement underlie transcriptional dysregulation in MED12 mutant uterine leiomyomas
Altered chromatin landscape and enhancer engagement underlie transcriptional dysregulation in MED12 mutant uterine leiomyomas
Somatic mutations in MED12 have been implicated as the causal genetic lesion in the majority of uterine leiomyomas. Here, the authors profile the chromatin landscape of matched normal and leiomyoma tissues and find that changes in enhancer acetylation, enhancer-promoter interaction strength, differential enhancer usage and transcription factor AP-1 occupancy are significant drivers of transcriptional dysregulation in MED12 mutant leiomyomas.
- Mthabisi B. Moyo
- , J. Brandon Parker
- & Debabrata Chakravarti
-
Article
| Open Access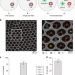 Inference and effects of barcode multiplets in droplet-based single-cell assays
Inference and effects of barcode multiplets in droplet-based single-cell assays
It is assumed that single-cell analyses capture one barcode per cell. Here, the authors show that up to 21% of cell barcodes on the 10X Chromium scATAC-seq assay may be derived from barcode multiplets.
- Caleb A. Lareau
- , Sai Ma
- & Jason D. Buenrostro
-
Article
| Open Access Chromatin mapping and single-cell immune profiling define the temporal dynamics of ibrutinib response in CLL
Chromatin mapping and single-cell immune profiling define the temporal dynamics of ibrutinib response in CLL
Ibrutinib, a Bruton tyrosine kinase inhibitor, provides effective treatment for chronic lymphocytic leukemia (CLL). Here, the authors describe time-dependent molecular changes to malignant cells and to the immune system in patients undergoing ibrutinib therapy, with can be used for therapy monitoring.
- André F. Rendeiro
- , Thomas Krausgruber
- & Christoph Bock
-
Article
| Open Access Epigenetics meets proteomics in an epigenome-wide association study with circulating blood plasma protein traits
Epigenetics meets proteomics in an epigenome-wide association study with circulating blood plasma protein traits
DNA methylation is associated with complex traits and the expression of genes and proteins. Here, Zaghlool et al. perform epigenome-wide association studies for 1,123 plasma proteins, replicate obtained protein (p)QTMs in an independent cohort and find overlap of pQTMs with expression QTMs and previously reported disease associations.
- Shaza B. Zaghlool
- , Brigitte Kühnel
- & Karsten Suhre
-
Article
| Open Access Edematous severe acute malnutrition is characterized by hypomethylation of DNA
Edematous severe acute malnutrition is characterized by hypomethylation of DNA
The edematous form of severe acute childhood malnutrition (ESAM) presents with more severe multi-organ dysfunction than non-edematous SAM (NESAM). Here the authors assess genome-wide DNA methylation in buccal cells of SAM children and find that ESAM is characterized by hypomethylation at genes associated with disorders of nutrition and metabolism, including fatty liver and diabetes.
- Katharina V. Schulze
- , Shanker Swaminathan
- & Neil A. Hanchard
-
Article
| Open Access Atlas of quantitative single-base-resolution N6-methyl-adenine methylomes
Atlas of quantitative single-base-resolution N6-methyl-adenine methylomes
N6-methyladenosine (m6A) and N6,2′-O-dimethyladenosine (m6Am) are eukaryotic mRNA modifications. Here the authors develop m6A-Crosslinking-Exonuclease-sequencing to map quantitative methylome changes at single-base-resolution after individually knocking out each known methyltransferase or demethylase.
- Casslynn W. Q. Koh
- , Yeek Teck Goh
- & W. S. Sho Goh
-
Article
| Open Access The epigenomic landscape of transposable elements across normal human development and anatomy
The epigenomic landscape of transposable elements across normal human development and anatomy
Although most are silenced, certain transposable elements (TEs) have been co-opted by the host. Here, the authors quantify the epigenomic status of TEs using data from the Roadmap Epigenomics Project, provide a systematic profile of TE activity across normal human tissues and development, finding that TEs encompass a quarter of the human regulatory epigenome, with 47% of TEs in regulatory states.
