Featured
-
-
Article
| Open Access Xist exerts gene-specific silencing during XCI maintenance and impacts lineage-specific cell differentiation and proliferation during hematopoiesis
Xist exerts gene-specific silencing during XCI maintenance and impacts lineage-specific cell differentiation and proliferation during hematopoiesis
Here the authors investigate the functional relevance of X-chromosome inactivation (XCI) regulator Xist in hematopoiesis. They find that Xist loss leads to changes in the ratio of hematopoietic progenitor cells and results in chromatin accessibility and transcriptional upregulation on the inactive X chromosome, including XCI escape genes known to be associated with cell cycle and immune response.
- Tianqi Yang
- , Jianhong Ou
- & Eda Yildirim
-
Article
| Open Access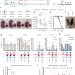 Balanced gene dosage control rather than parental origin underpins genomic imprinting
Balanced gene dosage control rather than parental origin underpins genomic imprinting
Here the authors investigate whether for imprinted genes the parent-of-origin of the expressed allele or rather appropriate gene dosage is more important for normal development. Using the differentially methylated region of Dlk1-Dio3 gene involved in imprinting, they show that correct parent-of-origin imprinting pattern is secondary to balanced gene dosage.
- Ariella Weinberg-Shukron
- , Raz Ben-Yair
- & Yonatan Stelzer
-
Article
| Open Access Activation of Xist by an evolutionarily conserved function of KDM5C demethylase
Activation of Xist by an evolutionarily conserved function of KDM5C demethylase
Here the authors show eutherian mammals co-opted the histone demethylase KDM5C during sex-chromosome evolution to induce X-chromosome inactivation by upregulating Xist expression selectively in females.
- Milan Kumar Samanta
- , Srimonta Gayen
- & Sundeep Kalantry
-
Article
| Open Access Elastic dosage compensation by X-chromosome upregulation
Elastic dosage compensation by X-chromosome upregulation
The concerted dynamics of X-chromosome upregulation and X-chromosome inactivation, which collectively balance X-chromosome expression, are not well understood. Using allelic single-cell genomics, the authors characterize the dynamics of X-chromosome upregulation and inactivation along mouse embryonic and stem cell development, calling to question keys aspects of the established model of mammalian dosage compensation.
- Antonio Lentini
- , Huaitao Cheng
- & Björn Reinius
-
Article
| Open Access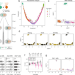 Integrated analysis of Xist upregulation and X-chromosome inactivation with single-cell and single-allele resolution
Integrated analysis of Xist upregulation and X-chromosome inactivation with single-cell and single-allele resolution
X-chromosome inactivation (XCI) ensures dosage compensation between the sexes. Here the authors perform allele-specific single-cell RNA sequencing in differentiating mouse embryonic stem cells to provide a detailed profile of the onset of XCI.
- Guido Pacini
- , Ilona Dunkel
- & Edda G. Schulz
-
Article
| Open Access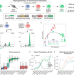 Chromosome compartments on the inactive X guide TAD formation independently of transcription during X-reactivation
Chromosome compartments on the inactive X guide TAD formation independently of transcription during X-reactivation
Both A/B compartments and TADs are thought to be absent from the inactive X chromosome, but to be re-established with transcriptional reactivation and chromatin opening during X-reactivation. Here, the authors characterise gene reactivation, chromatin opening and chromosome topology during X-reactivation, observe A/B-like compartments on the inactive X that guide TAD formation independently of transcription during X-reactivation.
- Moritz Bauer
- , Enrique Vidal
- & Bernhard Payer
-
Article
| Open Access Global chromatin conformation differences in the Drosophila dosage compensated chromosome X
Global chromatin conformation differences in the Drosophila dosage compensated chromosome X
In Drosophila, dosage compensation involves a twofold transcriptional upregulation of the single male chromosome X. Here the authors show that global conformational differences are specifically present in the male X chromosome and detectable using Hi-C data, indicating that dosage compensation affects global chromosome structure.
