Abstract
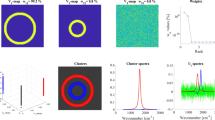
Principle component analysis (PCA) was used to extract components from the solid-state nuclear magnetic resonance (NMR) spectra of bacterial cellulose (BC). Polymers such as cellulose have several domain structures, and their structure and dynamics are reflected in the variety of solid-state spectra derived from different parameters. The complexity of the obtained spectra makes the analysis of spectra from relaxation measurements difficult. Multivariate analysis, such as PCA, has been suggested as an option to improve NMR spectral analyses. In this study, we attempted to extract peak components from cross polarization (CP) experiment data from the variable contact time spectra of BC by using PCA. The extracted peaks were annotated according to previous reports, and the existence of crystalline Iα was clearly recognized. The important components of the BC structure (that is, crystalline Iα, amorphous form and mobility) were separated in the PCA-loading plot and CP curve.
Similar content being viewed by others
Introduction
Solid-state nuclear magnetic resonance (NMR) is used to analyze the structure and dynamics of polymers. In solid-state NMR, cross polarization (CP)/magic angle spinning (MAS)1 is the most fundamental measurement. The intensity and line shape of the spectra obtained by CP/MAS can be edited by changing the contact time (CT) for the CP. The spectral change (M(CT)) is dependent on two parameters, time constant of cross polarization (TCH) and proton spin-lattice relaxation time in a rotating frame (T1ρH),2 relative to the local proton environment and molecular motion, respectively (M(CT)=M0/λ−1{1−exp(λ CT/TCH)exp(−CT/T1ρH)}, λ=1+TCH/T1ρC−TCH/T1ρH). Thus, CP experiments with variable contact time (CPVC) have been used to obtain information about the local structure and mobility during polymer analysis.3
Multivariate analysis is a potential method for the analysis of large data sets. As one of several multivariate analysis techniques, principle component analysis (PCA) has been widely used (for example, as an analytical tool for metabolomics).4 For complex samples such as biomass, PCAs are especially useful for the analysis of NMR spectra because these spectra contain a considerable amount of intensity information that can be analyzed as multivariate variables.5 On the other hand of such possibilities, these methods are necessary to collect spectra from samples with different pretreatment conditions or samples obtained from different species in order to construct the data matrix for the statistical calculation. However, there are some cases difficult to prepare different condition samples, and preparing such sample is very time-consuming; thus, alternative approaches for collecting spectra may be occasionally required.
Cellulose is the most abundant polymer in nature and one of the most versatile. The native cellulose structure exists in crystalline types Iα and Iβ and in amorphous forms. These different forms and the inherent redundancy of the polymerized molecules yield diverse NMR spectra. This diversity is the main reason for the difficulty of the NMR spectral analysis of cellulose. In previous reports, spectral analysis has been performed by preparation with reagents6 and by changing the measurement temperature.7 PCA has also been used in the spectral analysis of cellulose to investigate the effects of acid hydrolysis and solvent exchange on the spectral features of different cellulose substrates,8 but to our knowledge there is no previous example of the application of PCA to NMR spectral data collected during experiments in which only one parameter was changed in a pulse sequence, such as a relaxation time measurement sequence.
In the present work, we demonstrate PCA of the CPVC spectra of bacterial cellulose (BC), which contain information about dynamics. This investigation is the first example to analyze NMR spectra of the same sample collected by changing a parameter during a pulse sequence. Additionally, we investigate the information about the local cellulose structure obtained in this experiment.
Materials and methods
Materials
13C BC was prepared by stationary cultivation of Acetobacter xylinum in Hestrin-Schramm medium9 supplemented with 13C6-glucose (13C>99%), which was purchased from Cambridge Isotope Laboratories (Andover, MA, USA). The pellicles were cut into pieces with a food cutter.