- Erica C. Pehrsson
- , Mayank N. K. Choudhary
- & Ting Wang
-
Article
| Open Access Revealing Hi-C subcompartments by imputing inter-chromosomal chromatin interactions
Revealing Hi-C subcompartments by imputing inter-chromosomal chromatin interactions
Genome-wide mapping of chromatin interactions reveals various levels of 3D genome organization. Here, the authors develop SNIPER, a computational method for identifying subcompartments using Hi-C data with moderate coverage.
- Kyle Xiong
- & Jian Ma
-
Article
| Open Access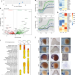 A genome-wide assessment of the ancestral neural crest gene regulatory network
A genome-wide assessment of the ancestral neural crest gene regulatory network
An understanding of the ancestral state of the neural crest (NC) gene regulatory network (GRN) gives insight into vertebrate evolution. Here, the authors use transcriptomic and chromatin accessibility analyses of the lamprey NC, as well as cross-species enhancer assays, to identify GRN elements conserved throughout vertebrates.
- Dorit Hockman
- , Vanessa Chong-Morrison
- & Tatjana Sauka-Spengler
-
Article
| Open Access SCALE method for single-cell ATAC-seq analysis via latent feature extraction
SCALE method for single-cell ATAC-seq analysis via latent feature extraction
Single-cell ATAC-seq data is challenging to analyse for reasons such as high dimensionality and sparsity. Here, the authors develop SCALE, a deep learning method that leverages latent feature extraction for various tasks of scATACseq data analysis.
- Lei Xiong
- , Kui Xu
- & Qiangfeng Cliff Zhang
-
Article
| Open Access Ageing affects DNA methylation drift and transcriptional cell-to-cell variability in mouse muscle stem cells
Ageing affects DNA methylation drift and transcriptional cell-to-cell variability in mouse muscle stem cells
Age-related tissue alterations have been associated with a decline in stem cell number and function. Here the authors report a single cell multi-omics study of mouse muscle stem cells, combining single cell transcriptome and DNA methylome profiling and find that aged cells have a global increase of uncoordinated transcriptional heterogeneity biased towards genes regulating cell-niche interactions.
- Irene Hernando-Herraez
- , Brendan Evano
- & Wolf Reik
-
Article
| Open Access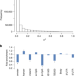 Genome-wide identification of DNA methylation QTLs in whole blood highlights pathways for cardiovascular disease
Genome-wide identification of DNA methylation QTLs in whole blood highlights pathways for cardiovascular disease
Differentially methylated CpGs can inform on disease mechanisms and be useful as biomarkers. Here, the authors perform GWAS for DNA methylation in whole blood, cis- and trans-meQTL mapping, followed by Mendelian randomization analysis that links meQTLs with cardiovascular diseases.
- Tianxiao Huan
- , Roby Joehanes
- & Daniel Levy
-
Article
| Open Access DNA methylation loss promotes immune evasion of tumours with high mutation and copy number load
DNA methylation loss promotes immune evasion of tumours with high mutation and copy number load
Demethylation of the genome is found in cancer. Here, the authors show that genomic demethylation entails changes in promoter methylation and gene expression associated with immune escape and suggest that the epigenetic alterations may be an important determinant of responses to immunotherapy.
- Hyunchul Jung
- , Hong Sook Kim
- & Jung Kyoon Choi
-
Article
| Open Access A high-resolution 3D epigenomic map reveals insights into the creation of the prostate cancer transcriptome
A high-resolution 3D epigenomic map reveals insights into the creation of the prostate cancer transcriptome
In prostate cancer, chromatin structure can impact the transcriptome. Here, the authors develop high resolution chromatin interaction maps in prostate cancer cells using in situ Hi-C, revealing prostate cancer-specific TADs and enhancer-promoter loops surrounding the androgen receptor (AR) locus.