- Koustav Pal
- , Mattia Forcato
- & Francesco Ferrari
-
Article
| Open Access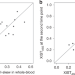 Heritability of skewed X-inactivation in female twins is tissue-specific and associated with age
Heritability of skewed X-inactivation in female twins is tissue-specific and associated with age
Skewing of X chromosome inactivation (XCI) occurs when the silencing of one parental X chromosome is non-random. Here, Zito et al. report XCI patterns in lymphoblastoid cell lines, blood, subcutaneous adipose tissue samples and skin samples of monozygotic and dizygotic twins and find XCI skew to associate with tissue and age.
- Antonino Zito
- , Matthew N. Davies
- & Kerrin S. Small
-
Article
| Open Access MAPCap allows high-resolution detection and differential expression analysis of transcription start sites
MAPCap allows high-resolution detection and differential expression analysis of transcription start sites
The position, shape and number of transcription start sites (TSS) regulate gene expression. Here authors present MAPCap, a method for high-resolution detection and differential expression analysis of TSS, and apply MAPCap to early fly development, detecting stage and sex-specific promoter and enhancer activity.
- Vivek Bhardwaj
- , Giuseppe Semplicio
- & Asifa Akhtar
-
Article
| Open Access Systematic allelic analysis defines the interplay of key pathways in X chromosome inactivation
Systematic allelic analysis defines the interplay of key pathways in X chromosome inactivation
Xist RNA is the master regulator of X chromosome inactivation. Here the authors describe a systematic analysis of Xist-mediated allelic silencing in mouse ESC models and define the contribution of different pathways that regulate gene silencing.
- Tatyana B. Nesterova
- , Guifeng Wei
- & Neil Brockdorff
-
Article
| Open Access The effect of X-linked dosage compensation on complex trait variation
The effect of X-linked dosage compensation on complex trait variation
Dosage compensation (DC) on the X chromosome has predictable effects on genetic and phenotypic trait variance. Here, the authors use information for 20 quantitative traits in the UK Biobank and across-tissue gene expression to compare X-linked heritability and the effects of trait-associated SNPs between the sexes.
- Julia Sidorenko
- , Irfahan Kassam
- & Peter M. Visscher
-
Article
| Open Access PRC1 collaborates with SMCHD1 to fold the X-chromosome and spread Xist RNA between chromosome compartments
PRC1 collaborates with SMCHD1 to fold the X-chromosome and spread Xist RNA between chromosome compartments
The inactive X (Xi)-specific S1/S2 chromosome compartments are merged by SMCHD1, but how the S1/S2 structure is constructed is unclear. The authors find that PRC1 drives the formation of S1/S2s and that the stepwise folding process of the Xi facilitates Xist RNA spreading between Xi compartments.
- Chen-Yu Wang
- , David Colognori
- & Jeannie T. Lee
-
Article
| Open Access The non-canonical SMC protein SmcHD1 antagonises TAD formation and compartmentalisation on the inactive X chromosome
The non-canonical SMC protein SmcHD1 antagonises TAD formation and compartmentalisation on the inactive X chromosome
The inactive X chromosome (Xi) has an atypical structure, with global loss of TADs, A/B compartments and formation of mega-domains. Here the authors show that the non-canonical SMC family protein, SmcHD1, important for developmental gene silencing on Xi, antagonises TAD formation and compartmentalization on the Xi in a transcription independent way.
- Michal R. Gdula
- , Tatyana B. Nesterova
- & Neil Brockdorff
-
Article
| Open Access Megadomains and superloops form dynamically but are dispensable for X-chromosome inactivation and gene escape
Megadomains and superloops form dynamically but are dispensable for X-chromosome inactivation and gene escape
The mammalian inactive X-chromosome (Xi) is organized into megadomains and superloops directed by the noncoding loci, Dxz4 and Firre. Here the authors provide evidence that megadomains do not precede Xist expression or Xi gene silencing, and suggest that Dxz4, Firre, and megadomains are dispensable for Xi silencing and escape from X-inactivation.