Solid-state NMR spectroscopy
Solid-state NMR experiments were performed with a DRX-500 standard-bore spectrometer (500 MHz Bruker-BioSpin, Billerica, MA, USA) with a Bruker MAS VTN 500SB BL4 probe. The rotor was filled with ∼100 mg of 13C BC. The MAS frequency was fixed at 8 kHz. The contact time for cross-polarization (CP)1 for each CP experiment with variable contact time (CPVC, Figure 1a) was varied from 10 μs to 5 ms with 12 points, and the recycle delay was set to 3 s. Fourier transforms were performed using TOPSPIN software, version 1.3 (Bruker-BioSpin).
(a) Cross-polarization experiment with variable contact times and (b) the observed spectra of 13C BC. As shown in a, the CTs used in each experiment were 10, 20, 30, 50, 100, 250, 500, 750, 1000, 2000, 3000 and 5000 μs. The normalized spectra for the sum of the area from 55 to 110 p.p.m. are presented in c.
Dipolar-assisted rotational resonance (DARR)10, 11, 12 spectrum was measured with a mixing time of 5 ms, a recycle delay of 3 s and a MAS frequency of 8 kHz. DARR recoupling was accomplished by applying a continuous 1H wave irradiation with a 1H radio frequency intensity equal to twice the spinning frequency (that is, 16 kHz). A total of 256 t1 increments were collected for an f1 spectral width of 298.91 p.p.m.
CP-incredible natural abundance double quantum transfer experiment (CP-INADEQUATE)13 was performed with an echo time of 3.6 ms, a recycle delay of 3 s and a MAS frequency of 8 kHz. A total of 256 t1 increments were collected for an f1 spectral width of 597.82 p.p.m. A variable-amplitude CP sequence was used for the two-dimensional (2D) experiments, and the contact time was 1.2 ms. Quadrature detection for the 2D experiments was achieved by using the time-proportional phase increment method. Glycine (specifically, the 13C chemical shift of its carbonyl carbon at 176.03 p.p.m.) was used as the external reference.
Statistical calculations
The R package was used to perform PCA, and the basic strategy and method for the statistical calculation of spectroscopic data has been reported elsewhere.14, 15, 16, 17 The measured spectra (Figure 1b) were converted to text data using the Bruker command ‘convbin2asc’. The data matrix for PCA was constructed from the text data. Before performing PCA, the text data corresponding to each spectrum were first normalized by the summed areas from 55–110 p.p.m. (755 points, Figure 1c). PCA was performed with the normalized matrix. For the second PCA (Supplementary Figure S1), only the matrix data corresponding to contact times of 10–50 μs were used. In each loading plot, the components with a reject rate of <5% in the t-distribution were not considered.
Results
Extracted peaks from the PCA-loading plot of CPVC spectra
Figure 2 shows the PCA results from the CPVC spectra of 13C BC. In the score plot (Figure 2a), the plots from 100 to 5000 μs are located along PC1 from negative to positive. The plots from 10 to 50 μs are located along PC2 from negative to positive. The contribution of each component was 95.07% on PC1 and 3.08% on PC2. Changes in the peak intensities were extracted from the PCA-loading plot (Figure 2b). In the loading plot of PC1 (Figure 2b, dark gray plot), the loading values at 77.4, 69.3, 65.2 and 62.8 p.p.m. were negative, whereas those at 105.2, 89.1, 74.7, 72.4 and 71.3 p.p.m. were positive. In the loading plot of PC2 (Figure 2b, light gray plot), the loading values at 108.5 p.p.m., 105.8, 92.3, 90.4, 84.5, 75.3, 72.5 and 65.4 p.p.m. were negative, whereas those at 103.6, 87.9, 73.7, 70.2, 64.2 and 61.7 p.p.m. were positive.
PCA results of the CPVC spectra of 13C BC. In the score plot (a), components with contact times of 10–5000 μs are colored from light to dark. The contributions of PC1 and PC2 were 95.07% and 3.08%, respectively. In the loading plot (b), the loading plots of PC1 and PC2 are plotted as light gray and dark black, respectively. The bold bars show the 5% level of the rejection rate in the t-distribution of the loading values.