- Suhn Kyong Rhie
- , Andrew A. Perez
- & Peggy J. Farnham
-
Article
| Open Access Chromatin loops associated with active genes and heterochromatin shape rice genome architecture for transcriptional regulation
Chromatin loops associated with active genes and heterochromatin shape rice genome architecture for transcriptional regulation
Three-dimensional genome organization and its effect on transcription remain elusive in rice. Here, the authors map promoter–promoter interactions and heterochromatin interactions using ChIA-PET and reveal spatial correlation between the genetic regulation of eQTLs and e-traits.
- Lun Zhao
- , Shuangqi Wang
- & Xingwang Li
-
Article
| Open Access Cell-type-specific resolution epigenetics without the need for cell sorting or single-cell biology
Cell-type-specific resolution epigenetics without the need for cell sorting or single-cell biology
Compared to bulk data, cell-type-specific DNA methylation data provide higher resolution of epigenetic variation. Here, the authors introduce Tensor Composition Analysis, a novel computational approach for learning cell-type-specific DNA methylation from tissue-level bulk data, and show its application in epigenome-wide association studies.
- Elior Rahmani
- , Regev Schweiger
- & Eran Halperin
-
Article
| Open Access Genome and epigenome wide studies of neurological protein biomarkers in the Lothian Birth Cohort 1936
Genome and epigenome wide studies of neurological protein biomarkers in the Lothian Birth Cohort 1936
Plasma levels of neurological proteins have the potential to serve as biomarkers for neurological conditions. Here, Hillary et al. perform genome- and epigenome-wide association studies for 92 neurological proteins and identify 41 genomic loci for 33 proteins and 26 CpG sites for 9 proteins.
- Robert F. Hillary
- , Daniel L. McCartney
- & Riccardo E. Marioni
-
Article
| Open Access Zebrafish preserve global germline DNA methylation while sex-linked rDNA is amplified and demethylated during feminisation
Zebrafish preserve global germline DNA methylation while sex-linked rDNA is amplified and demethylated during feminisation
Germline cells transfer genetic information to offspring, and in zebrafish, drive sex determination. Here the authors report that, unlike mammals, the germline of zebrafish does not undergo genome-wide DNA methylation erasure, while amplifying and demethylating sex-linked rDNA during feminisation.
- Oscar Ortega-Recalde
- , Robert C. Day
- & Timothy A. Hore
-
Article
| Open Access Detection of cell-type-specific risk-CpG sites in epigenome-wide association studies
Detection of cell-type-specific risk-CpG sites in epigenome-wide association studies
Cellular heterogeneity is one of the major confounding factors in EWAS studies. Here the authors present a statistical method, HIgh REsolution (HIRE), which enables the detection of risk-CpG sites for individual cell types.
- Xiangyu Luo
- , Can Yang
- & Yingying Wei
-
Article
| Open Access Retention of paternal DNA methylome in the developing zebrafish germline
Retention of paternal DNA methylome in the developing zebrafish germline
Germ cells are the means of transferring genetic information to the next generation. Here the authors characterise the DNA methylomes of zebrafish primordial germ cells and find that, unlike mammals, the zebrafish germ cells do not undergo genome-wide DNA demethylation but rather retain paternal DNA methylation patterns
- Ksenia Skvortsova
- , Katsiaryna Tarbashevich
- & Ozren Bogdanovic
-
Article
| Open Access Chromatin interaction maps reveal genetic regulation for quantitative traits in maize
Chromatin interaction maps reveal genetic regulation for quantitative traits in maize
Spatial organization of regulatory elements and its impact on gene expression in plants remain unclear. Here, the authors construct maize chromatin interaction maps using chromatin interaction analysis by paired-end tag sequencing (ChIA-PET) and show their associations with gene expression and agronomic traits.