- John E. Froberg
- , Stefan F. Pinter
- & Jeannie T. Lee
-
Article
| Open Access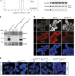 REX1 is the critical target of RNF12 in imprinted X chromosome inactivation in mice
REX1 is the critical target of RNF12 in imprinted X chromosome inactivation in mice
REX1 has been shown to regulate pluripotency of ESCs, genomic imprinting and preimplantation development in mice. Here the authors provide evidence that REX1 is the prime target of RNF12 E3 ubiquitin ligase and that Rex1 removal rescues the Rnf12 knockout phenotype in imprinted X chromosome inactivation in mice.
- Cristina Gontan
- , Hegias Mira-Bontenbal
- & Joost Gribnau
-
Article
| Open Access Female mice lacking Ftx lncRNA exhibit impaired X-chromosome inactivation and a microphthalmia-like phenotype
Female mice lacking Ftx lncRNA exhibit impaired X-chromosome inactivation and a microphthalmia-like phenotype
Although Ftx lncRNA has been linked to X-chromosome inactivation, its physiological roles in vivo remain unclear. Here the authors show that deletion of mouse Ftx causes eye abnormalities similar to human microphthalmia in a subset of female mice but rarely in males and provide evidence that Ftx plays a role in gene silencing on the inactive X chromosome.
- Yusuke Hosoi
- , Miki Soma
- & Shin Kobayashi
-
Article
| Open Access Autosomal genetic variation is associated with DNA methylation in regions variably escaping X-chromosome inactivation
Autosomal genetic variation is associated with DNA methylation in regions variably escaping X-chromosome inactivation
DNA methylation is critically involved in X chromosome inactivation (XCI) and dosage compensation, yet some X-chromosomal genes escape XCI. Here, Lujik et al. identify three autosomal genetic loci that associate with differential DNA methylation near genes that variably escape XCI in females.
- René Luijk
- , Haoyu Wu
- & Bastiaan T. Heijmans
-
Article
| Open Access Facultative dosage compensation of developmental genes on autosomes in Drosophila and mouse embryonic stem cells
Facultative dosage compensation of developmental genes on autosomes in Drosophila and mouse embryonic stem cells
In Drosophila the Male-Specific Lethal complex (MSLc) mediates upregulation of the single male X chromosome. Here the authors provide evidence that MSL2 also targets autosomal genes required for proper development and that MSL2 binds and similarly regulates mouse orthologues.
- Claudia Isabelle Keller Valsecchi
- , M. Felicia Basilicata
- & Asifa Akhtar
-
Article
| Open Access Orientation-dependent Dxz4 contacts shape the 3D structure of the inactive X chromosome
Orientation-dependent Dxz4 contacts shape the 3D structure of the inactive X chromosome
The inactive X chromosome condenses into a bipartite structure. Here the authors use cells with allelic deletions or inversions to show that the Dxz4 locus is necessary to maintain the bipartite structure and that Dxz4 orientation controls the distribution of contacts on the inactive X chromosome.
- G. Bonora
- , X. Deng
- & C. M. Disteche
-
Article
| Open Access Contribution of epigenetic landscapes and transcription factors to X-chromosome reactivation in the inner cell mass
Contribution of epigenetic landscapes and transcription factors to X-chromosome reactivation in the inner cell mass
X-chromosome inactivation is reversed in the mouse inner cell mass (ICM) through a mechanism that is not fully understood. Here, the authors investigate this process and characterize the contributions of the epigenetic landscape and transcription factors in X-linked gene reactivation dynamics.
- Maud Borensztein
- , Ikuhiro Okamoto
- & Edith Heard
-
Article
| Open Access Genetic and epigenetic features direct differential efficiency of Xist-mediated silencing at X-chromosomal and autosomal locations
Genetic and epigenetic features direct differential efficiency of Xist-mediated silencing at X-chromosomal and autosomal locations
Xist RNA is required for X chromosome inactivation but it is not well understood how Xist silences some regions more efficiently than others. Here, the authors induce ectopic Xist expression from multiple different X-linked and autosomal loci in cells to explore Xist function.