Figures 3a and b show the variation trend of the relative peak intensities with the contact time (the peaks were extracted from the PCA-loading plots of PC1 and PC2, respectively). The relative intensity of the maximum of each CP curve was set to be 1.0. As shown in Figure 3a, the region spanning from 100 to 5000 μs exhibits a remarkable variation in its CP curve trend. This trend corresponds to the results of the PCA score plot. In Figure 3b, there are no clear trends in the CP curves obtained. When we focused on the region from 10 to 100 μs (Figure 3c) and considered the PC2 direction of the PCA score plot, a slight trend appeared in the change from 20 to 50 μs, although it was not clear.
Relative intensity change in peaks extracted from the PCA-loading plot. The maximum value of each CP curve was fixed at 1.0. (a) Filled and open triangles show the relative intensities of peaks extracted from the PC1-loading plot with negative and positive loading values, respectively. (b) Filled and open circles show the relative intensities of peaks extracted from the PC2-loading plot with negative and positive loading values, respectively. (c) The contact time region from 10 to 100 μs in (b). (d) The contact time region from 10 to 100 μs of peaks extracted from another PCA using only the 10–50 μs CT matrix.
To remove the contribution from the PC1 axis, another PCA was performed using the matrix consisting of the spectra at 10, 20, 30 and 50 μs (Supplementary Figure S1). From PC1 of this PCA-loading plot (Supplementary Figure S1b, black plot), the chemical shifts at 105.3 p.p.m., 89.3, 74.9, 72.5 and 71.9 p.p.m. were extracted as peaks with negative loading values, whereas the chemical shifts at 103.3, 69.9, 64.6 and 62.2 p.p.m. were extracted as peaks with positive loading values. When we focused on the region from 10 to 100 μs of the extracted peaks from the second PCA (Figure 3d), the relative peak intensity trend was separated between peaks with positive and negative loading values. The extracted chemical shifts are shown in Table 1b.
Chemical shift assignments from 2D spectra
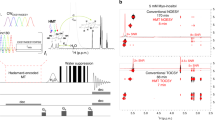
Table 1a shows the chemical shifts assigned from the 2D DARR10, 11, 12 spectrum (Figure 4a) and the 2D CP-INADEQUATE spectrum (Figure 4b) of 13C BC. In the 2D DARR spectrum (Figure 4a), the assigned chemical shifts at 104.9 (C1), 71.7 (C2), 74.5 (C3), 70.9 (C5) and 65.2 p.p.m. (C6) were similar to those of Iα crystalline material from previous reports.18, 19 In the 2D CP-INADEQUATE spectrum (Figure 4b), the chemical shift at 103.9 p.p.m. appeared as a new peak. The chemical shifts of CP-INADEQUATE are presented in Table 1a.
Peak assignment from 2D spectra of 13C BC (MAS=8 kHz, 298 K). Peaks in the 2D DARR spectrum (a) show the C-C correlations of relatively near carbons in the structure (the mixing time for this experiment was 5 ms), whereas peaks in the CP-INADEQUATE spectrum (b) show the C-C peaks with the JCC coupling (the echo time for this experiment was 3.6 ms).
Annotation of chemical shifts extracted from PCA-loading plot
The chemical shifts extracted from the PCA-loading plots were annotated (Table 1b). The extracted peaks at 105.2(3), 89.1(3), 74.7(9), 72.4(5), 71.9 and 65.2 p.p.m. were annotated as C1, C4, C3, C5, C2 and C6 of crystalline type Iα, respectively. The extracted peaks at 77.4, 69.9 and 69.3 p.p.m. could not be annotated from the assigned chemical shifts.
