Featured
-
-
Article
| Open Access Comparative genomics of Acinetobacter baumannii and therapeutic bacteriophages from a patient undergoing phage therapy
Comparative genomics of Acinetobacter baumannii and therapeutic bacteriophages from a patient undergoing phage therapy
A patient with a multidrug-resistant bacterial infection was successfully treated in 2016 using phage therapy. Here, the authors sequence the genomes of the therapeutic phages and three bacterial strains isolated before and during treatment, and show that the same mutations conferring phage resistance are found in in vitro-generated mutants and in phage-insensitive strains isolated from the patient.
- Mei Liu
- , Adriana Hernandez-Morales
- & Jason J. Gill
-
Article
| Open Access Mucin induces CRISPR-Cas defense in an opportunistic pathogen
Mucin induces CRISPR-Cas defense in an opportunistic pathogen
It is unknown what circumstances promote particular bacterial defenses against bacterial viruses (phages). Almeida & Hoikkala et al. show that mucin, derived from mucus, greatly accelerates CRISPR-Cas defenses against phage in an opportunistic pathogen.
- Gabriel Magno de Freitas Almeida
- , Ville Hoikkala
- & Lotta-Riina Sundberg
-
Article
| Open Access Gut virome profiling identifies a widespread bacteriophage family associated with metabolic syndrome
Gut virome profiling identifies a widespread bacteriophage family associated with metabolic syndrome
Here, the authors characterize gut viromes in a cohort of individuals with metabolic syndrome, which they find associated with a highly widespread family of gut bacteriophages they name Candidatus Heliusviridae.
- Patrick A. de Jonge
- , Koen Wortelboer
- & Hilde Herrema
-
Article
| Open Access Structural basis of template strand deoxyuridine promoter recognition by a viral RNA polymerase
Structural basis of template strand deoxyuridine promoter recognition by a viral RNA polymerase
Promoter recognition is a critical step in the initiation of transcription of DNA to RNA. Here, the authors describe a novel mechanism by which a phage-encoded RNA polymerase recognizes viral promoters containing deoxyuridines instead of thymidines.
- Alec Fraser
- , Maria L. Sokolova
- & Petr G. Leiman
-
Article
| Open Access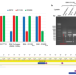 A truncated anti-CRISPR protein prevents spacer acquisition but not interference
A truncated anti-CRISPR protein prevents spacer acquisition but not interference
Phages can use ACR proteins that inhibit the adaptive immunity activities of bacterial CRISPR-Cas systems. Here, Philippe et al. show that these systems can block ACR-containing phages by targeting the acr gene, and this can select for phage mutants carrying a deletion within acr that does not block DNA cleavage (interference) but prevents the addition of new immunity (spacer acquisition).
- Cécile Philippe
- , Carlee Morency
- & Sylvain Moineau
-
Article
| Open Access Systematic and quantitative view of the antiviral arsenal of prokaryotes
Systematic and quantitative view of the antiviral arsenal of prokaryotes
Bacteria and archaea have developed multiple antiviral mechanisms. Here, Tesson et al. present a tool that automatically detects known antiviral systems in prokaryotic genomes, and show that variations in antiviral strategies correlate with genome size, viral threat, and lifestyle traits.
- Florian Tesson
- , Alexandre Hervé
- & Aude Bernheim
-
Article
| Open Access Bacteriophage treatment of disseminated cutaneous Mycobacterium chelonae infection
Bacteriophage treatment of disseminated cutaneous Mycobacterium chelonae infection
Increasing rates of multidrug-resistant bacterial infections has renewed interest in the therapeutic use of phages. Here the authors report an individual with cutaneous M. chelonae infection, and the improvement of disease upon treatment with a bacteriophage in combination with antimicrobial therapy.
- Jessica S. Little
- , Rebekah M. Dedrick
- & Graham F. Hatfull
-
Article
| Open Access Genome binning of viral entities from bulk metagenomics data
Genome binning of viral entities from bulk metagenomics data
Here, Johansen et al. develop an approach, Phages from Metagenomics Binning (PHAMB), that allows the binning of thousands of viral genomes directly from bulk metagenomics data, while simultaneously enabling clustering of viral genomes into accurate taxonomic viral populations, unveiling viral-microbial host interactions in the gut.