- Agnese Loda
- , Johannes H. Brandsma
- & Joost Gribnau
-
Article
| Open Access Evolution of dosage compensation under sexual selection differs between X and Z chromosomes
Evolution of dosage compensation under sexual selection differs between X and Z chromosomes
Complete sex chromosome dosage compensation is largely limited to male heterogametic species, with the majority of female heterogametic species displaying incomplete dosage compensation. Here, the authors show that sexual conflict over gene expression combined with sexual selection in males can explain this pattern.
- Charles Mullon
- , Alison E. Wright
- & Judith E. Mank
-
Article
| Open Access The role of maternal-specific H3K9me3 modification in establishing imprinted X-chromosome inactivation and embryogenesis in mice
The role of maternal-specific H3K9me3 modification in establishing imprinted X-chromosome inactivation and embryogenesis in mice
During mouse preimplantation phases, a repressive imprint is imposed on the maternal allele of Xist, which encodes a large non-coding RNA required for X-chromosome inactivation. Here the authors show that trimethylation of histone H3 at lysine 9 on Xist promoter chromatin is responsible for the maternally determined Xistrepression.
- Atsushi Fukuda
- , Junko Tomikawa
- & Akihiro Umezawa
-
Article |
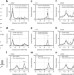 A prominent and conserved role for YY1 in Xist transcriptional activation
A prominent and conserved role for YY1 in Xist transcriptional activation
X-chromosome inactivation is a tightly regulated mechanism, which silences one of the two female X chromosomes. Here Makhlouf et al. show that the autosomal transcription factor YY1 directly promotes expression of the XistRNA—a master regulator of X-chromosome inactivation—at the onset of the inactivation process.
- Mélanie Makhlouf
- , Jean-François Ouimette
- & Claire Rougeulle
-
Article |
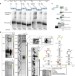 UNR facilitates the interaction of MLE with the lncRNA roX2 during Drosophila dosage compensation
UNR facilitates the interaction of MLE with the lncRNA roX2 during Drosophila dosage compensation
The process that balances expression of X-chromosomal genes between males and females is under tight regulatory control. Here, Militti et al. show that in Drosophila, the RNA-binding protein UNR functions during dosage compensation to promote the interaction between the RNA helicase MLE and the long non-coding RNA roX2.
- Cristina Militti
- , Sylvain Maenner
- & Fátima Gebauer
-
Article |
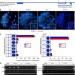 Differentiation-dependent requirement of Tsix long non-coding RNA in imprinted X-chromosome inactivation
Differentiation-dependent requirement of Tsix long non-coding RNA in imprinted X-chromosome inactivation
Imprinted mouse X-chromosome inactivation is controlled by two long non-coding RNAs, Tsix and Xist. Here, Maclary et al. demonstrate that Tsix is dispensable during the initiation and maintenance of X-inactivation in vivo and in vitro, but required to prevent Xist expression as trophectodermal progenitors differentiate.
- Emily Maclary
- , Emily Buttigieg
- & Sundeep Kalantry
-
Article
| Open Access Histone acetylation controls the inactive X chromosome replication dynamics
Histone acetylation controls the inactive X chromosome replication dynamics
How one copy of the X chromosome is silenced in replicating female somatic cells is poorly understood. Here, the authors demonstrate that the inactive X chromosome is replicated before constitutive heterochromatin and that histone hypoacetylation has a role in controlling replication of the inactive X chromosome.
- Corella S. Casas-Delucchi
- , Alessandro Brero
- & M. Cristina Cardoso
-
Article
| Open Access Chromatin remodelling complex dosage modulates transcription factor function in heart development
Chromatin remodelling complex dosage modulates transcription factor function in heart development
Inherited congenital heart defects are prevalent in the human population, but the molecular mechanisms are poorly understood. In this article, deficiency in the chromatin remodelling factor, Brg1, is shown to alter cardiac development in both mouse and zebrafish laboratory models.
- Jun K. Takeuchi
- , Xin Lou
- & Benoit G. Bruneau

 SMCHD1 has separable roles in chromatin architecture and gene silencing that could be targeted in disease
SMCHD1 has separable roles in chromatin architecture and gene silencing that could be targeted in disease