Discussion
PCA easily extracted the trend of peak intensity changes from the CPVC spectra
To the best of our knowledge, this study represents the first attempt to analyze NMR spectra consisting of dynamics information using PCA, and the trend of the CP curve was easily characterized by performing PCA on the CPVC spectra during a series of experiments on a single sample. The peaks at 105.2(3), 89.1(3), 74.7(9), 72.4(5) and 71.9 p.p.m. were extracted from both the first PC1 plot with a positive loading value and from the second PC1 plot with a negative loading value. They were annotated as carbons of crystalline type Iα (Table 1b), although the peak at 71.9 p.p.m. (71.3 p.p.m.) was not clearly in the same character as the loading values. The trends of the CP curve were the same over the entire contact time region (Figure 3a, open triangle) and the region from 10 to 50 μs (Figure 3d, filled triangle). This similarity means that their peaks have similar mobilities because they belong to the CH carbon of crystalline type Iα. Although the extracted peak at 65.2 p.p.m. was annotated as the same crystalline type Iα, it was extracted from the first PC1 plot with a negative loading value (Table 1b), and the trend of the CP curve was different from the peaks of the other Iα carbons, as described above (Figure 3a). This difference was mainly observed because the local proton environment was different (that is, CH2 in C6 and CH in the other carbons).
Changes in the contact time from 10 to 50 μs are reflected on PC2 of the score plot along with the contact time (Figure 2a), although the later change of the relative peak intensity did not follow a clear trend (Figure 3b). Because PC1 was dominant in the first PCA performed (Figure 3c, PC1 contribution of 95.07%), it may have hindered the extraction of the peak change from this data set; consequently, the second PCA (Supplementary Figure S1) was performed without the 100 to 5000-μs spectra, which contributed to PC1 of the first PCA. The peaks extracted from PC1 of the second PCA showed a clear trend in their CP curves (Figure 3d), and the peaks that appeared to be extracted from the noise contribution were removed from their peaks. The CP curves of the CH2 and CH signals have two stages: the first tens of microseconds are produced by the energy exchange between directly bonded 1H spins, and the second stage is associated with 1H spin diffusion.20 When the CP value of 100 μs was substituted into the equation for the spin diffusion time and distance in cellulose, the calculated distance was c.a. 0.26 nm.21 A comparison with the calculated distances from the cellulose crystalline Iα model (Supplementary Figure S2) suggests that the CP curves in Figure 3d are produced by direct-bonded 1H interaction. By selecting the data for analysis on the basis of the first PCA result, the change in the contact time in the range of 10 to 50 μs, in which energy exchange by direct-bonded 1H is the main effect, was characterized more clearly in the second PCA.
Possible implications for unannotated peaks
The peak at 62.8 p.p.m., annotated as amorphous form C6, exhibited a similar trend in the region from 100 to 2000 μs to that of the peak at 65.2 p.p.m., which is annotated as crystalline Iα (Figure 3a). These peaks also exhibited a weak trend in the decay region from 2000 to 5000 μs (Figure 3a), where T1ρH, which is associated with molecular motion, is dominant. This trend shows that the difference between the crystalline and amorphous structures of BC appears in the T1ρH region. Considering the unannotated peak at 77.4 p.p.m. (Figure 3a), the T1ρH region exhibited a similar decay to that of the peak at 62.8 p.p.m. This similarity suggests that the extracted peak at 77.4 p.p.m. could be a component from the amorphous region. The 2D DARR spectrum also shows the C6 amorphous peak, with a relatively short mixing time of 5 ms (Figure 4a), correlated with the 75.0-p.p.m. peak (Table 1a). Therefore, the existence of the C5 amorphous structure is suggested to occur at ∼75.0 p.p.m. In a previous report,22 the C5 chemical shift of cellulose was discussed using quantum chemistry calculations, and it was reported that the C5 chemical shift decreased by ∼5 p.p.m. when the side-group conformation changed from gt or gg to tg. Additionally, by considering the solid-state NMR results for the conformations of oligosaccharides and cellulose23 and chemical shift assignment,18, 24 it has been suggested that the C5 line appears at ∼76 p.p.m. in the disordered component, whereas the line for the crystalline component is observed as a doublet at ∼73 and 72 p.p.m. This downfield shift of the C5 carbon induced by the change from tg to gt is in good agreement with the theoretical prediction. Our suggestion concurs with this previous report,22 and we suggest that the C5 component at 77.4 p.p.m. extracted in our report is produced by the gt conformation of the C5-C6 side chain.
