Article
|
Open Access
Featured
-
-
Article
| Open Access Nicking mechanism underlying the DNA phosphorothioate-sensing antiphage defense by SspE
Nicking mechanism underlying the DNA phosphorothioate-sensing antiphage defense by SspE
SspABCD–SspE is a phosphorothioation-sensing bacterial defence system. Here, authors find that SspE exploits DNA phosphorothioation-binding preference to modulate its biochemical activities for the self/nonself discrimination and phage resistance.
- Haiyan Gao
- , Xinqi Gong
- & Lianrong Wang
-
Article
| Open Access Structural asymmetry governs the assembly and GTPase activity of McrBC restriction complexes
Structural asymmetry governs the assembly and GTPase activity of McrBC restriction complexes
The bacterial defense system McrBC is a two-component motor-driven nuclease complex that cleaves foreign DNA. Here, the authors present the structures of the GTP-specific AAA + motor protein McrB and two McrBC complexes and discuss the molecular mechanism of how McrC binding stimulates McrB GTP hydrolysis.
- Yiming Niu
- , Hiroshi Suzuki
- & Joshua S. Chappie
-
Article
| Open Access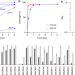 Structural basis for the promiscuous PAM recognition by Corynebacterium diphtheriae Cas9
Structural basis for the promiscuous PAM recognition by Corynebacterium diphtheriae Cas9
The RNA-guided DNA endonuclease Cas9 from Corynebacterium diphtheriae (CdCas9) recognizes a promiscuous protospacer adjacent motif (PAM). Here the authors provide insights into the CdCas9-mediated PAM recognition mechanism by determining the 2.9 Å crystal structure of CdCas9 in complex with the guide RNA and its target DNA, which is of interest for engineering of CRISPR-Cas9 genome-editor nucleases.
- Seiichi Hirano
- , Omar O. Abudayyeh
- & Osamu Nureki
-
Article
| Open Access Structural basis for the recognition of sulfur in phosphorothioated DNA
Structural basis for the recognition of sulfur in phosphorothioated DNA
DNA phosphorothioation (PT-DNA) is a DNA backbone sulfur modification that is recognized by the type-IV restriction endonuclease ScoMcrA. Here the authors provide insights into sulfur recognition by solving the crystal structure of the PT-DNA bound sulfur-binding domain (SBD) from ScoMcrA and they further show that SBD homologs are widely spread among prokaryotes.
- Guang Liu
- , Wencheng Fu
- & Xinyi He
-
Article
| Open Access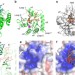 Crystal structure of APOBEC3A bound to single-stranded DNA reveals structural basis for cytidine deamination and specificity
Crystal structure of APOBEC3A bound to single-stranded DNA reveals structural basis for cytidine deamination and specificity
Cytidine deaminases are evolutionarily conserved enzymes that edit genomes by deaminating cytidine to uridine. Here the authors present the crystal structure of APOBEC3A with a single-stranded DNA substrate bound in the active site to shed light on the mechanism and specificity of substrate recognition.
- Takahide Kouno
- , Tania V. Silvas
- & Celia A. Schiffer
-
Article |
 Two-dimensional enzyme diffusion in laterally confined DNA monolayers
Two-dimensional enzyme diffusion in laterally confined DNA monolayers
Little is known about the effects of molecular crowding and confinement on biomolecule function. Castronovoet al. investigate the reactions of restriction enzymes with DNA confined in bushy matrices and find that the enzymes enter at the side of the matrix before diffusing two-dimensionally.
- Matteo Castronovo
- , Agnese Lucesoli
- & Giacinto Scoles

 Compact Cas9d and HEARO enzymes for genome editing discovered from uncultivated microbes
Compact Cas9d and HEARO enzymes for genome editing discovered from uncultivated microbes