- Joachim Johansen
- , Damian R. Plichta
- & Simon Rasmussen
-
Article
| Open Access Combination of pre-adapted bacteriophage therapy and antibiotics for treatment of fracture-related infection due to pandrug-resistant Klebsiella pneumoniae
Combination of pre-adapted bacteriophage therapy and antibiotics for treatment of fracture-related infection due to pandrug-resistant Klebsiella pneumoniae
In this case study of a patient with fracture-related pandrug-resistant Klebsiella pneumoniae infection after long-term antibiotic therapy, the authors use a combination therapy of pre-adapted bacteriophage and antibiotics resulting in clinical, microbiological and radiological improvement.
- Anaïs Eskenazi
- , Cédric Lood
- & Jean-Paul Pirnay
-
Article
| Open Access Resolving the structure of phage–bacteria interactions in the context of natural diversity
Resolving the structure of phage–bacteria interactions in the context of natural diversity
Understanding the interactions between bacteria and their viruses (phages) in natural communities is a major challenge. Here, the authors isolate and study large numbers of marine Vibrio bacteria and their phages, and find that lytic interactions are sparse and many phages are host-strain-specific, but nevertheless recombination between some phages is common.
- Kathryn M. Kauffman
- , William K. Chang
- & Libusha Kelly
-
Article
| Open Access Multi-species host range of staphylococcal phages isolated from wastewater
Multi-species host range of staphylococcal phages isolated from wastewater
The host range of bacteriophages defines their impact on bacterial ecology and diversity. Here, Göller et al. isolate 94 staphylococcal phages from wastewater and determine their host range on 117 staphylococci from 29 species, revealing a predominant multi-species host range and thus great potential for horizontal gene transfer.
- Pauline C. Göller
- , Tabea Elsener
- & Elena Gómez-Sanz
-
Article
| Open Access Bacterial chromosomal mobility via lateral transduction exceeds that of classical mobile genetic elements
Bacterial chromosomal mobility via lateral transduction exceeds that of classical mobile genetic elements
It is commonly thought that horizontal transfer of most bacterial chromosomal genes is limited, in comparison with the frequent transfer of mobile genetic elements. Humphrey et al. show that, actually, phage-mediated lateral transduction of core chromosomal genes can be more efficient than the transfer of mobile genetic elements via conjugation or generalized transduction.
- Suzanne Humphrey
- , Alfred Fillol-Salom
- & José R. Penadés
-
Article
| Open Access Lateral transduction is inherent to the life cycle of the archetypical Salmonella phage P22
Lateral transduction is inherent to the life cycle of the archetypical Salmonella phage P22
During the transition from lysogeny (a stable association between a phage and its bacterial host) to the lytic cycle, prophage excision can be followed or preceded by DNA replication and packaging. Here, the authors show that prophage excision is delayed in Salmonella phage P22, thus allowing the packaging and transfer of large fragments of host DNA via lateral transduction.
- Alfred Fillol-Salom
- , Rodrigo Bacigalupe
- & José R. Penadés
-
Article
| Open Access Shape shifter: redirection of prolate phage capsid assembly by staphylococcal pathogenicity islands
Shape shifter: redirection of prolate phage capsid assembly by staphylococcal pathogenicity islands
Phage-inducible chromosomal islands (PICIs) are a group of mobile genetic elements that hijack the replication and assembly machinery of helper bacteriophages. Here the authors describe a mechanism by which a group of PICIs from Staphylococcus aureus re-direct the assembly pathway of their helpers using a capsid protein homolog.
- N’Toia C. Hawkins
- , James L. Kizziah
- & Terje Dokland
-
Article
| Open Access Staphylococcal phages and pathogenicity islands drive plasmid evolution
Staphylococcal phages and pathogenicity islands drive plasmid evolution
Many plasmids can be transferred between bacterial cells via conjugation; however, the mechanisms underlying the transfer of non-conjugative plasmids are less clear. Here, Humphrey et al. show that staphylococcal phages and a family of pathogenicity islands (PICIs) can mediate intra- and inter-species plasmid transfer via generalised transduction.
- Suzanne Humphrey
- , Álvaro San Millán
- & José R. Penadés
-
Article
| Open Access Ecology of inorganic sulfur auxiliary metabolism in widespread bacteriophages
Ecology of inorganic sulfur auxiliary metabolism in widespread bacteriophages
Some bacteriophage encode auxiliary metabolic genes (AMGs) that impact host metabolism and biogeochemical cycling during infection. Here the authors identify hundreds of AMGs in environmental phage encoding sulfur oxidation genes and use their global distribution to infer phage-mediated biogeochemical impacts.