The peaks at 103.3 and 69.9 p.p.m. exhibited the same trend in Figure 3d, and are likely to represent the same component. However, these peaks and the peak at 64.6 p.p.m. could not be annotated because there are no previous assignments of cellulose, but there are assignments of plant cell wall pectin at ∼69 and 101 p.p.m.25
Normalization
As a result of this attempt, the spectral normalization (Figure 1c) of PCA was relevant for the extraction of the changes described above. At first, the PCA performed with the raw intensity data mainly showed a trend of increasing intensity with contact time (Supplementary Figure S3, PC1 98.49%). Then, to extract spectral differences other than the intensity change, each spectrum was normalized by dividing the intensity by the area. During this attempt, we assumed that the normalized spectra would emphasize the change in local proton environments and mobility because there was only a single variable in this attempt: the contact time. The CP curve for each chemical shift is known to change in a way that depends on its environment and mobility. When the PCA results and CPVC behaviors in the chemical shifts discussed above were compared, the PCA results from the normalization provided adequate trends in most chemical shift regions of the CPVC experiments.
In the score plot of the PCA (Figure 2a), the principal components at 20 and 250 μs were close to each other. In the raw and normalized spectra of those components and the 100 μs component (Supplementary Figure S4), the raw spectra (Supplementary Figure S4a) had different intensities over the entire chemical shift region, whereas the normalized spectra (Supplementary Figure S4c) had similar intensity bins between 20 and 250 μs (c.a. 64–95 p.p.m.), which was greater than the similarity between 100 and 250 μs (c.a. 55–64 p.p.m.). This effect was observed because the differences in the intensities were canceled out by the opposite changes of the normalized intensities (Supplementary Figure S4b). In the C4 amorphous region of the normalized spectra (Figure 1c and Supplementary Figure S4b, c.a. 84 p.p.m.), the differences among the various CT spectra were also canceled out by the normalization. The absolute intensity of the C4 amorphous region (Figure 1b and Supplementary Figure S4a, c.a. 84 p.p.m.) was small in each spectrum such that the difference between the spectra in the chemical shift region (Figure 1c and Supplementary Figure S4b, c.a. 84 p.p.m.) became much smaller than those in the other region after normalization. For more detailed information, the normalization should be improved.
Future improvement of this method
By performing the reported method of PCA on the CPVC spectra, the variation tendency over the entire spectrum was visualized, and the components that differed in CHn and crystallinity were partially extracted, whereas efficient methods for extracting information from the C4 region have not yet been developed. In a previous study, a less-ordered (para-crystalline) in core structure was investigated using partial signal suppression by spin-lattice relaxation.26 In this paper, the difference in the CP curves appeared in the T1ρH region. Therefore, the pulse sequence for collecting the spectra must be improved. In previous investigations of plant cell walls,25, 27 which are more complex examples of biomass systems containing cellulose, various types of relaxation times have been utilized to separate components such as cellulose, pectin and hemicellulose. In particular, proton spin relaxation editing using T1ρH has provided the best discrimination between cellulose and other cell wall components.28 Additionally, from a study using model systems and plant cell walls, it has been shown that long T1ρH components are associated with crystalline cellulose, implying that the proton spin diffusion between the cellulose crystalline microfibrils and the surface amorphous cellulose is less efficient on the time scale of T1ρH than on that of T1H.29 A further improved method for collecting spectra could be applied to support the assignments from the CP spectra, such as the example described above for the C5 amorphous structure signal candidate.
References
Pines, A. & Shattuck, T. W. C-13 proton NMR cross-polarization times in solid adamantane. J. Chem. Phys. 61, 1255–1256 (1974).
Ando, I. Koubunshi no kotai NMR, in Japanese 14–18 (Koudansha scientific, Tokyo, 1994).
Cory, D. G. & Ritchey, W. M. Inversion recovery cross-polarization NMR in solid semicrystalline polymers. Macromolecules 22, 1611–1615 (1989).
Seierstad, T., Roe, K., Sitter, B., Halgunset, J., Flatmark, K., Ree, A. H., Olsen, D. R., Gribbestad, I. S. & Bathen, T. F. Principal component analysis for the comparison of metabolic profiles from human rectal cancer biopsies and colorectal xenografts using high-resolution magic angle spinning (1)H magnetic resonance spectroscopy. Mol. Cancer 7 (2008) (http://www.molecular-cancer.com/content/7/1/33).