- Yong Peng
- , Dan Xiong
- & Xingwang Li
-
Article
| Open Access Long-range interactions between proximal and distal regulatory regions in maize
Long-range interactions between proximal and distal regulatory regions in maize
Chromatin interaction analysis by paired-end tag sequencing (ChIA-PET) can discover specific protein-centered chromatin interactions in high resolution. Here, the authors use ChIA-PET to reveal the complex and dynamic interactions between proximal and distal regulatory regions of genes in maize.
- En Li
- , Han Liu
- & Jinsheng Lai
-
Article
| Open Access An integrative cross-omics analysis of DNA methylation sites of glucose and insulin homeostasis
An integrative cross-omics analysis of DNA methylation sites of glucose and insulin homeostasis
Our understanding of the functional link between differential DNA methylation and type 2 diabetes and obesity remains limited. Here the authors present a blood-based EWAS of fasting glucose and insulin among 4808 non-diabetic Europeans and identify nine CpGs not previously implicated in glucose, insulin homeostasis and diabetes.
- Jun Liu
- , Elena Carnero-Montoro
- & Cornelia M. van Duijn
-
Article
| Open Access Integrated analysis of environmental and genetic influences on cord blood DNA methylation in new-borns
Integrated analysis of environmental and genetic influences on cord blood DNA methylation in new-borns
Environmental influences during prenatal development may have implications for health and disease later in life. Here, Czamara et al. assess DNA methylation in cord blood from new-born under various models including environmental and genetic effects individually and their additive or interaction effects.
- Darina Czamara
- , Gökçen Eraslan
- & Elisabeth B. Binder
-
Article
| Open Access Detection of DNA base modifications by deep recurrent neural network on Oxford Nanopore sequencing data
Detection of DNA base modifications by deep recurrent neural network on Oxford Nanopore sequencing data
DNA modification generates unique electric signals in Oxford Nanopore sequencing data but the signals can be complicated to decipher. Here, the authors develop a deep learning framework, DeepMod, to detect DNA base modifications including 5mC and 6mA using Nanopore sequencing data
- Qian Liu
- , Li Fang
- & Kai Wang
-
Article
| Open Access 6mA-DNA-binding factor Jumu controls maternal-to-zygotic transition upstream of Zelda
6mA-DNA-binding factor Jumu controls maternal-to-zygotic transition upstream of Zelda
N6-methyladenine (6mA) DNA modification is a dynamic epigenetic mark in Drosophila embryos, but how 6mA is decoded is unclear. Here, the authors show that the protein Jumu binds 6mA-marked DNA to regulate the maternal-to-zygotic transition, partly through regulation the expression of the 6mA marked pioneer factor zelda.
- Shunmin He
- , Guoqiang Zhang
- & Dahua Chen
-
Article
| Open Access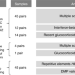 DNA methylation signatures of monozygotic twins clinically discordant for multiple sclerosis
DNA methylation signatures of monozygotic twins clinically discordant for multiple sclerosis
Monozygotic (MZ) twins are ideal to study the influence of non-genetic factors on complex phenotypes. Here, Souren et al. perform an EWAS in peripheral blood mononuclear cells from 45 MZ twins discordant for multiple sclerosis and identify disease and treatment-associated epigenetic markers.
- Nicole Y. Souren
- , Lisa A. Gerdes
- & Jörn Walter
-
Article
| Open Access Pancreatic islet chromatin accessibility and conformation reveals distal enhancer networks of type 2 diabetes risk
Pancreatic islet chromatin accessibility and conformation reveals distal enhancer networks of type 2 diabetes risk
Risk loci for type 2 diabetes (T2D) reside in pancreatic islet enhancers. Here, the authors generate high-resolution maps of islet chromatin conformation using Hi-C which they combine with ATAC-seq and ChIP-seq data to annotate candidate target genes of enhancers and validate IGF2BP2 activity in mouse islets.