- Kristopher Kieft
- , Zhichao Zhou
- & Karthik Anantharaman
-
Article
| Open Access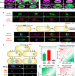 Viral speciation through subcellular genetic isolation and virogenesis incompatibility
Viral speciation through subcellular genetic isolation and virogenesis incompatibility
Virus speciation cannot be fully explained by the evolution of different host specificities. Here, Chaikeeratisak et al. identify ways viruses can remain genetically isolated despite co-infecting the same cell, providing insight into how new virus species evolve.
- Vorrapon Chaikeeratisak
- , Erica A. Birkholz
- & Joe Pogliano
-
Article
| Open Access Long-read metagenomics using PromethION uncovers oral bacteriophages and their interaction with host bacteria
Long-read metagenomics using PromethION uncovers oral bacteriophages and their interaction with host bacteria
Here, the authors profile the oral phageome of 4 healthy individuals via longread shotgun metagenomics using PromethION, a recently developed highthroughput nanopore sequencer, and uncover potential new candidate phages with enhanced scaffolding and their interaction with host bacteria.
- Koji Yahara
- , Masato Suzuki
- & Yusuke Okazaki
-
Article
| Open Access Rapid de novo evolution of lysis genes in single-stranded RNA phages
Rapid de novo evolution of lysis genes in single-stranded RNA phages
Leviviruses are phages with ssRNA genomes that encode a protein (Sgl) that induces host autolysis by interfering with bacterial cell wall synthesis. Identification of sgl genes is complicated by their small size and lack of sequence similarity. Here, Chamakura et al. use bioinformatic and experimental approaches to identify sgl genes in 244 leviviral genomes.
- Karthik R. Chamakura
- , Jennifer S. Tran
- & Ry Young
-
Article
| Open Access Architecture of the flexible tail tube of bacteriophage SPP1
Architecture of the flexible tail tube of bacteriophage SPP1
Bacteriophages of the Siphoviridae family have a long, flexible, non-contractile tail that has been difficult to characterize structurally. Here, the authors present the atomic structure of the tail tube of one of these phages, showing a hollow flexible tube formed by hexameric rings stacked by flexible linkers.
- Maximilian Zinke
- , Katrin A. A. Sachowsky
- & Adam Lange
-
Perspective
| Open Access Exploring the synthetic biology potential of bacteriophages for engineering non-model bacteria
Exploring the synthetic biology potential of bacteriophages for engineering non-model bacteria
Non-model bacteria offer unique and versatile metabolisms for synthetic biology. In this Perspective, the authors explore the limited availability of well-characterised biological parts in these species and argue that bacteriophages represent a diverse trove of orthogonal parts.
- Eveline-Marie Lammens
- , Pablo Ivan Nikel
- & Rob Lavigne
-
Article
| Open Access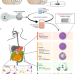 In situ reprogramming of gut bacteria by oral delivery
In situ reprogramming of gut bacteria by oral delivery
It is difficult to precisely target bacterial populations in the mammalian gut. Here the authors use encapsulated phages to deliver dCas9 to E. coli in the mouse gut to modulate RFP expression.
- Bryan B. Hsu
- , Isaac N. Plant
- & Pamela A. Silver
-
Article
| Open Access A distinct lineage of Caudovirales that encodes a deeply branching multi-subunit RNA polymerase
A distinct lineage of Caudovirales that encodes a deeply branching multi-subunit RNA polymerase
Viruses have been difficult to position in the Tree of Life using phylogenetic methods. This study uses an ancient enzyme multi-subunit RNA polymerase (RNAP) to reveal a novel viral group, the Caudovirales, and to suggest an ancient origin of RNAP in this group.
- Alaina R. Weinheimer
- & Frank O. Aylward
-
Article
| Open Access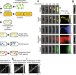 Emerging heterogeneous compartments by viruses in single bacterial cells
Emerging heterogeneous compartments by viruses in single bacterial cells
Here, the authors apply live-cell and in situ fluorescence imaging at the single-molecule level to examine lambda DNA replication in single cells, finding that individual phage DNAs sequester host factors to their own vicinity and confine their replicated DNAs into separate compartments, suggesting that phage decision-making transcripts are spatially organized in separate compartments to allow distinct subcellular decisions to develop.
- Jimmy T. Trinh
- , Qiuyan Shao
- & Lanying Zeng
-
Article
| Open Access Development of CRISPR-Cas13a-based antimicrobials capable of sequence-specific killing of target bacteria
Development of CRISPR-Cas13a-based antimicrobials capable of sequence-specific killing of target bacteria
CRISPR technology is emerging as a potential antimicrobial against antimicrobial-resistant bacteria. Here the authors develop a bacteriophage delivered Cas13a system for killing target bacteria and detecting bacterial genes.
