Abstract
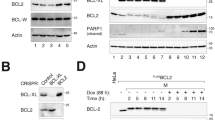
The pro-apoptotic molecule BAD binds BCL-xL or BCL2 and inactivates their survival function. In addition to their anti-apoptotic function, BCL2 and BCL-xL also delay cell cycle entry from quiescence. We found that the BH3-only molecule BAD also exerted a cell cycle effect. BAD expression resulted in failure to cell cycle block in growth arrest conditions. In low serum and in confluence, fibroblasts constitutively or inducibly expressing BAD persisted in S phase, continued to incorporate BrdU, and exhibited sustained cyclin E/cdk2 activity. Mutation analysis indicated that the cell cycle effect of BAD was not dependent on its phosphorylation status or subcellular localization, but strictly co-segregated with BCL-xL binding. bclx−/− MEFs expressing BAD and bad−/− MEFs both arrested in G0/G1 in low serum similar to wild-type controls, suggesting that the ability to overcome the G0/G1 checkpoint resulted from the presence of BAD/BCL-xL heterodimers, rather than the absence of BCL-xL or BAD. These data provide evidence that in addition to regulating apoptosis, the BAD/BCL-xL heterodimer has a novel cell cycle function.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Adams JM, Cory S . 1998 Science 281: 1322–1326
Ayllon V, Martinez AC, Garcia A, Cayla X, Rebollo A . 2000 EMBO J. 19: 2237–2246
Brady HJ, Gil-Gomez G, Kirberg J, Berns AJ . 1996 EMBO J. 15: 6991–7001
Chiang C-W, Harris G, Ellig C, Masters SC, Subramanian R, Shenolikar S, Wadzinski BE, Yang E . 2001 Blood 97: 1289–1297
Clarke AR, Maandag ER, van Roon M, van der Lugt NM, van der Valk M, Hooper ML, Berns A, te Riele H . 1992 Nature 359: 328–330
Datta SR, Dudek H, Tao X, Masters S, Fu H, Gotoh Y, Greenberg ME . 1997 Cell 91: 231–241
Datta SR, Katsove A, Hu L, Petros A, Fesik SW, Yaffe MB, Greenberg ME . 2000 Mol. Cell 6: 41–51
Donehower LA, Harvey M, Slagle BL, McArthur MJ, Montgomery Jr CA, Butel JS, Bradley A . 1992 Nature 356: 215–221
Furth PA, Bar-Peled U, Li M, Lewis A, Laucirica R, Jager R, Weiher H, Russell RG . 1999 Oncogene 18: 6589–6596
Gil-Gomez G, Berns A, Brady HJ . 1998 EMBO J. 17: 7209–7218
Gossen M, Bujard H . 1992 Proc. Natl. Acad. Sci. USA 89: 5547–5551
Hakem A, Sasaki T, Kozieradzki I, Penninger JM . 1999 J. Exp. Med. 189: 957–968
Harada H, Becknell B, Wilm M, Mann M, Huang LJ, Taylor SS, Scott JD, Korsmeyer SJ . 1999 Mol. Cell. 3: 413–422
Huang DC, O'Reilly LA, Strasser A, Cory S . 1997 EMBO J. 16: 4628–4638
Jacks T, Fazeli A, Schmitt EM, Bronson RT, Goodell MA, Weinberg RA . 1992 Nature 359: 295–300
Jacks T, Remington L, Williams BO, Schmitt EM, Halachmi S, Bronson RT, Weinberg RA . 1994 Curr. Biol. 4: 1–7
Kelekar A, Chang BS, Harlan JE, Fesik SW, Thompson CB . 1997 Mol. Cell Biol. 17: 7040–7046
Kelekar A, Thompson CB . 1998 Trends Cell. Biol. 8: 324–330
Korsmeyer SJ . 1999 Cancer Res. 59: 1693s–1700s
Krishan A . 1975 J. Cell. Biol. 66: 188–193
Lee EY, Chang CY, Hu N, Wang YC, Lai CC, Herrup K, Lee WH, Bradley A . 1992 Nature 359: 288–294
Li H, Zhu H, Xu CJ, Yuan J . 1998 Cell 94: 491–501
Lind EF, Wayne J, Wang QZ, Staeva T, Stolzer A, Petrie HT . 1999 J. Immunol. 162: 5374–5379
Linette GP, Hess JL, Sentman CL, Korsmeyer SJ . 1995 Blood 86: 1255–1260
Linette GP, Li Y, Roth K, Korsmeyer SJ . 1996 Proc. Natl. Acad. Sci. USA 93: 9545–9552
