Abstract
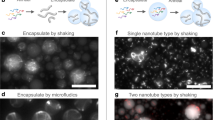
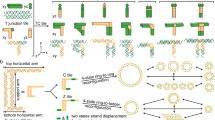
DNA nanotubes can template the growth of nanowires1, orient transmembrane proteins for nuclear magnetic resonance determination2, and can potentially act as stiff interconnects, tracks for molecular motors and nanoscale drug carriers3. Current methods for the construction of DNA nanotubes result in symmetrical and cylindrical assemblies that are entirely double-stranded2,4,5,6,7,8,9,10,11. Here, we report a modular approach to DNA nanotube synthesis that provides access to geometrically well-defined triangular and square-shaped DNA nanotubes. We also construct the first nanotube assemblies that can exist in double- and single-stranded forms with significantly different stiffness. This approach allows for parameters such as geometry, stiffness, and single- or double-stranded character to be fine-tuned, and could enable the creation of designer nanotubes for a range of applications, including the growth of nanowires of controlled shape, the loading and release of cargo, and the real-time modulation of stiffness and persistence length within DNA interconnects.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Yan, H., Park, S. H., Finkelstein, G., Reif, J. H. & LaBean, T. H. DNA-templated self-assembly of protein arrays and highly conductive nanowires. Science 301, 1882–1884 (2003).
Douglas, S. M., Chou, J. J. & Shih, W. M. DNA-nanotube-induced alignment of membrane proteins for NMR structure determination. Proc. Natl Acad. Sci. USA 104, 6644–6648 (2007).
Geng, Y. et al. Shape effects of filaments versus spherical particles in flow and drug delivery. Nature Nanotech. 2, 249–255 (2007).
Rothemund, P. W. K. et al. Design and characterization of programmable DNA nanotubes. J. Am. Chem. Soc. 126, 16344–16352 (2004).
Mitchell, J. C., Harris, J. R., Malo, J., Bath, J. & Turberfield, A. J. Self-assembly of chiral DNA nanotubes. J. Am. Chem. Soc. 126, 16342–16343 (2004).
Sherman, W. B. & Seeman, N. C. Design of minimally strained nucleic acid nanotubes. Biophys. J. 90, 4546–4557 (2006).
Endo, M., Seeman, C. & Majima, T. DNA tube structures controlled by a four-way-branched DNA connector. Angew. Chem. Int. Ed. 44, 6074–6077 (2005).
Park, S. H. et al. Three-helix bundle DNA tiles self-assemble into 2D lattice or 1D templates for silver nanowires. Nano Lett. 5, 693–696 (2005)
Liu, D., Park, S. H., Reif, J. H. & LaBean, T. H. DNA nanotubes self-assembled from triple-crossover tiles as templates for conductive nanowires. Proc. Natl Acad. Sci. USA 101, 717–722 (2004).
Mathieu, F. et al. Six-helix bundles designed from DNA. Nano Lett. 5, 661–665 (2005).
Kuzuya, A., Wang, R., Sha, R. & Seeman, N. C. Six-helix and eight-helix DNA nanotubes assembled from half-tubes. Nano Lett. 7, 1757–1763 (2007).
Aldaye, F. A. & Sleiman, H. F. Sequential self-assembly of a DNA hexagon as a template for the organization of gold nanoparticles. Angew. Chem. Int. Ed. 118, 2204–2209 (2006).
Aldaye, F. A. & Sleiman, H. F. Dynamic DNA templates for discrete gold nanoparticle assemblies: Control of geometry, modularity, write/erase and structural switching. J. Am. Chem. Soc. 129, 4130–4131 (2007).
Aldaye, F. A. & Sleiman, H. F. Modular access to structurally switchable 3D discrete DNA assemblies. J. Am. Chem. Soc. 129, 13376–13377 (2007).
Aldaye, F. A., Palmer, A. L. & Sleiman, H. F. Assembling materials with DNA as the guide. Science 321, 1795–1799 (2008).
Acknowledgements
The authors would like to thank the Natural Sciences and Engineering Research Council of Canada, the Canada Foundation for Innovation, the Centre for Self-Assembled Chemical Structures, and the Canadian Institute for Advanced Research for financial support, A.L. Palmer for help in manuscript preparation, and J. Hedberg for help in preparing the graphical illustrations. F.A.A is a McGill University Principal's Prize Fellow. P.K.L. and P.K. thank CIHR for a Chemical Biology Scholarship. H.F.S is a Cottrell Scholar of the Research Corporation.
Author information
Authors and Affiliations
Contributions
All authors discussed the results and commented on the manuscript. H.F.S. conceived and designed the project, analysed the data and co-wrote the paper. F.A.A conceived and designed the project, performed the experiments, analysed the data and co-wrote the paper. P.K.L performed the experiments and analysed the data. P.K. and G.C. performed the confocal fluorescence microscopy measurements. C.K.M. assisted in project design.
Corresponding author
Supplementary information
Supplementary Information
Supplementary information (PDF 3540 kb)
Rights and permissions
About this article
Cite this article
Aldaye, F., Lo, P., Karam, P. et al. Modular construction of DNA nanotubes of tunable geometry and single- or double-stranded character. Nature Nanotech 4, 349–352 (2009). https://doi.org/10.1038/nnano.2009.72
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nnano.2009.72
This article is cited by
-
DNA nanostructures as templates for biomineralization
Nature Reviews Chemistry (2021)
-
Gold nanocrystals with DNA-directed morphologies
Nature Communications (2016)
-
Stepwise growth of surface-grafted DNA nanotubes visualized at the single-molecule level
Nature Chemistry (2015)
-
DNA hydrogel-based supercapacitors operating in physiological fluids
Scientific Reports (2013)
-
Molecular logic computing model based on DNA self-assembly strand branch migration
Chinese Science Bulletin (2013)