- Kotaro Kiga
- , Xin-Ee Tan
- & Longzhu Cui
-
Article
| Open Access An amber obligate active site-directed ligand evolution technique for phage display
An amber obligate active site-directed ligand evolution technique for phage display
Most epigenetic regulator inhibitors target tunnels of active sites, rather than the peptide binding groove, leading to concerns with low selectivity. Here the authors use an amber obligate phage library to rapidly identify isoform-selective inhibitors of SIRT2.
- Jeffery M. Tharp
- , J. Trae Hampton
- & Wenshe Ray Liu
-
Article
| Open Access Acquisition, transmission and strain diversity of human gut-colonizing crAss-like phages
Acquisition, transmission and strain diversity of human gut-colonizing crAss-like phages
CrAss-like phages are bacterial viruses often found in the human gut. Here, Siranosian et al. analyze gut metagenomic data to evaluate the patterns of acquisition, transmission and strain diversity of these phages in mother-infant pairs and in patients undergoing fecal microbiota transplantation.
- Benjamin A. Siranosian
- , Fiona B. Tamburini
- & Ami S. Bhatt
-
Article
| Open Access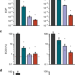 Type I-F CRISPR-Cas resistance against virulent phages results in abortive infection and provides population-level immunity
Type I-F CRISPR-Cas resistance against virulent phages results in abortive infection and provides population-level immunity
Bacteria use CRISPR-Cas systems to protect themselves against viral infections. Here, Watson et al. show that a type I CRISPR-Cas system can induce abortive viral infection, where infected cells do not survive but viral propagation is decreased, thus protecting the bacterial population.
- Bridget N. J. Watson
- , Reuben B. Vercoe
- & Peter C. Fineran
-
Article
| Open Access Coordination of cohabiting phage elements supports bacteria–phage cooperation
Coordination of cohabiting phage elements supports bacteria–phage cooperation
Bacterial pathogens often carry multiple phage-derived elements within their genome. Here, the authors show that two phage elements are co-regulated in Listeria monocytogenes, the first one controlling the induction of the second one, which in turn regulates virulence of their bacterial host.
- Tal Argov
- , Shai Ran Sapir
- & Anat A. Herskovits
-
Article
| Open Access Prophages and satellite prophages are widespread in Streptococcus and may play a role in pneumococcal pathogenesis
Prophages and satellite prophages are widespread in Streptococcus and may play a role in pneumococcal pathogenesis
Prophages are viral genomes integrated within bacterial genomes. Here, Rezaei Javan et al. identify nearly 800 prophages and satellite prophages in > 1300 Streptococcus genomes, and show that a satellite prophage is associated with virulence in a mouse model of pneumococcal infection.
- Reza Rezaei Javan
- , Elisa Ramos-Sevillano
- & Angela B. Brueggemann
-
Article
| Open Access High-throughput screen reveals sRNAs regulating crRNA biogenesis by targeting CRISPR leader to repress Rho termination
High-throughput screen reveals sRNAs regulating crRNA biogenesis by targeting CRISPR leader to repress Rho termination
Small non-coding RNAs (sRNA) regulate bacterial functions by finding nucleic acids and proteins. Here the authors identify PhrS sRNA in Pseudomonas as a positive regulator of CRISPR, and show PhrS acts by binding to CRISPR leader, thereby preventing Rho-mediated transcription termination and promoting anti-bacteriophage immunity.
- Ping Lin
- , Qinqin Pu
- & Min Wu
-
Article
| Open Access Structural basis for transcription antitermination at bacterial intrinsic terminator
Structural basis for transcription antitermination at bacterial intrinsic terminator
Bacteriophages reprogram the host transcriptional machinery. Here the authors provide insights into the mechanism of how bacteriophages regulate host transcription by determining the cryo-EM structures of two bacterial transcription elongation complexes bound with the bacteriophage master host-transcription regulator protein P7.
- Linlin You
- , Jing Shi
- & Yu Zhang
-
Article
| Open Access Inhibition of CRISPR-Cas9 ribonucleoprotein complex assembly by anti-CRISPR AcrIIC2
Inhibition of CRISPR-Cas9 ribonucleoprotein complex assembly by anti-CRISPR AcrIIC2
Anti-CRISPR proteins offer the means of regulating CRISPR-Cas9 activity. Here the authors present the structure and biochemical characterisation of AcrIIC2Nme alone and in complex with Cas9.