Luo X, Budihardjo I, Zou H, Slaughter C, Wang X . 1998 Cell 94: 481–490
Lutz RJ . 2000 Biochem. Soc. Trans. 28: 51–56
Mazel S, Burtrum D, Petrie HT . 1996 J. Exp. Med. 183: 2219–2226
McDonnell TJ, Korsmeyer SJ . 1991 Nature 349: 254–256
Murphy KL, Kittrell FS, Gay JP, Jager R, Medina D, Rosen JM . 1999 Oncogene 18: 6597–6604
Nagata S . 1999 Nat. Cell. Biol. 1: E143–E145
O'Reilly LA, Huang DC, Strasser A . 1996 EMBO J. 15: 6979–6990
Oda E, Ohki R, Murasawa H, Nemoto J, Shibue T, Yamashita T, Tokino T, Taniguchi T, Tanaka N . 2000 Science 288: 1053–1058
Pandey S, Wang E . 1995 J. Cell. Biochem. 58: 135–150
Pear WS, Nolan GP, Scott ML, Baltimore D . 1993 Proc. Natl. Acad. Sci. USA 90: 8392–8396
Puthalakath H, Huang DC, O'Reilly LA, King SM, Strasser A . 1999 Mol. Cell 3: 287–296
Schurmann A, Mooney AF, Sanders LC, Sells MA, Wang HG, Reed JC, Bokoch GM . 2000 Mol. Cell. Biol. 20: 453–461
Tan Y, Demeter MR, Ruan H, Comb MJ . 2000 J. Biol. Chem. 275: 25865–25869
Vairo G, Innes KM, Adams JM . 1996 Oncogene 13: 1511–1519
Vairo G, Soos TJ, Upton TM, Zalvide J, DeCaprio JA, Ewen ME, Koff A, Adams JM . 2000 Mol. Cell. Biol. 20: 4745–4753
Wang HG, Pathan N, Ethell IM, Krajewski S, Yamaguchi Y, Shibasaki F, McKeon F, Bobo T, Franke TF, Reed JC . 1999 Science 284: 339–343
Yang E, Zha J, Jockel J, Boise LH, Thompson CB, Korsmeyer SJ . 1995 Cell 80: 285–291
Zha J, Harada H, Osipov K, Jockel J, Waksman G, Korsmeyer SJ . 1997 J. Biol. Chem. 272: 24101–24104
Zha J, Harada H, Yang E, Jockel J, Korsmeyer SJ . 1996 Cell 87: 619–628
Zhou XM, Liu Y, Payne G, Lutz RJ, Chittenden T . 2000 J. Biol. Chem. 275: 25046–25051
Acknowledgements
We are grateful to Dr Stanley Korsmeyer for providing bad−/− mice and 10929 anti-BAD antibody, to Dr Kevin Roth for providing bclx+/− mice, and to Dr Lawrence Boise for anti-BCL-x antibodies. We thank Courtney Greider for assistance with kinase assays and Dr Jeffrey Rottman for expert computer assistance. We thank Drs Stephen Brandt, Scott Hiebert, and Mark Koury for helpful discussions and critical reading of the manuscript. Experiments were performed in part through use of the Vanderbilt University Medical Center Cell Imaging Resource supported by CA68485 and DK20593. This work was supported by NIH 1RO1CA78443 grant, and grants from Pfizer, Inc. and the V Foundation.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Chattopadhyay, A., Chiang, CW. & Yang, E. BAD/BCL-xL heterodimerization leads to bypass of G0/G1 arrest. Oncogene 20, 4507–4518 (2001). https://doi.org/10.1038/sj.onc.1204584
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1204584
Keywords
This article is cited by
-
Non-canonical BAD activity regulates breast cancer cell and tumor growth via 14-3-3 binding and mitochondrial metabolism
Oncogene (2019)
-
BAD overexpression inhibits cell growth and induces apoptosis via mitochondrial-dependent pathway in non-small cell lung cancer
Cancer Cell International (2013)
-
Digital gene expression profiling of primary acute lymphoblastic leukemia cells
Leukemia (2012)
-
In vitro analysis of ovarian cancer response to cisplatin, carboplatin, and paclitaxel identifies common pathways that are also associated with overall patient survival
British Journal of Cancer (2012)
-
The BH3-only protein Bad confers breast cancer taxane sensitivity through a nonapoptotic mechanism
Oncogene (2010)