- William W. Greenwald
- , Joshua Chiou
- & Kyle J. Gaulton
-
Article
| Open Access Differential methylation of enhancer at IGF2 is associated with abnormal dopamine synthesis in major psychosis
Differential methylation of enhancer at IGF2 is associated with abnormal dopamine synthesis in major psychosis
Dopamine dysregulation is centrally linked to major psychosis. Here, the authors characterise the hypomethylation of an enhancer within the insulin-like growth factor 2 gene in neurons of patients with major psychosis and provide evidence that this enhancer targets the tyrosine hydroxylase gene, responsible for dopamine synthesis.
- Shraddha Pai
- , Peipei Li
- & Viviane Labrie
-
Article
| Open Access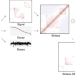 Binless normalization of Hi-C data provides significant interaction and difference detection independent of resolution
Binless normalization of Hi-C data provides significant interaction and difference detection independent of resolution
Analysis of Hi-C datasets is limited by the current existing methods for data normalization, with detection of features such as TADs and chromatin loops being inconsistent amongst different approaches. Here the authors develop Binless, a method that allows for reproducible normalization of Hi-C data independent of its resolution and compare how Binless performs in comparison with other methods.
- Yannick G. Spill
- , David Castillo
- & Marc A. Marti-Renom
-
Article
| Open Access Single-cell trajectories reconstruction, exploration and mapping of omics data with STREAM
Single-cell trajectories reconstruction, exploration and mapping of omics data with STREAM
The increasing accessibility of single cell omics technologies beyond transcriptomics demands parallel advances in analysis. Here, the authors introduce STREAM, a pipeline for reconstruction and visualization of differentiation trajectories from both single-cell RNA-seq and ATAC-seq data.
- Huidong Chen
- , Luca Albergante
- & Luca Pinello
-
Article
| Open Access Corrupted coordination of epigenetic modifications leads to diverging chromatin states and transcriptional heterogeneity in CLL
Corrupted coordination of epigenetic modifications leads to diverging chromatin states and transcriptional heterogeneity in CLL
In chronic lymphocytic leukemia (CLL), evolution is driven by transcriptional and epigenetic heterogeneity. Here, the authors integrate epigenomic analyses to show how intra-tumoral epigenetic diversity results in divergent chromatin states in CLL cells, increasing cell-to-cell transcriptional heterogeneity.
- Alessandro Pastore
- , Federico Gaiti
- & Dan A. Landau
-
Article
| Open Access RdDM-independent de novo and heterochromatin DNA methylation by plant CMT and DNMT3 orthologs
RdDM-independent de novo and heterochromatin DNA methylation by plant CMT and DNMT3 orthologs
Whether plants have true DNMT3 orthologs and their role in establishing DNA methylation are still unclear. Here, the authors show that DNMT3s are persistent through plant evolution and mediates both de novo and heterochromatin DNA methylation in the early divergent land plant Physcomitrella patens.
- Rafael Yaari
- , Aviva Katz
- & Nir Ohad
-
Article
| Open Access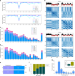 Temporal dynamic reorganization of 3D chromatin architecture in hormone-induced breast cancer and endocrine resistance
Temporal dynamic reorganization of 3D chromatin architecture in hormone-induced breast cancer and endocrine resistance
In breast cancer, the 3D architecture of the genome can impact gene regulation. Here, the authors use tethered chromatin conformation profiling to investigate 3D chromatin structure in models of hormone-induced breast cancer and endocrine resistance, finding dynamic temporal reorganisation.
- Yufan Zhou
- , Diana L. Gerrard
- & Victor X. Jin
-
Article
| Open Access Parent of origin genetic effects on methylation in humans are common and influence complex trait variation
Parent of origin genetic effects on methylation in humans are common and influence complex trait variation
Parent-of-origin effects (POE) are observed when there are different effects from alleles inherited from the two parents on phenotypic measures. Here, Zeng et al. study POE on DNA methylation in 5,101 individuals and identify genetic variants that associate with methylation variation via POE and their potential phenotypic consequences.