- Annoj Thavalingam
- , Zhi Cheng
- & Karen L. Maxwell
-
Article
| Open Access Structural assembly of the tailed bacteriophage ϕ29
Structural assembly of the tailed bacteriophage ϕ29
Mature particles of bacteriophage ϕ29 consist of a 33-MDa complex formed by over 450 subunits, assembled into a head and a short tail. Here, Xu et al. report the near-atomic structures of the ϕ29 prohead, the mature virion and the genome-emptied virion, providing insights into DNA packaging and release.
- Jingwei Xu
- , Dianhong Wang
- & Ye Xiang
-
Article
| Open Access Numerous cultivated and uncultivated viruses encode ribosomal proteins
Numerous cultivated and uncultivated viruses encode ribosomal proteins
Viruses can encode genes that regulate the host's translational machinery to their advantage. Here, the authors show that viruses encode ribosomal proteins that can be incorporated into the host’s ribosome and may affect translation.
- Carolina M. Mizuno
- , Charlotte Guyomar
- & Mart Krupovic
-
Article
| Open Access Identification and characterization of a direct activator of a gene transfer agent
Identification and characterization of a direct activator of a gene transfer agent
Gene transfer agents (GTAs) are ‘domesticated’ bacteriophages that can transfer any genes between bacteria. Here, Paul Fogg identifies a protein that directly regulates transcription of GTA genes and whose expression is in turn controlled by a global cell-cycle regulator and a quorum-sensing regulator.
- Paul C. M. Fogg
-
Article
| Open Access Nucleotide-dependent DNA gripping and an end-clamp mechanism regulate the bacteriophage T4 viral packaging motor
Nucleotide-dependent DNA gripping and an end-clamp mechanism regulate the bacteriophage T4 viral packaging motor
Packaging of viral DNA depends on strong molecular motors that are powered by ATP hydrolysis. Here, the authors develop a single-molecule assay to monitor how nucleotide binding regulates motor-DNA interactions and reveal a generic mechanism that prevents exit of the whole DNA from the viral capsid during packaging.
- Mariam Ordyan
- , Istiaq Alam
- & Douglas E. Smith
-
Article
| Open Access ΦCrAss001 represents the most abundant bacteriophage family in the human gut and infects Bacteroides intestinalis
ΦCrAss001 represents the most abundant bacteriophage family in the human gut and infects Bacteroides intestinalis
Bacteriophages of the crAssphage family have not yet been isolated, despite being highly abundant in the human gut. Here, Shkoporov et al. isolate in pure culture one of these viruses and show that it infects the human gut symbiont Bacteroides intestinalis.
- Andrey N. Shkoporov
- , Ekaterina V. Khokhlova
- & Colin Hill
-
Article
| Open Access Incomplete prophage tolerance by type III-A CRISPR-Cas systems reduces the fitness of lysogenic hosts
Incomplete prophage tolerance by type III-A CRISPR-Cas systems reduces the fitness of lysogenic hosts
CRISPR-Cas systems, such as type III-A CRISPR-Cas, provide an immune mechanism for prokaryotic hosts to resist parasites, including phages. Here, the authors show that maintenance of conditionally tolerant type III-A systems can affect the fitness of Staphylococcus aureus lysogens.
- Gregory W. Goldberg
- , Elizabeth A. McMillan
- & Luciano A. Marraffini
-
Article
| Open Access Bacteriophage T5 tail tube structure suggests a trigger mechanism for Siphoviridae DNA ejection
Bacteriophage T5 tail tube structure suggests a trigger mechanism for Siphoviridae DNA ejection
Host cell recognition is mediated by the phage tail tip proteins, which then triggers viral genome delivery via the phage tail. Here, the authors combine crystallography and cryoEM to structurally characterise the bacteriophage T5 tail tube structure before and after interaction with its host receptor.
- Charles-Adrien Arnaud
- , Grégory Effantin
- & Cécile Breyton
-
Article
| Open Access Internalization of a polysialic acid-binding Escherichia coli bacteriophage into eukaryotic neuroblastoma cells
Internalization of a polysialic acid-binding Escherichia coli bacteriophage into eukaryotic neuroblastoma cells
Eukaryotic organisms are continuously exposed to bacteriophages, but these are not thought to enter non-phagocytic cells. Here, Lehti et al. show that a bacteriophage can bind to a specific receptor on the surface of human neuroblastoma cells in vitro, and be internalized via the endolysosomal route.