Sekiyama, Y., Chikayama, E. & Kikuchi, J. Evaluation of a semipolar solvent system as a step toward heteronuclear multidimensional NMR-based metabolomics for (13)C-labelled bacteria, plants, and animals. Anal. Chem., Washington, DC, U. S. 83, 719–726 (2011).
Yamamoto, H., Horii, F. & Hirai, A. Structural studies of bacterial cellulose through the solid-phase nitration and acetylation by CP/MAS C-13 NMR spectroscopy. Cellulose 13, 327–342 (2006).
Yamamoto, H. & Horii, F. Cp Mas C-13 NMR analysis of the crystal transformation induced for valonia cellulose by annealing at high-temperatures. Macromolecules 26, 1313–1317 (1993).
Wickholm, K., Larsson, P. T. & Iversen, T. Assignment of non-crystalline forms in cellulose I by CP/MAS C-13 NMR spectroscopy. Carbohydr. Res. 312, 123–129 (1998).
Hestrin, S. & Schramm, M. Synthesis of cellulose by Acetobacter xylinum. II. Preparation of freeze-dried cells capable of polymerizing glucose to cellulose. Biochem. J. 58, 345–352 (1954).
Ohashi, R. & Takegoshi, K. Asymmetric (13)C-(13)C polarization transfer under dipolar-assisted rotational resonance in magic-angle spinning NMR. J. Chem. Phys. 125, 214503 (2006).
Takegoshi, K., Nakamura, S. & Terao, T. C-13-H-1 dipolar-driven C-13-C-13 recoupling without C-13 rf irradiation in nuclear magnetic resonance of rotating solids. J. Chem. Phys. 118, 2325–2341 (2003).
Takegoshi, K., Nakamura, S. & Terao, T. C-13-H-1 dipolar-assisted rotational resonance in magic-angle spinning NMR. Chem. Phys. Lett. 344, 631–637 (2001).
Lesage, A., Auger, C., Caldarelli, S. & Emsley, L. Determination of through-bond carbon-carbon connectivities in solid-state NMR using the INADEQUATE experiment. J. Am. Chem. Soc. 119, 7867–7868 (1997).
Nakanishi, Y., Fukuda, S., Chikayama, E., Kimura, Y., Ohno, H. & Kikuchi, J. Dynamic omics approach identifies nutrition-mediated microbial interactions. J. Proteome Res. 10, 824–836 (2011).
Sekiyama, Y., Chikayama, E. & Kikuchi, J. Profiling polar and semipolar plant metabolites throughout extraction processes using a combined solution-state and high-resolution magic angle spinning NMR approach. Anal. Chem., (Washington, DC, U. S.)82, 1643–1652 (2010).
Fukuda, S., Nakanishi, Y., Chikayama, E., Ohno, H., Hino, T. & Kikuchi, J. Evaluation and characterization of bacterial metabolic dynamics with a novel profiling technique, real-time metabolotyping. PLoS One 4, e4893 (2009).
Tian, C., Chikayama, E., Tsuboi, Y., Kuromori, T., Shinozaki, K., Kikuchi, J. & Hirayama, T. Top-down phenomics of Arabidopsis thaliana: metabolic profiling by one- and two-dimensional nuclear magnetic resonance spectroscopy and transcriptome analysis of albino mutants. J. Biol. Chem. 282, 18532–18541 (2007).
Kono, H., Erata, T. & Takai, M. Determination of the through-bond carbon-carbon and carbon-proton connectivities of the native celluloses in the solid state. Macromolecules 36, 5131–5138 (2003).
Hesse-Ertelt, S., Witter, R., Ulrich, A. S., Kondo, T. & Heinze, T. Spectral assignments and anisotropy data of cellulose I-alpha: C-13-NMR chemical shift data of cellulose I-alpha determined by INADEQUATE and RAI techniques applied to uniformly C-13-labeled bacterial celluloses of different Gluconacetobacter xylinus strains. Magn. Reson. Chem. 46, 1030–1036 (2008).