- Yanni Zeng
- , Carmen Amador
- & Chris S. Haley
-
Article
| Open Access Dissecting features of epigenetic variants underlying cardiometabolic risk using full-resolution epigenome profiling in regulatory elements
Dissecting features of epigenetic variants underlying cardiometabolic risk using full-resolution epigenome profiling in regulatory elements
Obesity and related metabolic complications represent an important health burden. Here the authors carry out a methylC-capture sequencing-based epigenome-wide association study to link circulating plasma lipid levels, CpG methylation and cardiometabolic risk across adipose and blood tissues.
- Fiona Allum
- , Åsa K. Hedman
- & Elin Grundberg
-
Article
| Open Access Subtle changes in chromatin loop contact propensity are associated with differential gene regulation and expression
Subtle changes in chromatin loop contact propensity are associated with differential gene regulation and expression
It is currently unclear how quantitative changes in chromatin loop propensity contribute to differential gene regulation. Here, the authors use phased Hi-C, RNA-seq, and ChIP-seq to show that subtle changes in loop propensity associate with differential gene regulation across cell types and haplotypes.
- William W. Greenwald
- , He Li
- & Kelly A. Frazer
-
Article
| Open Access Deconvolution of single-cell multi-omics layers reveals regulatory heterogeneity
Deconvolution of single-cell multi-omics layers reveals regulatory heterogeneity
Heterogeneity in gene expression and epigenetic states exists across individual cells. Here, the authors develop scCAT-seq, a technique for simultaneously performing ATAC-seq and RNA-seq within the same single cell.
- Longqi Liu
- , Chuanyu Liu
- & Xun Xu
-
Article
| Open Access Diverse motif ensembles specify non-redundant DNA binding activities of AP-1 family members in macrophages
Diverse motif ensembles specify non-redundant DNA binding activities of AP-1 family members in macrophages
Transcription factors of the AP-1 family can play diverse roles despite recognizing the same DNA sequence. Here the authors investigate the DNA binding activities of AP-1 members in mouse macrophages and apply a machine learning approach to identify motifs predicted to drive factor-specific binding profiles.
- Gregory J. Fonseca
- , Jenhan Tao
- & Christopher K. Glass
-
Article
| Open Access Replication timing and epigenome remodelling are associated with the nature of chromosomal rearrangements in cancer
Replication timing and epigenome remodelling are associated with the nature of chromosomal rearrangements in cancer
The connection between DNA replication timing and changes that occur to the epigenome in cancer are still poorly understood. Here, the authors perform Repli-Seq and integrated epigenome analyses and find that genomic regions that undergo long-range epigenetic deregulation in prostate cancer also show concordant differences in replication timing.
- Qian Du
- , Saul A. Bert
- & Susan J. Clark
-
Article
| Open Access Enhancer hijacking activates oncogenic transcription factor NR4A3 in acinic cell carcinomas of the salivary glands
Enhancer hijacking activates oncogenic transcription factor NR4A3 in acinic cell carcinomas of the salivary glands
Acinic cell carcinoma (AciCC) is a rare salivary gland carcinoma that is poorly understood. Here the authors perform genomic, transcriptomic and epigenomic profiling of AciCC and find highly recurrent and specific rearrangements [t(4;9)(q13;q31)], which lead to enhancer hijacking that activates oncogenic transcription factor NR4A3.
- Florian Haller
- , Matthias Bieg
- & Abbas Agaimy
-
Article
| Open Access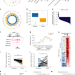 Aberrant enhancer hypomethylation contributes to hepatic carcinogenesis through global transcriptional reprogramming
Aberrant enhancer hypomethylation contributes to hepatic carcinogenesis through global transcriptional reprogramming
There are distinct hypermethylation patterns in gene promoters in hepatocellular carcinomas (HCCs). Here, the authors show that the enhancer of C/EBPβ is recurrently hypomethylated in human HCCs, recapitulating this in a transgenic murine model and linking aberrant enhancer hypomethylation to hepatocarcinogenesis.