- Timo A. Lehti
- , Maria I. Pajunen
- & Jukka Finne
-
Article
| Open Access Long-term genomic coevolution of host-parasite interaction in the natural environment
Long-term genomic coevolution of host-parasite interaction in the natural environment
Arms races between phage and bacteria are well known from lab experiments, but insight from field systems is limited. Here, the authors show changes in the resistance and CRISPR loci of bacteria and the infectivity, host range and genome size of phage over multiple years in an aquaculture environment.
- Elina Laanto
- , Ville Hoikkala
- & Lotta-Riina Sundberg
-
Article
| Open Access Single-peptide DNA-dependent RNA polymerase homologous to multi-subunit RNA polymerase
Single-peptide DNA-dependent RNA polymerase homologous to multi-subunit RNA polymerase
Although all known RNA polymerases have multiple subunits, unrelated single-subunit polymerases have also been described. Here, the authors describe a single-subunit RNA polymerase from the SPβ prophage ofBacillus subtilis, which shares homology to multi-subunit enzymes.
- David Forrest
- , Katherine James
- & Nikolay Zenkin
-
Article
| Open Access De novo evolved interference competition promotes the spread of biofilm defectors
De novo evolved interference competition promotes the spread of biofilm defectors
The production of secreted polymers in bacterial biofilms is costly, and therefore mechanisms preventing invasion of non-producing mutants are hypothesized. Here, the authors show that non-producers can evolve the ability to better incorporate into biofilms via phage-mediated interference.
- Marivic Martin
- , Anna Dragoš
- & Ákos T. Kovács
-
Article
| Open Access Cell fate decisions emerge as phages cooperate or compete inside their host
Cell fate decisions emerge as phages cooperate or compete inside their host
The bacteriophage lambda and its hostEscherichia coli provide a model system to study cell-fate decisions. Here, Trinh et al. develop a four-colour fluorescence system at the single-cell/single-virus/single-viral-DNA level and find phages cooperate during lysogenization and compete during lysis.
- Jimmy T. Trinh
- , Tamás Székely
- & Lanying Zeng
-
Article
| Open Access A cocktail of three virulent bacteriophages prevents Vibrio cholerae infection in animal models
A cocktail of three virulent bacteriophages prevents Vibrio cholerae infection in animal models
There has been a renewed interest in the potential use of bacterial viruses (phages) for treatment or prevention of bacterial infections. Here the authors show that a cocktail of three phages can prevent cholera-like diarrhoea in animals infected withVibrio cholerae.
- Minmin Yen
- , Lynne S. Cairns
- & Andrew Camilli
-
Article
| Open Access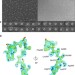 Portal protein functions akin to a DNA-sensor that couples genome-packaging to icosahedral capsid maturation
Portal protein functions akin to a DNA-sensor that couples genome-packaging to icosahedral capsid maturation
Tailed bacteriophages assemble empty precursor capsids known as procapsids that are subsequently filled with viral DNA by a genome-packaging motor. Here the authors present a structure-based analysis that suggests the signal for termination of genome packaging is achieved through a DNA-dependent symmetrization of portal protein.
- Ravi K. Lokareddy
- , Rajeshwer S. Sankhala
- & Gino Cingolani
-
Article
| Open Access Bacterial viruses enable their host to acquire antibiotic resistance genes from neighbouring cells
Bacterial viruses enable their host to acquire antibiotic resistance genes from neighbouring cells
Prophages are quiescent bacterial viruses that, when activated, produce viral particles and kill their host cells. Here, Haaber et al. show that these viral particles can mediate the transfer of antibiotic resistance genes from neighbouring cells back to the remaining prophage-containing cells.
- Jakob Haaber
- , Jørgen J. Leisner
- & Hanne Ingmer
-
Article
| Open Access The solution structure of an anti-CRISPR protein
The solution structure of an anti-CRISPR protein
Recently, anti-CRISPR proteins have been identified. Here, the authors report the solution structure of one of these proteins, and use mutational analysis to provide some insight into its function.
- Karen L. Maxwell
- , Bianca Garcia
- & Alan R. Davidson

 Personalized bacteriophage therapy to treat pandrug-resistant spinal Pseudomonas aeruginosa infection
Personalized bacteriophage therapy to treat pandrug-resistant spinal Pseudomonas aeruginosa infection