Xiaoling, W., Shanmin, Z. & Xuewen, W. Two-stage feature of Hartmann-Hahn cross relaxation in magic-angle sample spinning. Phys. Rev. B: Condens. Matter Mater. Phys. 37, 9827–9829 (1988).
Masuda, K., Adachi, M., Hirai, A., Yamamoto, H., Kaji, H. & Horii, F. Solid-state C-13 and H-1 spin diffusion NMR analyses of the microfibril structure for bacterial cellulose. Solid State Nucl. Magn. Reson. 23, 198–212 (2003).
Suzuki, S., Horii, F. & Kurosu, H. Theoretical investigations of (13)C chemical shifts in glucose, cellobiose, and native cellulose by quantum chemistry calculations. J. Mol. Struct. 921, 219–226 (2009).
Horii, F., Hirai, A. & Kitamaru, R. Solid-state C-13-Nmr study of conformations of oligosaccharides and cellulose—conformation of Ch2oh Group About the Exo-Cyclic C-C Bond. Polym. Bull. 10, 357–361 (1983).
Kono, H., Yunoki, S., Shikano, T., Fujiwara, M., Erata, T. & Takai, M. CP/MAS (13)C NMR study of cellulose and cellulose derivatives. 1. Complete assignment of the CP/MAS (13)C NMR spectrum of the native cellulose. J. Am. Chem. Soc. 124, 7506–7511 (2002).
Tang, H. R., Belton, P. S., Ng, A. & Ryden, P. C-13 MAS NMR studies of the effects of hydration on the cell walls of potatoes and Chinese water chestnuts. J. Agric. Food Chem. 47, 510–517 (1999).
Larsson, P. T., Wickholm, K., Iversen, T. & CP, A /MAS C-13 NMR investigation of molecular ordering in celluloses. Carbohydr. Res. 302, 19–25 (1997).
Tekely, P. & Vignon, M. R. Proton T1 and T2 relaxation-times of wood components using C-13 Cp Mas R. J. Polym. Sci., Part C: Polym. Lett. 25, 257–261 (1987).
Newman, R. H., Davies, L. M. & Harris, P. J. Solid-state C-13 nuclear magnetic resonance characterization of cellulose in the cell walls of Arabidopsis thaliana leaves. Plant Physiol. 111, 475–485 (1996).
Tang, H. R., Wang, Y. L. & Belton, P. S. C-13 CPMAS studies of plant cell wall materials and model systems using proton relaxation-induced spectral editing techniques. Solid State Nucl. Magn. Reson. 15, 239–248 (2000).
Acknowledgements
We thank Yoshiyuki Ogata, Tetsuya Mori and Yuuri Tsuboi for helpful discussions and providing highly cleaned BC. This research was supported in part by Grants-in-Aid for Scientific Research for challenging exploratory research from the Ministry of Education, Culture, Sports, Science and Technology, Japan (MEXT to JK). This work was also supported in part by grants from the New Energy and Industrial Technology Development Organization (NEDO to JK).
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on Polymer Journal website
Supplementary information
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Okushita, K., Komatsu, T., Chikayama, E. et al. Statistical approach for solid-state NMR spectra of cellulose derived from a series of variable parameters. Polym J 44, 895–900 (2012). https://doi.org/10.1038/pj.2012.82
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/pj.2012.82
Keywords
This article is cited by
-
Effective degradation of cellulose by Microwave irradiation in alkaline solution
Cellulose (2021)
-
Isolation and Characterization of Nanocrystalline Cellulose from Totally Chlorine Free Oil Palm Empty Fruit Bunch Pulp
Journal of Polymers and the Environment (2017)
-
Organosolv pretreatment of sorghum bagasse using a low concentration of hydrophobic solvents such as 1-butanol or 1-pentanol
Biotechnology for Biofuels (2016)
-
Modification of plant cell wall structure accompanied by enhancement of saccharification efficiency using a chemical, lasalocid sodium
Scientific Reports (2016)