- Lei Xiong
- , Feng Wu
- & Ka-Fai To
-
Article
| Open Access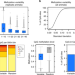 DNA methylation in mice is influenced by genetics as well as sex and life experience
DNA methylation in mice is influenced by genetics as well as sex and life experience
DNA methylation is an epigenetic mark involved in gene regulation. Here the authors investigate the extent to which genetics, sex and pregnancy influence genomic DNA methylation in mice, providing evidence of the stability of CpG methylation across generation and suggest that CpG methylation may serve as an epigenetic record of life events in somatic tissues at loci whose expression is linked to the relevant biology.
- Sara A. Grimm
- , Takashi Shimbo
- & Paul A. Wade
-
Article
| Open Access Transposable elements are regulated by context-specific patterns of chromatin marks in mouse embryonic stem cells
Transposable elements are regulated by context-specific patterns of chromatin marks in mouse embryonic stem cells
Transposable elements (TEs) fulfill essential but poorly understood roles in genome organization and gene expression control. Here the authors show that the regulation of TEs occurs through overlapping epigenetic mechanisms that control the expression and chromatin signatures at TEs.
- Jiangping He
- , Xiuling Fu
- & Andrew P. Hutchins
-
Article
| Open Access Human genome-wide measurement of drug-responsive regulatory activity
Human genome-wide measurement of drug-responsive regulatory activity
Quantification of genomic responses to environmental stimuli by current genome-scale assays is limited to indirect measurements or requires knowledge of the transcription factors involved. Here, the authors use genome-wide high-throughput reporter assays to agnostically map enhancer activity in response to glucocorticoid treatment across the human genome.
- Graham D. Johnson
- , Alejandro Barrera
- & Timothy E. Reddy
-
Article
| Open Access High-resolution genome-wide functional dissection of transcriptional regulatory regions and nucleotides in human
High-resolution genome-wide functional dissection of transcriptional regulatory regions and nucleotides in human
Millions of enhancers are predicted, but their validation remains challenging. Here, the authors report genome-wide enhancer function quantification and high-resolution dissection for millions of accessible DNA fragments, revealing driver nucleotides and helping interpret non-coding disease variants.
- Xinchen Wang
- , Liang He
- & Manolis Kellis
-
Article
| Open Access A rapid and robust method for single cell chromatin accessibility profiling
A rapid and robust method for single cell chromatin accessibility profiling
ATAC-seq is widely used to identify regulatory regions in the genome. Here the authors develop a simple and robust plate-based single-cell ATAC-seq method that works in fresh and cryopreserved cells.
- Xi Chen
- , Ricardo J. Miragaia
- & Sarah A. Teichmann
-
Article
| Open Access Local and global chromatin interactions are altered by large genomic deletions associated with human brain development
Local and global chromatin interactions are altered by large genomic deletions associated with human brain development
Copy number variants in the human genome (CNVs) are associated with neurodevelopmental and psychiatric disorders such as schizophrenia and autism. Here the authors investigate how the large deletion CNV on chromosome 22q11.2 alters chromatin organization.
- Xianglong Zhang
- , Ying Zhang
- & Alexander E. Urban
-
Article
| Open Access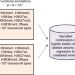 A semi-supervised approach for predicting cell-type specific functional consequences of non-coding variation using MPRAs
A semi-supervised approach for predicting cell-type specific functional consequences of non-coding variation using MPRAs
Predicting the functional consequences of non-coding genetic variants is a challenge. Here, He et al. present GenoNet, a semi-supervised method that combines information from experimentally confirmed regulatory variants with cell type- and tissue specific annotation for function prediction.
- Zihuai He
- , Linxi Liu
- & Iuliana Ionita-Laza

 Candidate silencer elements for the human and mouse genomes
Candidate silencer elements for the human and mouse genomes